Back
How to Study Microbiology for USMLE Step 1 2026: High-Yield Organisms, Mechanisms and Exam Strategy
Master USMLE Step 1 microbiology with this complete 2026 guide. Learn high-yield organisms, mechanisms, and proven study strategies to score high on 15-20% of the exam.

How to Study Microbiology for USMLE Step 1 2026: High-Yield Organisms, Mechanisms and Exam Strategy
You are staring at a 600-page microbiology textbook, and USMLE Step 1 is 12 weeks away. The sheer volume feels overwhelming — bacteria, viruses, fungi, parasites, all with their specific mechanisms, treatments, and clinical presentations. Here's what most students dont realize: microbiology makes up 15-20% of Step 1, but only about 200 high-yield facts actually get tested repeatedly.
The exam doesnt care if you can recite every detail about Mycobacterium marinum. It wants you to instantly recognize that a patient with swimming pool exposure and slowly healing skin lesions has atypical mycobacterial infection. The difference? Pattern recognition built through mechanism-first learning, not textbook memorization.
Most students waste 4-6 weeks passively re-reading microbiology chapters. This guide gives you a laser-focused strategy: which organisms to prioritize, how to learn mechanisms that stick, and how to build automatic bug identification by exam day.
Why Microbiology is High-Yield (and Why Students Struggle)
USMLE Step 1 microbiology questions test 3 core skills:
1. Organism identification from clinical vignettes (gram stain, growth characteristics, epidemiology)
2. Mechanism understanding (toxins, virulence factors, antibiotic targets)
3. Clinical correlation (which bug causes which syndrome)
The problem? Most students approach it backwards. They memorize organism lists without understanding the underlying mechanisms that explain why certain bugs cause specific diseases. When Step 1 presents a novel scenario — say, a patient with honey-colored crusts and bullous lesions — they cant connect staphylococcal exfoliative toxin mechanism to the clinical presentation.
Effective microbiology study builds these connections systematically. Start with high-yield organisms, layer in mechanisms, then drill clinical correlations until pattern recognition becomes automatic.
The High-Yield Organism Priority System
Not all microorganisms are created equal on Step 1. Focus your energy on these categories, in order:
Tier 1: Must-Know Organisms (80% of microbiology points)
Gram-Positive Bacteria:
Staphylococcus aureus (MRSA mechanisms, toxins, clinical syndromes)
Streptococcus pyogenes (Group A strep, necrotizing fasciitis, rheumatic fever)
Streptococcus pneumoniae (encapsulated, vaccine strains, resistance patterns)
Enterococcus (VRE mechanisms, intrinsic resistance)
Listeria monocytogenes (intracellular growth, neonatal meningitis)
Gram-Negative Bacteria:
E. coli (ETEC, EHEC, UPEC — know your toxin mechanisms)
Salmonella (typhoidal vs. non-typhoidal, intracellular survival)
Neisseria species (gonorrhoeae vs. meningitidis differentiation)
Haemophilus influenzae (type b vs. non-typeable)
Pseudomonas aeruginosa (multidrug resistance, biofilm formation)
Practice with targeted bacterial microbiology questions to cement these core organisms before moving to lower-yield bugs.
Tier 2: High-Yield Viruses and Intracellular Organisms
DNA Viruses:
HSV-1/HSV-2 (latency mechanisms, clinical presentations)
EBV (oncogenic mechanisms, heterophile antibodies)
CMV (congenital infection patterns, retinitis)
HPV (oncogenic types, vaccine coverage)
RNA Viruses:
Influenza (antigenic drift vs. shift, pandemic potential)
HIV (reverse transcriptase, resistance mechanisms)
Hepatitis B and C (chronic infection, hepatocellular carcinoma)
Intracellular Bacteria:
Chlamydia species (elementary vs. reticulate bodies)
Rickettsia (Rocky Mountain spotted fever, epidemic typhus)
Mycobacterium tuberculosis (cord factor, granuloma formation)
Drill these with viral microbiology lessons and use spaced repetition through microbiology flashcards to lock in the clinical correlations.
