Nucleic Acid Biochemistry — MCQs

On this page
A patient presents with skin cancer and hyperpigmentation that worsens with sunlight exposure. Which of the following DNA repair mechanisms is most likely defective in this condition?
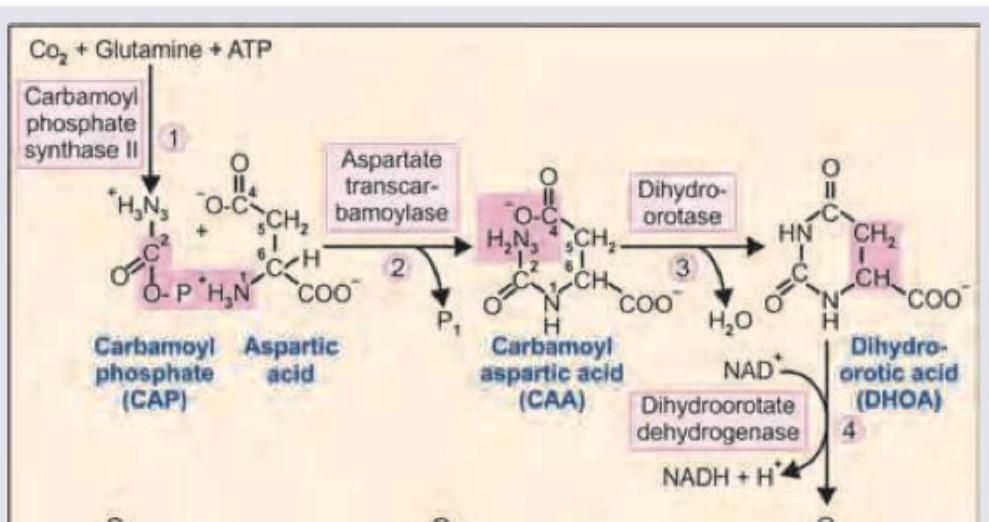
What is the treatment of the condition shown due to absence of enzyme marked as $X$ ?
Which type of bonds are represented by the dotted lines in the image? (AIIMS Nov 2017)

A patient presents with a skin rash that is exaggerated on sun exposure. What is the repair mechanism involved in this condition?
Deamination of methylated cytosine forms which of the following?
Which bases are present in human DNA?
Which organelle contains its own DNA apart from the nucleus?
Which is a product of purine metabolism?
Which part of DNA is most susceptible to radiation?
What will happen to DNA if salt is added to it: (PGI Dec 2008)
Practice by Chapter
Nucleotide Structure and Function
Practice Questions
DNA Structure and Replication
Practice Questions
RNA Structure and Types
Practice Questions
Transcription: RNA Synthesis
Practice Questions
Post-Transcriptional Modifications
Practice Questions
Translation: Protein Synthesis
Practice Questions
Genetic Code and Codon Usage
Practice Questions
Regulation of Gene Expression
Practice Questions
Mutations and DNA Repair
Practice Questions
Purine Metabolism and Disorders
Practice Questions
Pyrimidine Metabolism and Disorders
Practice Questions
Nucleotide Degradation and Salvage Pathways
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
