Molecular Biology and Genomics — MCQs

On this page
DNA denaturation is measured by absorbance at what wavelength?
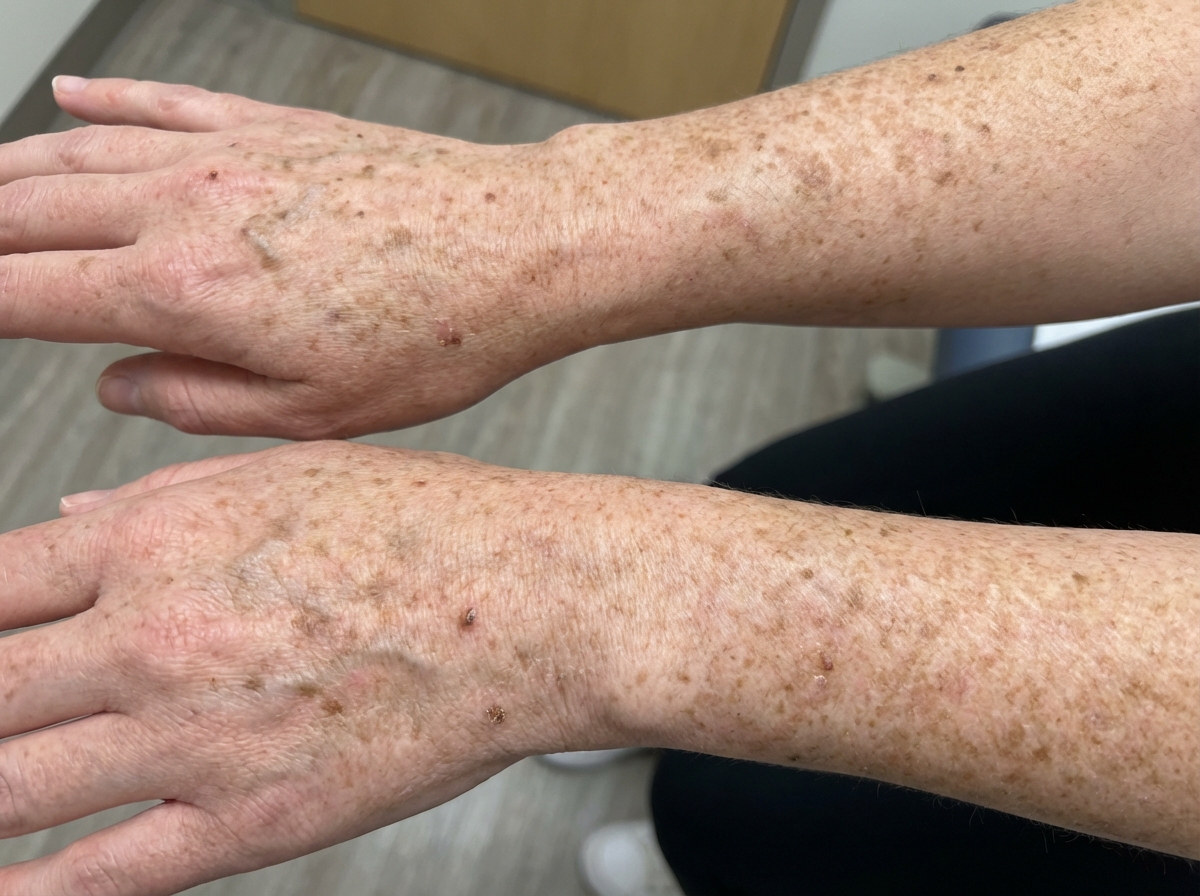
This patient presented with a defect. Which cellular process is most likely affected?
The Nobel prize in Physiology or Medicine for 2007 was jointly awarded to Mario R. Capecchi, Martin J. Evans, and Oliver Smithies for their discoveries of?
What are genes composed of?
Satellite sequences during G0 phase are seen in which of the following?
Which eukaryotic DNA polymerase is involved in proofreading and DNA repair during replication?
An 8-year-old boy presents with progressive proximal muscle weakness and difficulty rising from the floor. Examination reveals calf pseudohypertrophy and a positive Gower's sign. What is the most common type of mutation in the gene responsible for this condition?
Aminoacyl-tRNA is required for which of the following processes?
In the following partial sequence of mRNA, a mutation of the template DNA results in a change in codon 91 to UAA. What is the type of mutation?
Which technique is used to analyze DNA obtained from cancer biopsies when tumor cells are often contaminated with large numbers of admixed stromal cells?
Practice by Chapter
DNA Replication and Repair Mechanisms
Practice Questions
Transcription Factors and Gene Regulation
Practice Questions
Epigenetics and DNA Methylation
Practice Questions
RNA Processing and Splicing
Practice Questions
miRNA and RNA Interference
Practice Questions
Protein Synthesis and Post-Translational Modifications
Practice Questions
Genomics and Human Genome Project
Practice Questions
Single Nucleotide Polymorphisms
Practice Questions
Gene Therapy Approaches
Practice Questions
CRISPR-Cas9 and Genome Editing
Practice Questions
DNA Fingerprinting and Forensics
Practice Questions
Molecular Basis of Genetic Diseases
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
