Next-Generation Sequencing in Microbiology — MCQs

Which one of the following is an emerging viral disease?
What is the best investigation for identifying malaria species?
Which of the following techniques is primarily used for RNA analysis?
Best method for the detection of mutations with low allele frequency is:
All of the following are pattern recognition receptors for extracellular or ingested microbes, except:
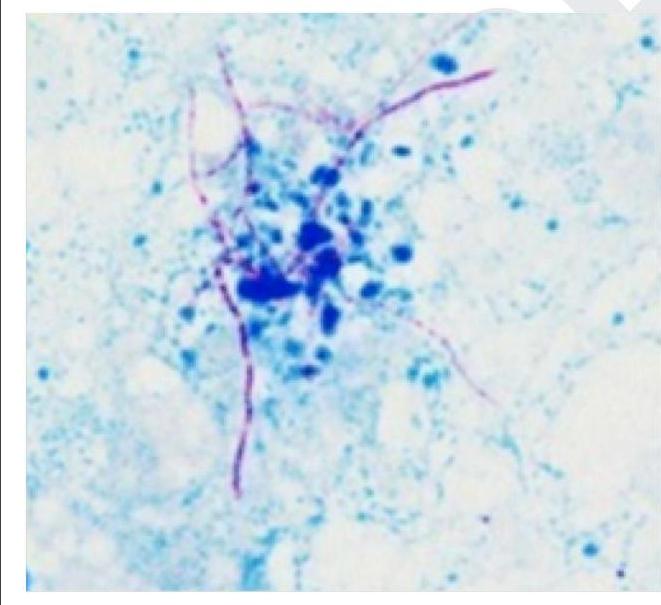
Which of the following is a gram-positive organism that shows the following appearance on Ziehl-Neelsen staining?
Gene amplification is achieved through
DNA amplification is done by all, except:
Influenza virus causes new epidemic by (3-5 yrs)-
What does Polymerase Chain Reaction (PCR) detect?
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
