Molecular Biology Techniques in Microbiology — MCQs

On this page
Two different specimens from two patients are received. Which of the following techniques can confirm they are from different individuals (in the context of Mycobacterium tuberculosis)?
What is the technique shown in the graph that demonstrates antibody-antigen interaction and response kinetics?
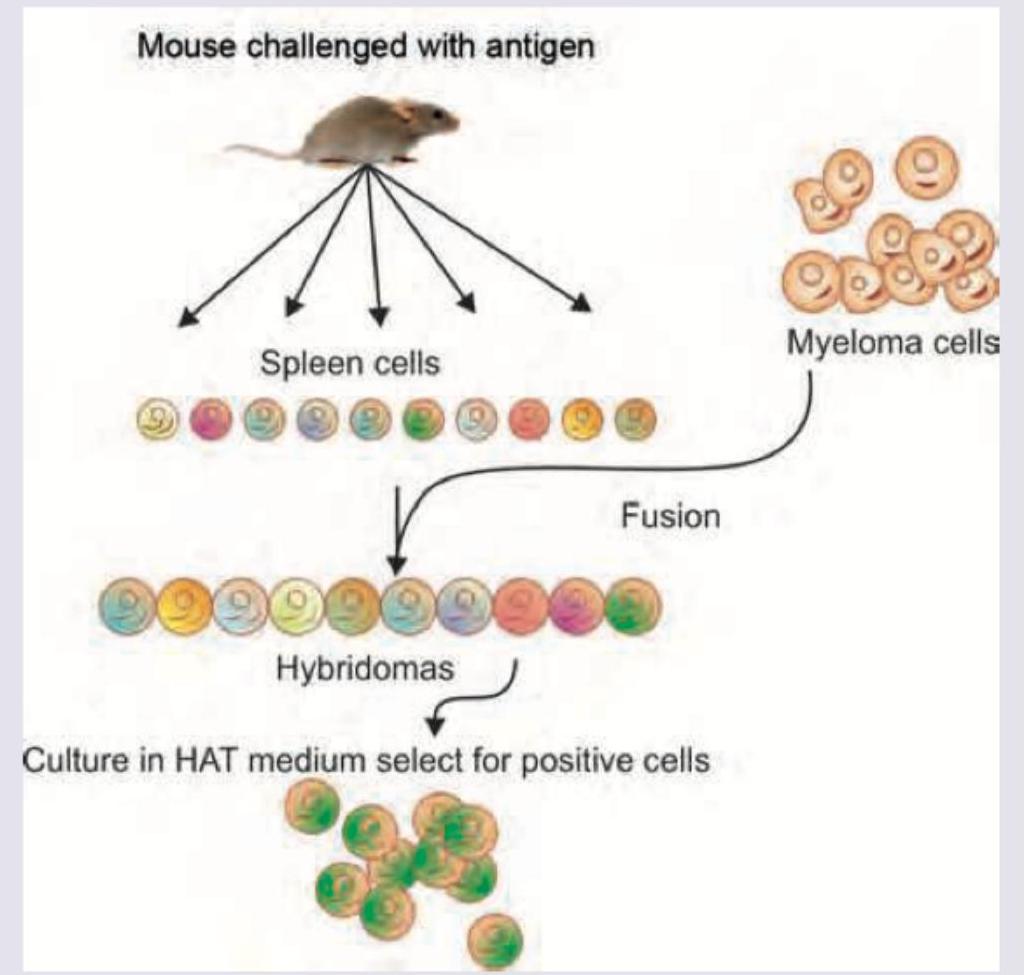
The technique shown in the picture is used for: (AIIMS Nov 2018)
Which virus can be identified by a PCR method and is endemic to India?
Which molecular biology technique is used to amplify a specific DNA sequence and is essential for diagnosing infections by detecting pathogen DNA in clinical samples?
A laboratory technician is preparing to perform PCR for the detection of a viral pathogen. The sample is suspected to contain RNA viruses. What modification should be made to the standard PCR protocol?
By which method foreign DNA is introduced into a cell by a virus or viral vector?
Which of the following is a primary cell line?
Practice by Chapter
Nucleic Acid Extraction Methods
Practice Questions
Polymerase Chain Reaction
Practice Questions
Real-Time PCR
Practice Questions
DNA Sequencing Techniques
Practice Questions
Next-Generation Sequencing in Microbiology
Practice Questions
Pulsed-Field Gel Electrophoresis
Practice Questions
Whole Genome Sequencing
Practice Questions
MALDI-TOF Mass Spectrometry
Practice Questions
Microarrays in Microbial Diagnostics
Practice Questions
Molecular Typing Methods
Practice Questions
Bioinformatics Tools in Microbiology
Practice Questions
Metagenomics
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
