Antimicrobial Resistance

On this page
🦠 AMR Fundamentals: The Microbial Arms Race
Antimicrobial resistance represents one of medicine's most urgent threats, transforming once-treatable infections into potential death sentences and eroding a century of therapeutic progress. You'll master how bacteria evolve sophisticated defense mechanisms, recognize resistance patterns before lab confirmation, interpret diagnostic tools that guide precision therapy, and deploy evidence-based treatment algorithms when first-line agents fail. This lesson equips you to combat resistance at the bedside and understand the global forces accelerating this crisis, building the clinical judgment essential for modern infectious disease management.
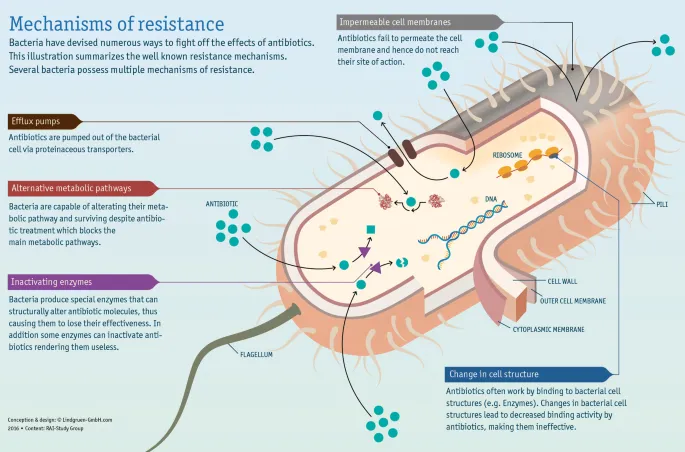
Antimicrobial resistance represents the evolutionary adaptation of microorganisms to survive exposure to antimicrobial agents. This phenomenon occurs when bacteria, viruses, fungi, or parasites develop mechanisms to neutralize drugs designed to kill or inhibit their growth. The World Health Organization estimates that resistant infections cause 700,000 deaths annually worldwide, with projections reaching 10 million deaths by 2050 without intervention.
📌 Remember: ESCAPE pathogens - Enterococcus faecium, Staphylococcus aureus, Clostridium difficile, Acinetobacter baumannii, Pseudomonas aeruginosa, Enterobacteriaceae. These six organisms account for >80% of healthcare-associated resistant infections globally.
The resistance spectrum encompasses multiple categories based on drug susceptibility patterns:
-
Multidrug-resistant (MDR): Non-susceptible to ≥1 agent in ≥3 antimicrobial categories
- Affects 15-20% of hospital isolates in Indian ICUs
- Associated with 2-3x increased mortality rates
- Treatment costs increase by $18,000-29,000 per patient
- Hospital stay extends by 6.4-12.7 days on average
-
Extensively drug-resistant (XDR): Non-susceptible to ≥1 agent in all but ≤2 categories
- Prevalence: 3-8% of MDR isolates progress to XDR
- Mortality rates: 40-60% for XDR infections
- Limited to 2-3 effective antibiotic classes
- Requires combination therapy in >90% of cases
-
Pandrug-resistant (PDR): Non-susceptible to all agents in all antimicrobial categories
- Represents <1% of clinical isolates but 100% mortality without novel agents
- Emergence documented in Acinetobacter, Pseudomonas, and Klebsiella species
| Resistance Category | Antibiotic Classes Affected | Prevalence in India | Mortality Rate | Treatment Options |
|---|---|---|---|---|
| Susceptible | None | 60-70% isolates | <5% | Standard therapy |
| MDR | ≥3 classes | 15-25% isolates | 15-25% | Limited options |
| XDR | All but ≤2 classes | 3-8% isolates | 40-60% | Salvage therapy |
| PDR | All classes | <1% isolates | >90% | Experimental drugs |
| Colistin-resistant | Last-resort agents | 2-5% isolates | 70-80% | No effective therapy |
The resistance burden varies significantly across geographic regions and healthcare settings. Indian surveillance data reveals carbapenem resistance rates of 15-30% in Enterobacteriaceae, 40-60% in Acinetobacter species, and 20-35% in Pseudomonas aeruginosa. These rates exceed global averages by 2-3 fold, reflecting intensive antibiotic use and infection control challenges.
