Molecular Biology and Genomics — MCQs

On this page
Which of the following is necessarily present in an expression vector but not in a cloning vector?
Acetylation of histones results in which of the following modification?
A patient presents with a condition highly suggestive of a defect in which of the following cellular mechanisms?
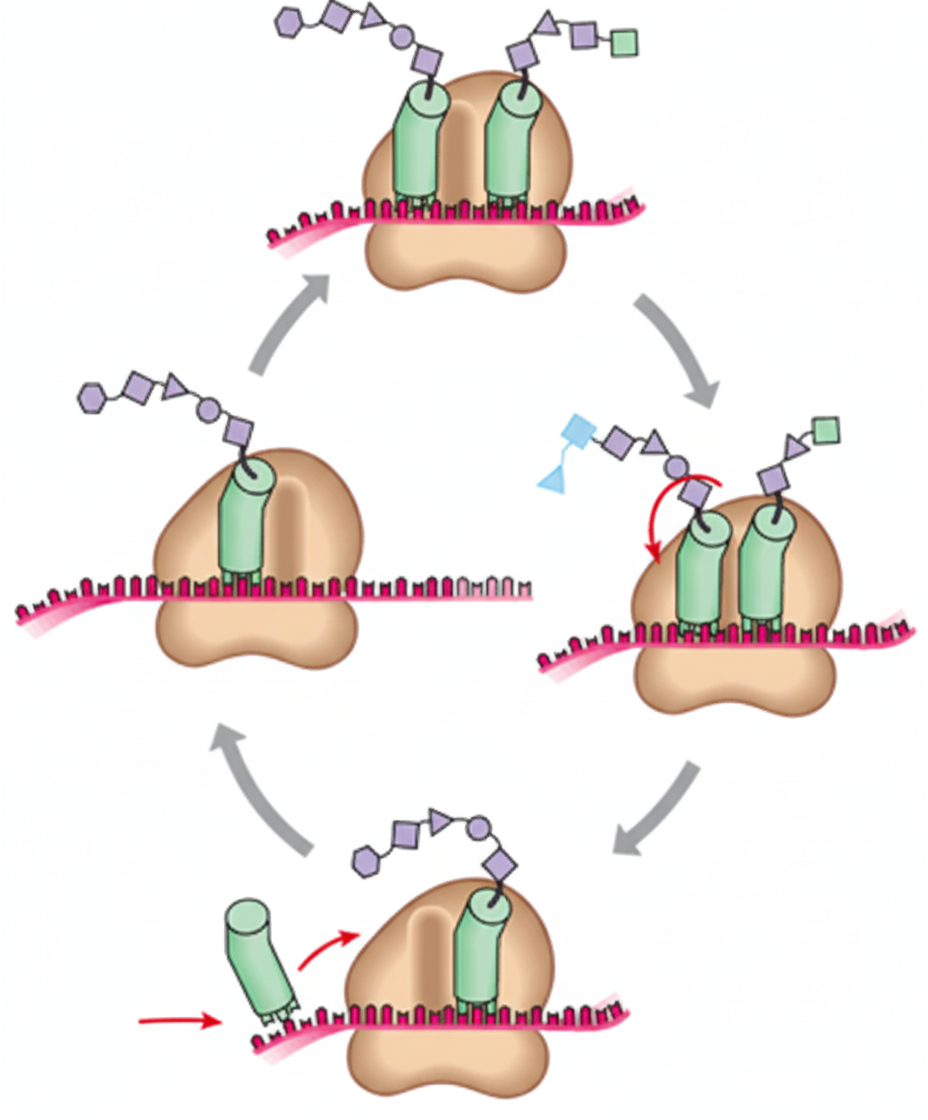
Which of the following statements is true about ribosomes?
Defect in snRNPs causes which of the following?
What property of DNA synthesis does the Sanger method of DNA sequencing take advantage of to generate a sequencing ladder?
Genomic imprinting is seen in which of the following conditions?
DNA replication follows which of the following models?
Which of the following represents the most characteristic function of a Type II Restriction Enzyme?
Which of the following statements regarding satellite DNA is incorrect?
Practice by Chapter
DNA Replication and Repair Mechanisms
Practice Questions
Transcription Factors and Gene Regulation
Practice Questions
Epigenetics and DNA Methylation
Practice Questions
RNA Processing and Splicing
Practice Questions
miRNA and RNA Interference
Practice Questions
Protein Synthesis and Post-Translational Modifications
Practice Questions
Genomics and Human Genome Project
Practice Questions
Single Nucleotide Polymorphisms
Practice Questions
Gene Therapy Approaches
Practice Questions
CRISPR-Cas9 and Genome Editing
Practice Questions
DNA Fingerprinting and Forensics
Practice Questions
Molecular Basis of Genetic Diseases
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
