Molecular Biology and Genomics — MCQs

On this page
Recombinant human insulin is made by -
DNA fingerprinting was discovered by:
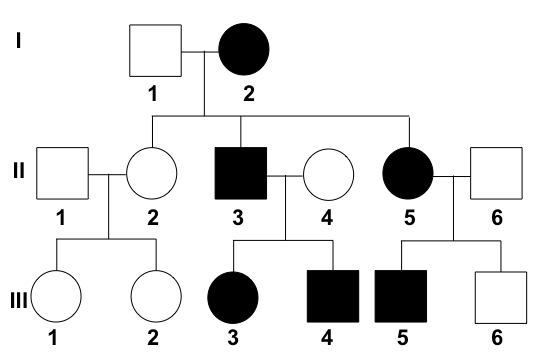
Examine this pedigree chart carefully. What type of transmission does it depict?
The term restriction map primarily refers to the mapping of sites of
In CRISPR-Cas9 system, which repair mechanism is predominantly used for genome editing?
Which of the following doesn't occur in 5' to 3' direction?
Assertion: DNA methylation leads to gene silencing. Reason: Methylation prevents binding of transcription factors to promoter regions.
Which type of mutation can act as a suppressor to restore the wild-type phenotype in organisms carrying a mutant gene?
What are the potential implications of using CRISPR-Cas9 technology to correct mutations in the CFTR gene for the treatment of cystic fibrosis?
How does a defect in the nucleotide excision repair (NER) pathway contribute to the development of xeroderma pigmentosum?
Practice by Chapter
DNA Replication and Repair Mechanisms
Practice Questions
Transcription Factors and Gene Regulation
Practice Questions
Epigenetics and DNA Methylation
Practice Questions
RNA Processing and Splicing
Practice Questions
miRNA and RNA Interference
Practice Questions
Protein Synthesis and Post-Translational Modifications
Practice Questions
Genomics and Human Genome Project
Practice Questions
Single Nucleotide Polymorphisms
Practice Questions
Gene Therapy Approaches
Practice Questions
CRISPR-Cas9 and Genome Editing
Practice Questions
DNA Fingerprinting and Forensics
Practice Questions
Molecular Basis of Genetic Diseases
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
