Molecular Biology and Genomics — MCQs

On this page
Which type of mutation usually involves mutation of a gene coding for the following molecule?
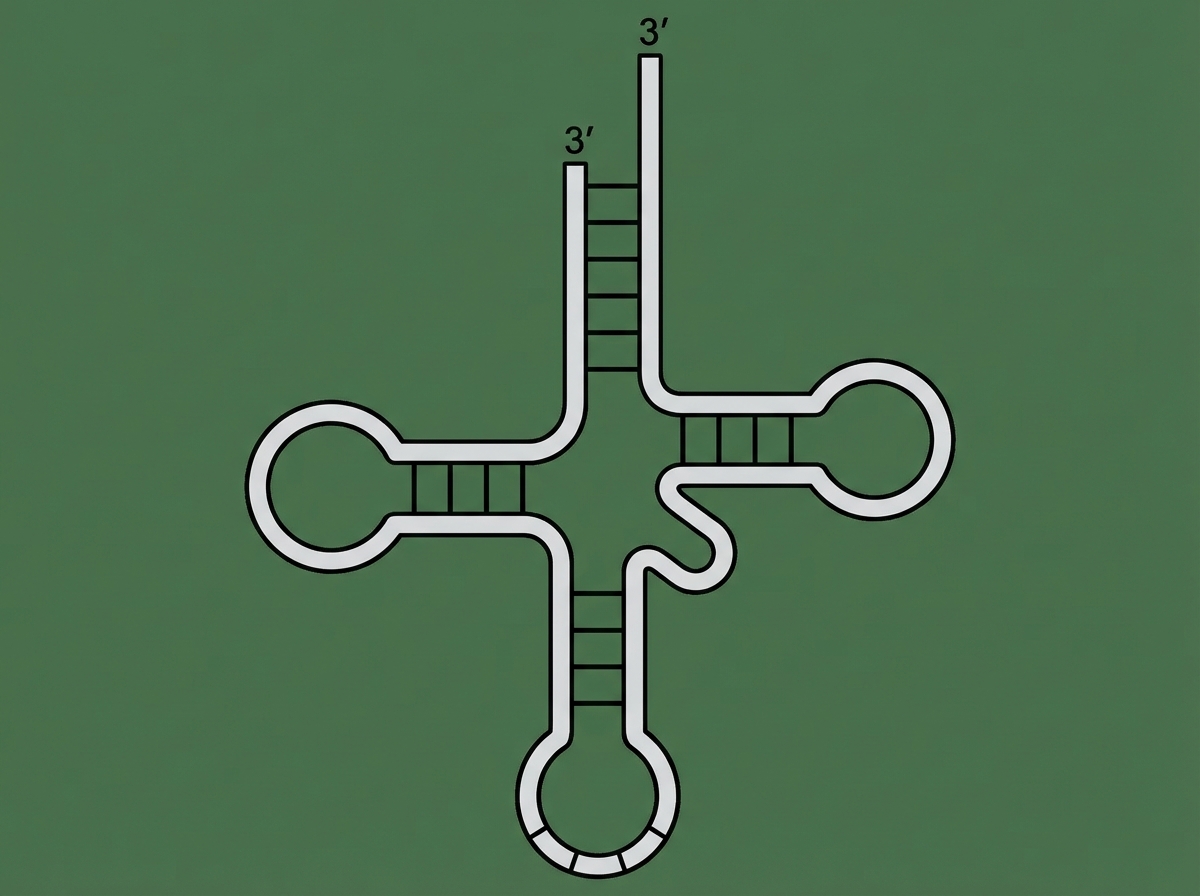
What is the most likely effect of a 2bp insertion in the middle of an intron in a typical eukaryotic gene?
Which of the following is the cofactor for prokaryotic DNA ligase?
Which of the following is NOT a characteristic of the genetic code?
Gene therapy is most effectively given in which of the following conditions?
Single gene defect causing multiple unrelated problems is known as:
What percentage of mitochondrial DNA constitutes total cellular DNA?
Okazaki fragments formed during DNA replication are structurally:
The DNA nucleotide sequence to which RNA polymerase binds to initiate transcription is called the:
In humans, what is the sequence of telomeres?
Practice by Chapter
DNA Replication and Repair Mechanisms
Practice Questions
Transcription Factors and Gene Regulation
Practice Questions
Epigenetics and DNA Methylation
Practice Questions
RNA Processing and Splicing
Practice Questions
miRNA and RNA Interference
Practice Questions
Protein Synthesis and Post-Translational Modifications
Practice Questions
Genomics and Human Genome Project
Practice Questions
Single Nucleotide Polymorphisms
Practice Questions
Gene Therapy Approaches
Practice Questions
CRISPR-Cas9 and Genome Editing
Practice Questions
DNA Fingerprinting and Forensics
Practice Questions
Molecular Basis of Genetic Diseases
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