Mechanism-First Learning Strategy
Step 1 questions dont ask "Which organism causes pneumonia?" They present a 45-year-old smoker with rapid-onset pneumonia and ask about the most likely causative organism based on gram stain and clinical course. Your job: connect the dots between mechanism and presentation.
Toxin Mechanisms (Extremely High-Yield)
ADP-Ribosylating Toxins: Understanding ADP-ribosylation explains multiple organisms at once. When you see Oncourse AI's Mnemonic challenge present "DEPTH" for ADP-ribosylating toxins (Diphtheria, E. coli heat-labile, Pertussis, Heat-labile enterotoxin), you are not just memorizing a list — you are grouping by shared mechanism. All these toxins increase cAMP through the same pathway. Cell Wall Synthesis Inhibitors: Beta-lactam antibiotics target peptidoglycan synthesis. But why dont they work against Mycoplasma? No cell wall. Why do they work against gram-positives better than gram-negatives? Thicker peptidoglycan layer, less efflux pump activity. Understanding mechanism explains the clinical choice.
Virulence Factor Categories
Group organisms by how they evade immune responses:
Capsule Formation (Encapsulated Organisms): "Some Killers Have Pretty Nice Capsules" — Streptococcus pneumoniae, Klebsiella pneumoniae, Haemophilus influenzae, Pseudomonas aeruginosa, Neisseria meningitidis, Cryptococcus neoformans. The capsule prevents phagocytosis, explaining why these cause severe invasive disease.
When studying encapsulated organisms through bacterial lessons, ask Rezzy AI tutor to explain capsule formation mechanisms rather than just memorizing the list. Understanding polysaccharide capsule structure explains why these organisms cause meningitis and sepsis.
Immune Evasion Strategies:
Antigenic variation (influenza, HIV, Neisseria)
Molecular mimicry (Group A strep M protein resembling cardiac myosin)
Complement inhibition (factor H binding by Neisseria)
Building Your Study System
Phase 1: Foundation Building (Weeks 1-2)
Start with organism identification patterns. Use the "bug chart" approach:
Organism | Gram Stain | Key Test | Clinical Syndrome |
|---|---|---|---|
S. aureus | Gram+ cocci, clusters | Catalase+, Coagulase+ | Skin/soft tissue, endocarditis |
S. pyogenes | Gram+ cocci, chains | Catalase-, Beta-hemolytic | Pharyngitis, necrotizing fasciitis |
S. pneumoniae | Gram+ diplococci | Alpha-hemolytic, Optochin sensitive | Pneumonia, meningitis |
Dont just read this chart — actively test yourself. Look at clinical vignettes and predict the organism before seeing answer choices.
Phase 2: Mechanism Integration (Weeks 3-4)
Layer mechanisms onto your organism knowledge. For each high-yield bug, know:
Primary virulence factor
Mechanism of pathogenicity
Antibiotic target and resistance mechanisms
Clinical syndrome explanation
Using Synapses Game during this phase helps cement mechanism-based groupings. When you group organisms by shared features under time pressure, you are building the same cognitive pathways Step 1 questions test.
Phase 3: Clinical Correlation (Weeks 5-6)
Connect organisms to clinical presentations. Practice with case-based questions that require you to:
Identify organism from clinical clues
Predict antibiotic choice based on organism and patient factors
Recognize complications and explain their mechanisms
Common Pitfalls and How to Avoid Them
Mistake 1: Confusing Similar Organisms
Listeria vs. Group B Strep: Both cause neonatal meningitis, but only Listeria shows tumbling motility and grows at 4°C. Both are beta-hemolytic, but Listeria is catalase-positive. Solution: Create comparison tables for commonly confused organisms. Focus on the 1-2 distinguishing features that Step 1 questions actually test.
Mistake 2: Neglecting Fungi and Parasites Until Too Late
Many students spend 80% of their time on bacteria and viruses, cramming fungi and parasites in the final week. But these represent 15-20% of microbiology questions and have predictable patterns.