💡 Master This: Resistance development requires three critical elements - genetic determinant (resistance gene), selective pressure (antibiotic exposure), and transmission opportunity (healthcare contact). Eliminating any component breaks the resistance cycle and prevents emergence.
Understanding resistance classification enables rational antibiotic selection and guides empirical therapy decisions. Connect these fundamental patterns through resistance mechanisms to predict which antibiotics remain effective against specific pathogens.
🦠 AMR Fundamentals: The Microbial Arms Race
⚙️ Resistance Mechanisms: The Bacterial Defense Arsenal
Enzymatic inactivation represents the most common resistance mechanism, accounting for 60-70% of clinically significant resistance. Bacteria produce enzymes that chemically modify or destroy antimicrobial agents before they reach their targets.
- Beta-lactamases: Hydrolyze beta-lactam ring in penicillins, cephalosporins, carbapenems
- >1,000 variants identified with distinct substrate specificities
- ESBLs (Extended-Spectrum Beta-Lactamases) affect 15-25% of E. coli isolates
- Confer resistance to 3rd/4th generation cephalosporins
- Remain susceptible to carbapenems and beta-lactamase inhibitors
- Carbapenemases hydrolyze all beta-lactams including carbapenems
- KPC, NDM, OXA-48 families predominate globally
- Associated with 40-70% mortality in bloodstream infections
📌 Remember: SPACE organisms produce AmpC beta-lactamases - Serratia, Pseudomonas, Acinetobacter, Citrobacter, Enterobacter. These chromosomal enzymes are inducible by beta-lactam exposure and confer resistance to penicillins, cephalosporins, and aztreonam.
Target modification alters the antimicrobial binding site, reducing drug affinity while preserving essential cellular functions. This mechanism affects 20-30% of resistant isolates and often involves point mutations in target genes.

-
Penicillin-binding protein (PBP) alterations:
- PBP2a in MRSA reduces methicillin affinity by >1000-fold
- mecA gene encodes modified PBP with low beta-lactam affinity
- Confers resistance to all beta-lactams except ceftaroline
- Requires vancomycin or linezolid for treatment
-
DNA gyrase mutations in fluoroquinolone resistance:
- gyrA and parC gene mutations reduce quinolone binding
- Single mutation increases MIC by 4-8 fold
- Double mutations confer high-level resistance (MIC >32 mg/L)
- Cross-resistance occurs across all fluoroquinolones
| Resistance Mechanism | Target Antibiotic | Genetic Basis | Resistance Level | Clinical Impact |
|---|---|---|---|---|
| Beta-lactamase | Beta-lactams | bla genes | High (MIC >256) | Severe - Limited options |
| PBP modification | Methicillin | mecA/mecC | Complete | Critical - MRSA emergence |
| Gyrase mutation | Fluoroquinolones | gyrA/parC | Moderate-High | Significant - Cross-resistance |
| rRNA methylation | Aminoglycosides | rmtA-H | High (MIC >512) | Severe - Synergy loss |
| Efflux overexpression | Multiple classes | Pump regulators | Moderate | Broad - MDR phenotype |
Permeability reduction decreases antibiotic penetration into bacterial cells through porin loss or membrane modifications. This mechanism contributes to 10-15% of resistance cases and often combines with other mechanisms.
- Porin deficiency in Enterobacteriaceae:
- OmpC/OmpF loss reduces carbapenem penetration by >90%
- Combined with ESBL production creates carbapenem resistance
- Ertapenem most affected due to large molecular size
- Meropenem/Imipenem retain partial activity
💡 Master This: Efflux pump overexpression creates multidrug resistance by actively removing antibiotics from bacterial cells. AcrAB-TolC system in Enterobacteriaceae exports fluoroquinolones, chloramphenicol, tetracyclines, and beta-lactams simultaneously, explaining MDR phenotypes in clinical isolates.
Understanding these molecular mechanisms enables prediction of cross-resistance patterns and guides combination therapy selection. Connect resistance mechanisms through detection methods to identify optimal diagnostic approaches for clinical practice.
