Enzymes — MCQs

On this page
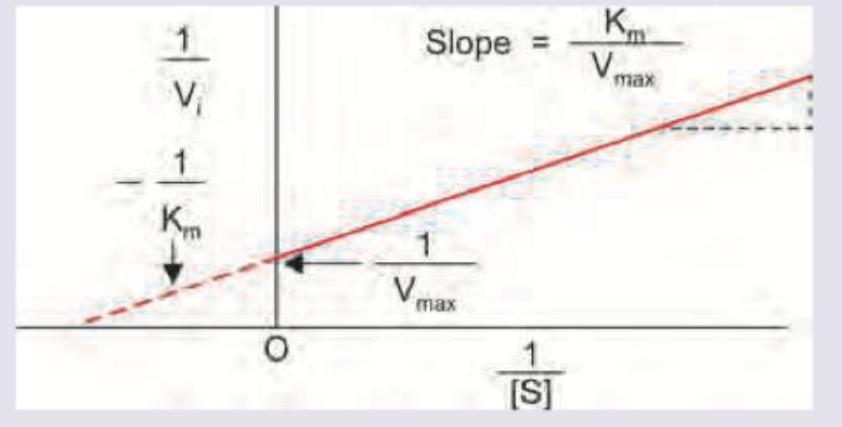
Name the plot dealing with kinetics of enzyme inhibition?
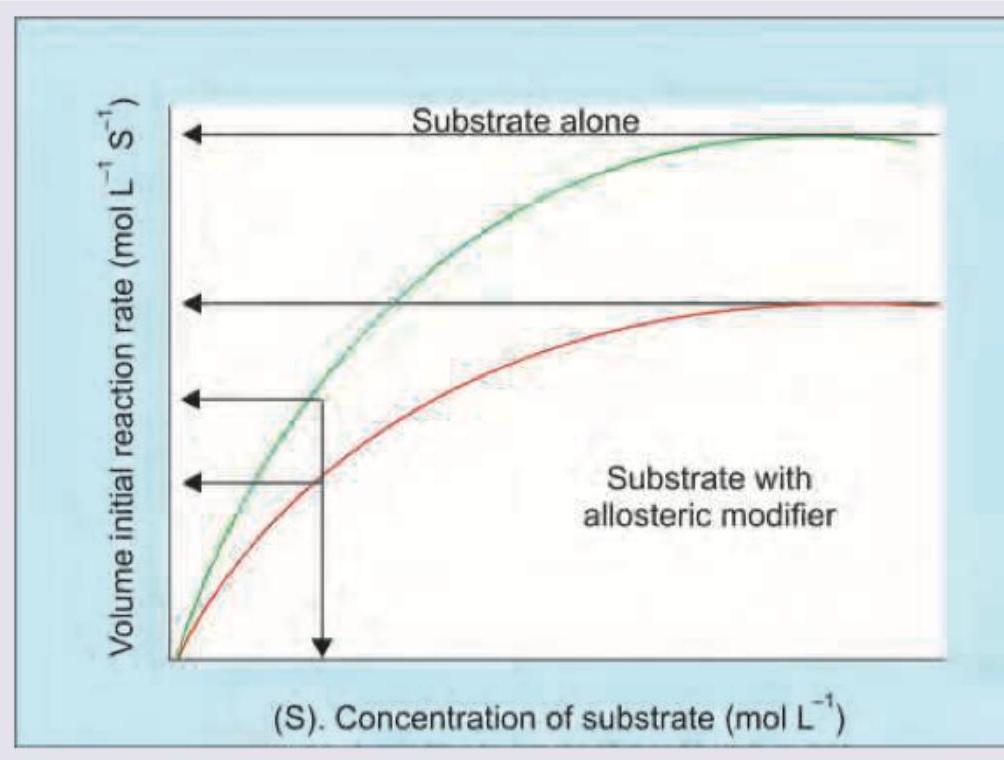
The curve below characterizes an allosteric enzyme system. Which of the following is correct about the given curve?
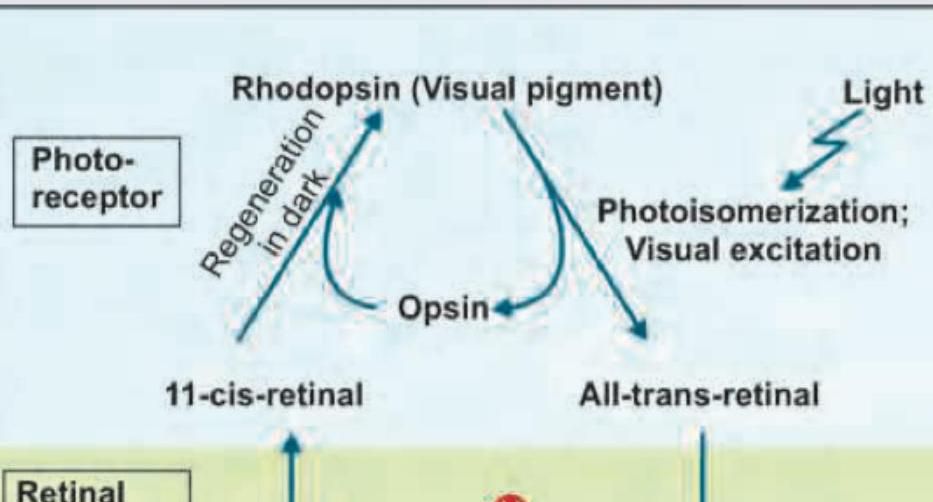
Name the enzyme involved in this cycle:
Identify the false statement regarding suicide inhibition
Fluoride, used in the collection of blood samples, inhibits which enzyme?
Zinc is cofactor of which enzyme?
Km increases, but Vmax remains same. This is which type of inhibition?
Enzymes, which play an important role in calcification, are:
Arsenic inhibits all except :
SGPT is found in:
Practice by Chapter
Enzyme Classification and Nomenclature
Practice Questions
Enzyme Kinetics and Michaelis-Menten Equation
Practice Questions
Enzyme Inhibition: Competitive and Non-competitive
Practice Questions
Allosteric Regulation
Practice Questions
Coenzymes and Cofactors
Practice Questions
Isoenzymes and Clinical Significance
Practice Questions
Enzyme Regulation: Covalent Modification
Practice Questions
Enzyme Regulation: Zymogen Activation
Practice Questions
Enzyme Induction and Repression
Practice Questions
Ribozymes and Catalytic RNA
Practice Questions
Enzyme Diagnostic Applications
Practice Questions
Enzyme Therapy and Inhibitors as Drugs
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