High-Yield Fungi:
Candida species (pseudohyphae, germ tube test)
Cryptococcus (encapsulated yeast, India ink positive)
Histoplasma (Ohio River Valley, macrophage infection)
Coccidioides (Southwest US, spherules)
High-Yield Parasites:
Plasmodium species (malaria lifecycle, drug resistance)
Toxoplasma gondii (cat exposure, CNS lesions in AIDS)
Giardia lamblia (camping exposure, trophozoites)
Mistake 3: Passive Re-reading Instead of Active Recall
Reading microbiology notes for the 5th time creates familiarity, not mastery. Step 1 requires instant pattern recognition under time pressure.
Better approach: Use active recall methods. See a clinical vignette, predict the organism, then check your answer. Use spaced repetition through microbiology flashcards to ensure long-term retention.
Antibiotic Mechanisms and Resistance Patterns
Understanding antimicrobial mechanisms helps predict coverage and resistance patterns without memorizing every drug-bug combination.
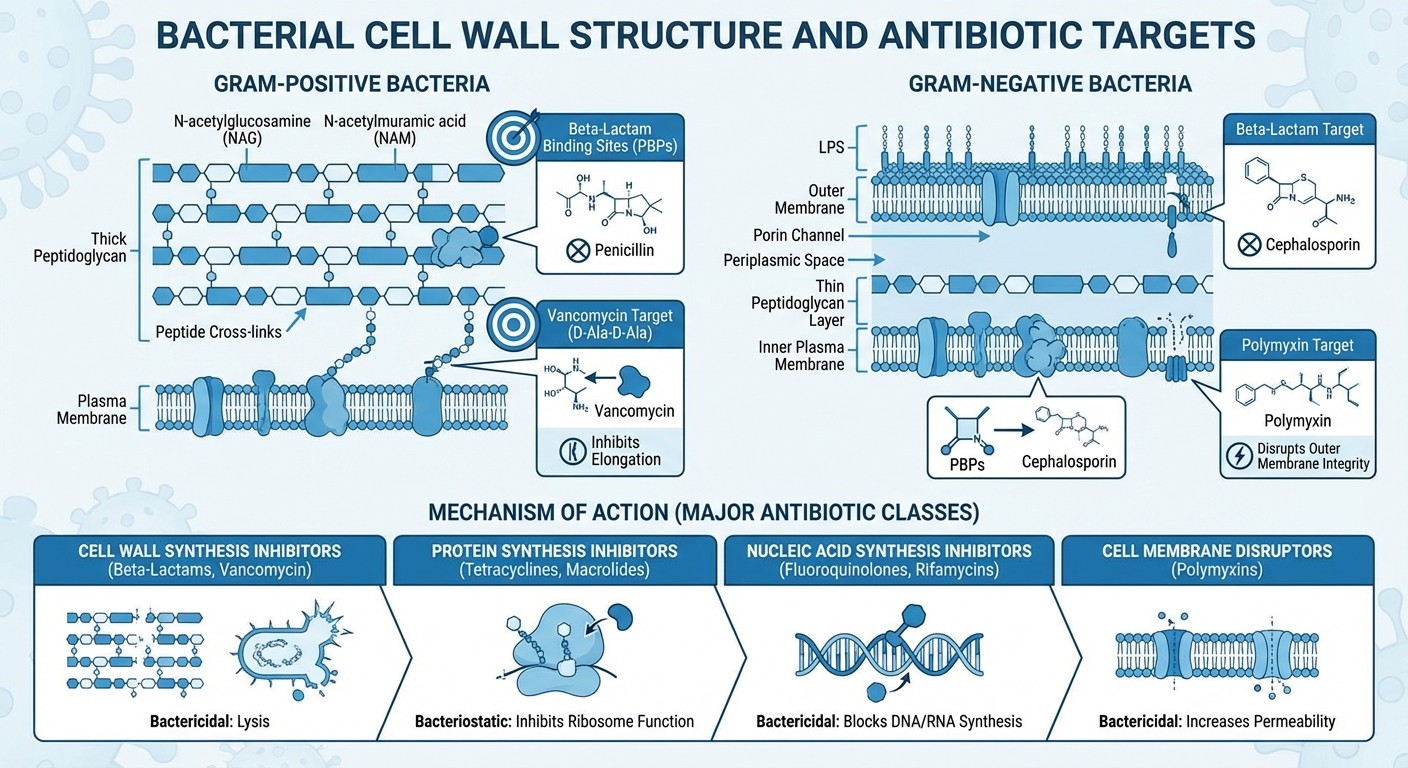
Cell Wall Synthesis Inhibitors
Beta-lactams (penicillins, cephalosporins, carbapenems):
Target: Peptidoglycan cross-linking
Spectrum: Gram-positives > gram-negatives (with exceptions)
Resistance: Beta-lactamases, altered PBPs
Clinical pearl: Dont work against Mycoplasma, Chlamydia (no cell wall)
Vancomycin:
Target: D-ala-D-ala peptide binding
Spectrum: Gram-positives only (cant cross gram-negative outer membrane)
Resistance: D-ala-D-lac substitution (VRE mechanism)
Study antibiotic resistance mechanisms to understand why certain bugs develop resistance to specific drug classes.
Protein Synthesis Inhibitors
30S Ribosome Inhibitors:
Streptomycin (irreversible)
Tetracyclines (reversible)
Aminoglycosides (bactericidal)
50S Ribosome Inhibitors:
Chloramphenicol (bacteriostatic)
Macrolides (bacteriostatic)
Lincomycin/Clindamycin
Understanding ribosome targeting explains why some antibiotics are bactericidal vs. bacteriostatic and helps predict drug interactions.