⚙️ Resistance Mechanisms: The Bacterial Defense Arsenal
🎯 Clinical Recognition: Resistance Pattern Mastery
ESBL Detection Patterns represent the most clinically significant resistance recognition challenge, affecting 15-30% of Enterobacteriaceae isolates in Indian hospitals. The characteristic pattern reveals cephalosporin resistance with carbapenem susceptibility.
- Classic ESBL Pattern Recognition:
- Resistant: Ceftriaxone, ceftazidime, cefotaxime, aztreonam
- Susceptible: Carbapenems, cephamycins (cefoxitin), tigecycline
- Ceftazidime MIC >2 mg/L suggests ESBL production
- Cefotaxime MIC >1 mg/L confirms extended-spectrum activity
- Variable: Fluoroquinolones (60-80% co-resistance), aminoglycosides (40-70% co-resistance)
- Trimethoprim-sulfamethoxazole resistance in >90% of ESBL producers
- Piperacillin-tazobactam may retain activity (30-50% susceptible)
📌 Remember: ESBL Confirmation - Ceftazidime vs Ceftazidime-clavulanate, Cefotaxime vs Cefotaxime-clavulanate. ≥3 doubling dilution decrease (≥8-fold MIC reduction) with inhibitor confirms ESBL production per CLSI guidelines.
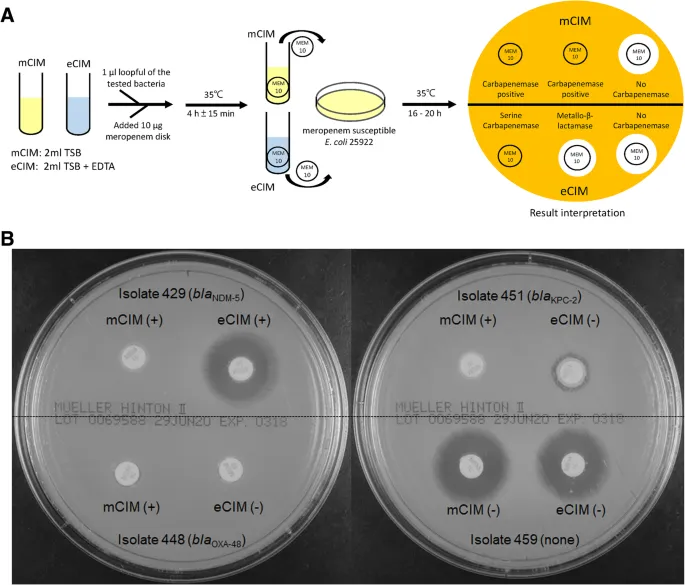
Carbapenemase Recognition requires distinguishing true carbapenemase production from carbapenem resistance due to porin loss plus ESBL. This distinction determines infection control measures and treatment strategies.
- Carbapenemase Pattern Signatures:
- Class A (KPC): Ertapenem MIC >8 mg/L, meropenem MIC 4-16 mg/L
- Boronic acid inhibition suggests KPC production
- Ceftazidime-avibactam remains active against >95% of KPC producers
- Class B (NDM, VIM, IMP): All carbapenems MIC >8 mg/L
- EDTA inhibition confirms metallo-beta-lactamase
- Aztreonam retains activity (not hydrolyzed by MBLs)
- Class D (OXA-48): Ertapenem MIC >8 mg/L, meropenem MIC 2-8 mg/L
- Temocillin often remains susceptible
- Colistin required for severe infections
- Class A (KPC): Ertapenem MIC >8 mg/L, meropenem MIC 4-16 mg/L
AmpC Beta-lactamase Recognition involves identifying inducible versus constitutive expression patterns that affect therapeutic selection and resistance development risk.
| Organism Pattern | Cefoxitin | Ceftazidime | Cefepime | Carbapenem | Interpretation |
|---|---|---|---|---|---|
| Wild-type | S | S | S | S | No resistance |
| ESBL | S | R | R | S | ESBL producer |
| AmpC | R | R | S | S | AmpC producer |
| Carbapenemase | R | R | R | R | Carbapenemase |
| Porin + ESBL | R | R | R | I/R | Combined mechanism |
MRSA Pattern Recognition extends beyond methicillin resistance to include associated resistance markers that guide empirical therapy and infection control decisions.
