Molecular Biology and Genomics — MCQs

On this page
Termination of RNA molecule synthesis is signaled by a specific sequence in the DNA template strand, which is recognized by which termination protein?
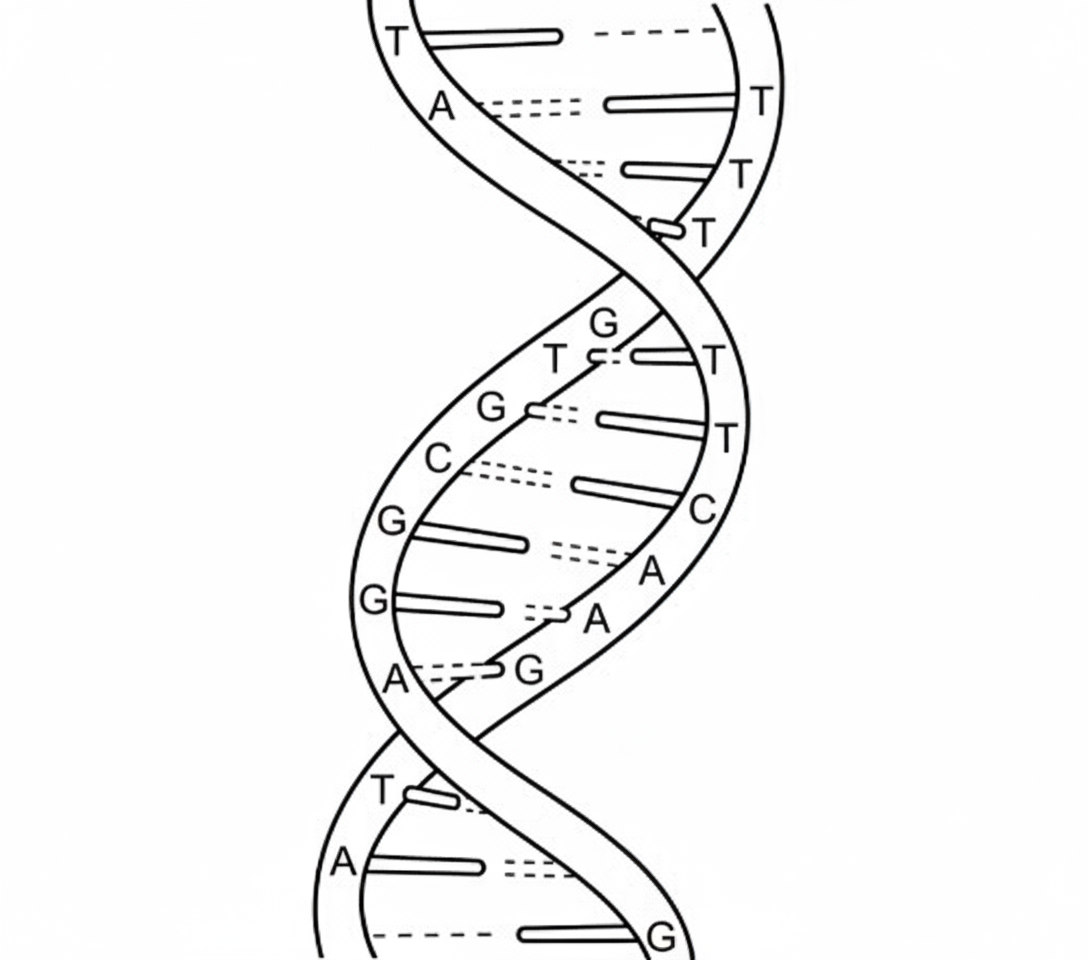
What is the bond between the strands in the given diagram?
Presence of which of the following in the expression vector ensures an increase in the yield of recombinant protein produced?
True about telomerase?
Which of the following statements is incorrect?
What is the function of reverse transcriptase?
During the translation process, which enzyme is responsible for the proofreading of mRNA?
In which direction does DNA replication and transcription occur?
Apolipoprotein B-48 and apolipoprotein B-100 are expressed as two different apoproteins due to a difference in which of the following?
The Shine-Dalgarno sequence in prokaryotes is primarily associated with which molecular process?
Practice by Chapter
DNA Replication and Repair Mechanisms
Practice Questions
Transcription Factors and Gene Regulation
Practice Questions
Epigenetics and DNA Methylation
Practice Questions
RNA Processing and Splicing
Practice Questions
miRNA and RNA Interference
Practice Questions
Protein Synthesis and Post-Translational Modifications
Practice Questions
Genomics and Human Genome Project
Practice Questions
Single Nucleotide Polymorphisms
Practice Questions
Gene Therapy Approaches
Practice Questions
CRISPR-Cas9 and Genome Editing
Practice Questions
DNA Fingerprinting and Forensics
Practice Questions
Molecular Basis of Genetic Diseases
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