Step 1 Question Patterns and Approach
Pattern 1: Organism Identification from Clinical Vignette
Example approach: 25-year-old sexually active female presents with dysuria and urethral discharge. Gram stain shows gram-negative intracellular diplococci. Your thought process: 1. Intracellular diplococci = Neisseria 2. Sexually transmitted = N. gonorrhoeae (not N. meningitidis)
3. Confirm with oxidase positive, grows on chocolate agar
Pattern 2: Mechanism-Based Questions
Example: Patient develops diarrhea after antibiotic treatment. Which mechanism explains C. difficile toxin pathogenicity? Your approach:
1. Know that C. diff toxins A and B are glycosyltransferases
2. They inactivate Rho proteins
3. This disrupts actin cytoskeleton → intestinal epithelial damage
When encountering mechanism questions, use Rezzy AI to explore signaling pathways and ensure you understand the connection between molecular mechanism and clinical presentation.
Pattern 3: Treatment Selection
Example: Newborn develops meningitis. CSF shows gram-positive rods. Most appropriate treatment? Your approach: 1. Gram-positive rods in neonate = Listeria 2. Ampicillin is drug of choice (Listeria has intrinsic cephalosporin resistance)
3. Add gentamicin for synergy in severe cases
The 2-Week Intensive Review Strategy
If you have limited time, focus on these high-yield rapid-fire facts:
Week 1: Core Organisms and Mechanisms
Day 1-2: Gram-positive cocci (Staph, Strep species)
Day 3-4: Gram-negative rods (Enterobacteriaceae, Pseudomonas)
Day 5-6: Intracellular organisms and acid-fast bacteria
Day 7: Review and practice questions
Week 2: Integration and Clinical Correlation
Day 1-2: Viruses (DNA and RNA families)
Day 3-4: Fungi and parasites
Day 5: Antibiotic mechanisms and resistance
Day 6-7: Mixed practice and weak area review
Use bacterial questions and viral questions to test your knowledge daily during this intensive phase.