- MRSA Resistance Associations:
- Healthcare-associated MRSA: Fluoroquinolone resistance >90%, clindamycin resistance >80%
- Vancomycin MIC creep: >50% isolates with MIC 1.5-2 mg/L
- Linezolid and daptomycin remain >95% susceptible
- Community-associated MRSA: Clindamycin susceptible >70%, TMP-SMX susceptible >85%
- PVL toxin positive in >80% of CA-MRSA strains
- Doxycycline effective for skin/soft tissue infections
- Healthcare-associated MRSA: Fluoroquinolone resistance >90%, clindamycin resistance >80%
💡 Master This: D-test positivity indicates inducible clindamycin resistance despite apparent susceptibility. Erythromycin-resistant, clindamycin-susceptible isolates with D-zone around clindamycin disk will develop resistance during therapy in >60% of cases.
Recognition of these resistance patterns enables rapid therapeutic optimization and appropriate infection control measures. Connect pattern recognition through systematic diagnostic approaches to master resistance detection methodologies.
🎯 Clinical Recognition: Resistance Pattern Mastery
🔬 Diagnostic Precision: Resistance Detection Mastery

Disk Diffusion Method (Kirby-Bauer) provides qualitative results through zone diameter interpretation, offering cost-effective screening for routine susceptibility testing. This method requires standardized conditions and careful quality control for reliable results.
- Critical Technical Parameters:
- Mueller-Hinton agar pH 7.2-7.4, divalent cation concentration 20-25 mg/L Mg²⁺, 10-12.5 mg/L Ca²⁺
- Inoculum density: 0.5 McFarland standard (1.5 × 10⁸ CFU/mL)
- Disk application within 15 minutes of inoculum preparation
- Incubation: 35°C ± 2°C for 16-20 hours in ambient air
- Zone measurement: Nearest whole millimeter using transmitted light
- Swarming Proteus: Measure thin film at swarm edge
- Double zones: Measure inner zone for trimethoprim-sulfamethoxazole
📌 Remember: Quality Control Strains - Escherichia coli ATCC 25922, Staphylococcus aureus ATCC 25923, Pseudomonas aeruginosa ATCC 27853. These ESP strains must produce zone diameters within CLSI acceptable ranges for each antibiotic tested.
Broth Microdilution represents the gold standard for quantitative susceptibility testing, providing precise MIC values essential for dosing optimization and resistance mechanism detection.
- MIC Interpretation Categories:
- Susceptible (S): Standard dosing regimens achieve therapeutic concentrations exceeding MIC
- Clinical success rates >90% with appropriate dosing
- PK/PD targets: fT>MIC ≥40% for beta-lactams, AUC/MIC >125 for fluoroquinolones
- Susceptible-Dose Dependent (SDD): Higher doses or alternative schedules required
- Extended infusion beta-lactams or increased frequency
- Success rates 70-85% with optimized dosing
- Intermediate (I): Uncertain clinical efficacy with standard dosing
- Buffer zone accounting for technical variation
- Alternative agents preferred when available
- Resistant (R): Clinical failure likely despite maximal dosing
- Success rates <60% even with combination therapy
- Alternative antimicrobials required
- Susceptible (S): Standard dosing regimens achieve therapeutic concentrations exceeding MIC
Specialized Phenotypic Tests target specific resistance mechanisms requiring enhanced detection beyond routine susceptibility testing.
| Test Method | Target Mechanism | Principle | Interpretation | Clinical Utility |
|---|---|---|---|---|
| ESBL Confirmation | Extended-spectrum beta-lactamases | Clavulanate synergy | ≥3 dilution decrease | Carbapenem selection |
| AmpC Detection | AmpC beta-lactamases | Boronic acid inhibition | ≥3 dilution decrease | Cefepime preference |
| Carbapenemase Detection | Carbapenemases | Multiple inhibitors | Mechanism-specific | Infection control |
| D-test | Inducible clindamycin resistance | Erythromycin induction | Flattened zone | Clindamycin avoidance |
| Vancomycin Screen | Vancomycin resistance | High-concentration agar | Any growth | VRE detection |
Genotypic Detection Methods identify resistance genes directly, providing rapid results and mechanism identification for targeted therapy and epidemiological tracking.
-
PCR-based Detection:
- Real-time PCR: Results in 2-4 hours from positive blood culture
- mecA/mecC detection for MRSA identification
- vanA/vanB genes for VRE screening
- Multiplex PCR panels: Simultaneous detection of multiple genes
- Carbapenemase genes: blaKPC, blaNDM, blaOXA-48, blaVIM, blaIMP
- ESBL genes: blaCTX-M, blaSHV, blaTEM variants
- Real-time PCR: Results in 2-4 hours from positive blood culture
-
Whole Genome Sequencing (WGS):
- Comprehensive resistance profiling within 24-48 hours
- Outbreak investigation through phylogenetic analysis
- Single nucleotide polymorphism tracking for transmission chains
- Resistance gene evolution and horizontal transfer detection
💡 Master This: Phenotype-genotype discordance occurs in 5-15% of cases due to gene expression regulation, novel variants, or multiple mechanisms. Phenotypic testing remains essential for clinical decision-making, while genotypic methods provide rapid screening and epidemiological data.
Mastering resistance detection methodologies enables optimal test selection and accurate result interpretation. Connect diagnostic precision through treatment algorithms to develop evidence-based therapeutic strategies.
🔬 Diagnostic Precision: Resistance Detection Mastery
⚖️ Treatment Algorithms: Evidence-Based Resistance Management
ESBL-Producing Enterobacteriaceae Management requires carbapenem-based therapy for serious infections, with alternative agents reserved for specific clinical scenarios based on susceptibility patterns and infection severity.
- Carbapenem Selection Criteria:
- Meropenem: Pseudomonas coverage required, CNS infections, severe sepsis
- Dosing: 1-2g every 8 hours, extended infusion (3-4 hours) for MIC >2 mg/L
- PK/PD target: fT>MIC ≥40% for bacteriostatic, ≥70% for bactericidal effect
- Ertapenem: No Pseudomonas risk, outpatient therapy, once-daily dosing
- Dosing: 1g every 24 hours, renal adjustment for CrCl <30 mL/min
- Tissue penetration: Excellent for intra-abdominal, skin/soft tissue infections
- Imipenem-cilastatin: Alternative when meropenem unavailable
- CNS toxicity risk with high doses or renal impairment
- Dosing: 500mg-1g every 6-8 hours based on infection severity
- Meropenem: Pseudomonas coverage required, CNS infections, severe sepsis
📌 Remember: Carbapenem-Sparing Strategies for ESBL infections - Piperacillin-tazobactam for UTI/mild infections, Tigecycline for intra-abdominal, Fosfomycin for UTI, Nitrofurantoin for cystitis. PTFN alternatives reduce carbapenem pressure and preserve efficacy.
Carbapenemase-Producing Organism (CPO) Treatment represents the most challenging resistance scenario, requiring combination therapy and novel agents for optimal outcomes.
| CPO Type | First-Line Therapy | Alternative Options | Combination Partners | Success Rate |
|---|---|---|---|---|
| KPC | Ceftazidime-avibactam | Meropenem-vaborbactam | Polymyxin B/E | 75-85% |
| NDM | Aztreonam + ceftazidime-avibactam | Cefiderocol | Tigecycline | 65-75% |
| OXA-48 | Ceftazidime-avibactam | Meropenem + polymyxin | Fosfomycin | 70-80% |
| VIM/IMP | Cefiderocol | Aztreonam + avibactam | Polymyxin B | 60-70% |
| Mixed | Cefiderocol | Combination therapy | Multiple agents | 50-65% |
- Ceftazidime-avibactam: Active against Class A (KPC) and Class C (AmpC) beta-lactamases
- Dosing: 2.5g every 8 hours, extended infusion recommended
- Renal adjustment: Dose reduction for CrCl <50 mL/min
- Meropenem-vaborbactam: KPC-specific activity, enhanced against porin-deficient strains
- Dosing: 4g every 8 hours as 3-hour infusion
- Nephrotoxicity: Lower risk compared to polymyxins
- Cefiderocol: Siderophore cephalosporin with broad activity against all carbapenemases
- Iron-dependent uptake bypasses porin resistance
- Dosing: 2g every 8 hours as 3-hour infusion
MRSA Treatment Algorithms incorporate vancomycin MIC values, infection type, and patient factors to optimize therapeutic selection and dosing strategies.