Memory Techniques That Actually Work
The TORCH Complex
Toxoplasmosis, Other (syphilis, varicella, parvovirus), Rubella, CMV, HSV — congenital infections with overlapping presentations but distinct features:
Toxoplasma: calcifications, chorioretinitis
Syphilis: snuffles, saber shins
Rubella: PDA, cataracts, deafness
CMV: periventricular calcifications
HSV: temporal lobe involvement
Encapsulated Organism Mnemonic
"SHiN SKiS" — Streptococcus pneumoniae, Haemophilus influenzae type b, Neisseria meningitidis, Salmonella, Klebsiella pneumoniae, Streptococcus agalactiae. These require opsonization for phagocytosis, explaining severe disease in asplenic patients.
The daily Mnemonic challenge reinforces these memory devices through active retrieval rather than passive review.
Practice Question Analysis Framework
For every microbiology question you get wrong:
1. Identify the knowledge gap: Was it organism recognition, mechanism understanding, or clinical correlation?
2. Review the mechanism: Why does this organism cause this clinical presentation?
3. Connect to similar organisms: What other bugs could cause similar presentations? How do you differentiate?
4. Test yourself again: Can you now recognize this pattern in a different clinical context?
Track your performance across organism categories using antibiotic resistance questions to identify weak areas that need targeted review.
Exam Day Strategy
Timing: 90 seconds per microbiology question
30 seconds: Read and identify key clinical clues
45 seconds: Connect clues to organism/mechanism
15 seconds: Eliminate wrong answers and confirm choice
Key Clinical Clues to Recognize Instantly
Travel history: Malaria (tropics), Histoplasma (Ohio Valley), Coccidioides (Southwest)
Occupational exposure: Anthrax (wool), Brucella (livestock), Tularemia (rabbits)
Immunocompromised status: Opportunistic infections have predictable patterns
Antibiotic history: Think C. difficile, resistant organisms
When You are Stuck
1. Use gram stain and morphology to narrow possibilities
2. Apply epidemiologic clues (age, location, exposures)
3. Consider the most common organism for that clinical syndrome
4. Eliminate options that dont fit the basic microbiology (gram stain, oxygen requirements)
Frequently Asked Questions
How long should I spend on microbiology during Step 1 prep?
Allocate 3-4 weeks for comprehensive microbiology review, with daily 2-3 hour study blocks. The subject is high-yield but manageable if you focus on mechanism-based learning rather than rote memorization.
Should I memorize every antibiotic coverage pattern?
No. Learn the major antibiotic classes and their mechanisms. Coverage patterns follow logically from mechanism understanding. For example, knowing that vancomycin cant cross gram-negative outer membranes explains why it only covers gram-positives.
How do I differentiate similar organisms like Strep species?
Focus on the 1-2 distinguishing features that Step 1 actually tests. For Strep: Group A (beta-hemolytic, bacitracin sensitive), Group B (beta-hemolytic, bacitracin resistant), Pneumoniae (alpha-hemolytic, optochin sensitive), Viridans (alpha-hemolytic, optochin resistant).
What if I cant remember all the viral oncogenes?
Focus on the highest-yield associations: EBV + Burkitt lymphoma, HPV 16/18 + cervical cancer, HTLV-1 + adult T-cell leukemia, HBV/HCV + hepatocellular carcinoma. Know the mechanism (viral integration, p53 inactivation) rather than memorizing every viral protein.
How important are fungi and parasites?
They represent 15-20% of microbiology questions. Dont ignore them, but focus on the most common organisms and their distinguishing features. Geographic distribution is particularly high-yield for fungi.
Should I use multiple question banks for microbiology?
Quality over quantity. Master one comprehensive question bank with detailed explanations rather than superficially covering multiple sources. Focus on understanding why wrong answers are wrong, not just getting questions correct.
Microbiology mastery for Step 1 comes from systematic pattern recognition, not textbook memorization. Build your foundation with high-yield organisms, understand the mechanisms that explain clinical presentations, and drill these connections until they become automatic. Your future patients — and your Step 1 score — will thank you.
Prepare smarter with Oncourse AI — adaptive MCQs, spaced repetition, and AI explanations built for USMLE Step 1. Download free on Android and iOS.