⭐ Clinical Pearl: Vancomycin AUC/MIC ratio >400 correlates with clinical success and reduced resistance development. Bayesian dosing software enables individualized dosing to achieve target AUC 400-600 mg·h/L while minimizing nephrotoxicity risk.
Combination Therapy Principles guide synergistic drug selection for difficult-to-treat resistant infections, particularly CPO and XDR pathogens.
- Synergy Testing Indications:
- Pandrug-resistant organisms with no susceptible agents
- High-risk infections: Bloodstream, CNS, endocarditis
- Time-kill studies demonstrate >2 log₁₀ reduction vs monotherapy
- Checkerboard assays identify synergistic combinations (FIC ≤0.5)
- Combination strategies:
- Beta-lactam + polymyxin: Membrane disruption enhances drug penetration
- Double carbapenem: Meropenem + ertapenem for KPC producers
- Aztreonam + avibactam: MBL-active combination under investigation
💡 Master This: Heteroresistance affects treatment outcomes in 15-25% of apparently susceptible isolates. Population analysis profiling identifies resistant subpopulations that emerge during monotherapy, supporting combination approaches for critical infections.
Evidence-based treatment algorithms enable systematic resistance management with improved clinical outcomes. Connect therapeutic strategies through stewardship principles to develop comprehensive resistance prevention programs.
⚖️ Treatment Algorithms: Evidence-Based Resistance Management
🌐 Global Impact: The Resistance Pandemic Architecture
Global Resistance Burden demonstrates exponential growth across all antimicrobial classes, with developing countries experiencing disproportionate impact due to healthcare infrastructure limitations and unregulated antibiotic access.
- Economic Impact Projections:
- Current annual costs: $35 billion in healthcare expenses, $20 billion in productivity losses
- 2050 projections: $100 trillion cumulative GDP impact without intervention
- 10 million deaths annually by 2050 (exceeding cancer mortality)
- Healthcare costs increase by $1.2 trillion globally
- Low-middle income countries: 75% of resistance burden with <25% of surveillance capacity
- Infant mortality increases by 214,000 deaths annually
- Surgical procedures become prohibitively risky without effective prophylaxis
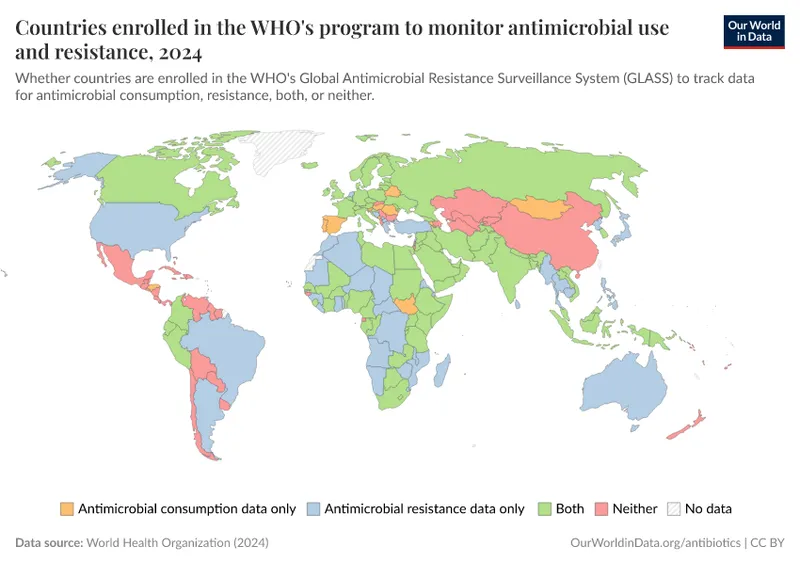
📌 Remember: GLASS (Global Antimicrobial Resistance and Use Surveillance System) - Global Laboratory Antimicrobial Surveillance System. WHO initiative tracking resistance patterns in >100 countries using standardized methodologies for international comparison.
One Health Resistance Dynamics reveal complex interactions between human, animal, and environmental antimicrobial use that drive resistance selection and transmission across ecological boundaries.
- Agricultural Antimicrobial Use:
- Global consumption: 131,000 tons annually in food animals (4x human use)
- Growth promotion: 60-80% of agricultural antibiotics used for non-therapeutic purposes
- Colistin use in agriculture drives mcr-1 gene emergence
- Fluoroquinolone use selects for ESBL-producing E. coli in food chains
- Resistance transmission: 25-40% of human ESBL infections linked to food animal sources
- ST131 E. coli clone spreads between poultry and human populations
- Enterococcus resistance transfers from agricultural to clinical settings
Environmental Resistance Reservoirs serve as mixing vessels for resistance gene exchange and long-term persistence of antimicrobial compounds that maintain selective pressure.
| Environmental Source | Resistance Contribution | Key Pathogens | Intervention Strategies |
|---|---|---|---|
| Hospital Wastewater | High-level resistance genes | CPO, VRE, MRSA | Advanced treatment systems |
| Agricultural Runoff | Volume-driven selection | ESBL E. coli, Enterococcus | Manure management |
| Pharmaceutical Manufacturing | Extreme concentrations | Multi-class resistance | Discharge regulations |
| Aquaculture Systems | Closed-system amplification | Vibrio, Aeromonas | Reduced antimicrobial use |
| Urban Water Systems | Community-wide exposure | Opportunistic pathogens | Water treatment upgrades |
Resistance Transmission Networks operate through healthcare systems, international travel, and global trade that rapidly disseminate resistant clones across continental boundaries.
-
Healthcare-Associated Transmission:
- Hand hygiene compliance: <50% in many settings enables cross-transmission
- Contact precautions: Reduce transmission by >80% when properly implemented
- Single-room isolation for CPO and VRE carriers
- Dedicated equipment and staff cohorting during outbreaks
- Screening programs: Active surveillance identifies asymptomatic carriers
- Admission screening detects 5-15% CPO carriage in high-risk populations
- Contact tracing prevents secondary transmission in >90% of identified contacts
-
International Spread Patterns:
- Medical tourism: 15-20% of travelers acquire resistant organisms during healthcare abroad
- Refugee populations: Conflict zones serve as resistance amplification centers
- Syrian conflict: OXA-48 and NDM spread throughout European healthcare systems
- Rohingya crisis: Extensively resistant TB strains cross international borders
💡 Master This: Resistance gene mobility occurs through plasmids, transposons, and integrons that transfer multiple resistance determinants simultaneously. IncX3 plasmids carrying blaNDM genes spread globally within 5 years of initial detection, demonstrating rapid resistance dissemination potential.
Understanding global resistance architecture enables coordinated intervention strategies that address systemic drivers while preserving antimicrobial effectiveness. Connect international perspectives through stewardship frameworks to develop sustainable resistance management approaches.
🌐 Global Impact: The Resistance Pandemic Architecture
🎯 Resistance Mastery: Clinical Command Arsenal
Essential Resistance Recognition Matrix provides rapid pattern identification for common resistance phenotypes encountered in clinical practice.
| Clinical Scenario | Resistance Pattern | First-Line Therapy | Alternative Options | Key Monitoring |
|---|---|---|---|---|
| UTI + ESBL | Ceph-R, Carb-S | Ertapenem 1g q24h | Fosfomycin, Nitrofurantoin | Clinical response 48-72h |
| Pneumonia + MRSA | Meth-R, Van-S | Vancomycin AUC-guided | Linezolid, Ceftaroline | Trough levels, Nephrotoxicity |
| Sepsis + CPO | Carb-R, Pol-S | Ceftazidime-avibactam | Polymyxin combinations | Resistance emergence |
| Endocarditis + VRE | Van-R, Amp-R | Daptomycin + gentamicin | Linezolid + rifampin | Bactericidal activity |
| Meningitis + PRSP | Pen-R, Ceph-I | Vancomycin + ceftriaxone | Meropenem monotherapy | CSF penetration |
Pharmacokinetic Optimization Strategies maximize antimicrobial effectiveness against resistant pathogens through dosing modifications and administration techniques.
- Extended Infusion Protocols:
- Beta-lactams: 3-4 hour infusions increase fT>MIC from 40% to >70%
- Meropenem: 2g over 3 hours every 8 hours for MIC 4-8 mg/L
- Piperacillin-tazobactam: 4.5g over 4 hours every 8 hours
- Vancomycin: AUC-guided dosing targets 400-600 mg·h/L
- Loading dose: 25-30 mg/kg for severe infections
- Maintenance: 15-20 mg/kg every 8-12 hours based on renal function
- Beta-lactams: 3-4 hour infusions increase fT>MIC from 40% to >70%
Combination Therapy Decision Matrix guides synergistic drug selection for difficult-to-treat resistant infections.
- Synergy Prediction Rules:
- Beta-lactam + polymyxin: Membrane disruption enhances beta-lactam penetration
- Meropenem + polymyxin B for NDM-producing Enterobacteriaceae
- Ceftazidime + colistin for XDR Pseudomonas
- Double beta-lactam: Enzyme saturation strategy for carbapenemase producers
- Meropenem + aztreonam for MBL producers
- Ceftazidime-avibactam + aztreonam for NDM + OXA-48
- Aminoglycoside synergy: Cell wall disruption enhances uptake
- Ampicillin + gentamicin for Enterococcus endocarditis
- Vancomycin + gentamicin for MRSA endocarditis
- Beta-lactam + polymyxin: Membrane disruption enhances beta-lactam penetration
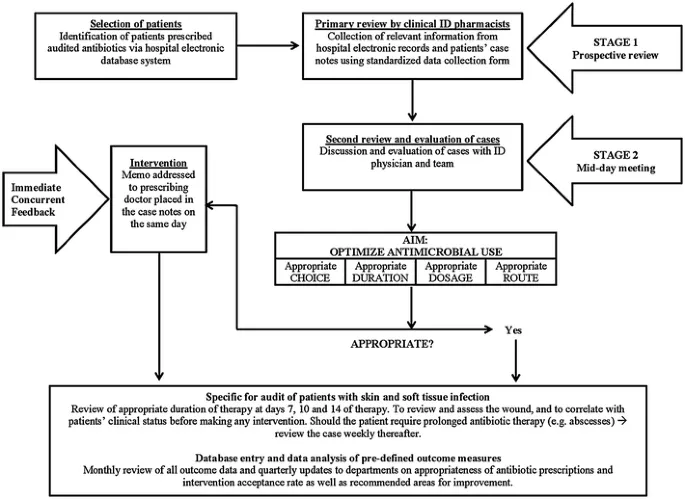
Stewardship Integration Points embed resistance management into routine clinical workflows for sustainable practice improvement.
- Prescribing Decision Support:
- Electronic alerts for broad-spectrum antibiotic orders
- Automatic stop dates for empirical therapy (default 72 hours)
- De-escalation prompts when culture results available
- Dose optimization recommendations based on renal function
- Restriction protocols for last-resort agents
- Infectious disease consultation required for polymyxins
- Prior authorization for novel beta-lactam combinations
⭐ Clinical Pearl: Rapid diagnostic testing reduces time to optimal therapy by 24-48 hours, improving clinical outcomes and reducing resistance selection. Blood culture identification and susceptibility testing within 6-8 hours enables early de-escalation in >70% of cases.
Resistance Prevention Protocols integrate infection control measures with antimicrobial optimization to prevent resistance emergence and transmission.
| Prevention Strategy | Implementation | Effectiveness | Monitoring Metrics |
|---|---|---|---|
| Contact Precautions | CPO, VRE, MRSA isolation | >80% transmission reduction | Acquisition rates |
| Hand Hygiene | 5 moments compliance monitoring | >70% infection prevention | Compliance rates |
| Environmental Cleaning | Enhanced disinfection protocols | >60% contamination reduction | ATP monitoring |
| Antimicrobial Cycling | Class rotation every 3-6 months | Variable resistance impact | Susceptibility trends |
| Selective Decontamination | Topical antimicrobials for high-risk patients | >50% infection reduction | Resistance emergence |
This clinical command arsenal provides immediate access to evidence-based tools for resistance management. Regular practice with these frameworks develops pattern recognition and decision-making speed essential for optimal patient care in the resistance era.
🎯 Resistance Mastery: Clinical Command Arsenal
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app
























