Biochemical Techniques — MCQs

On this page
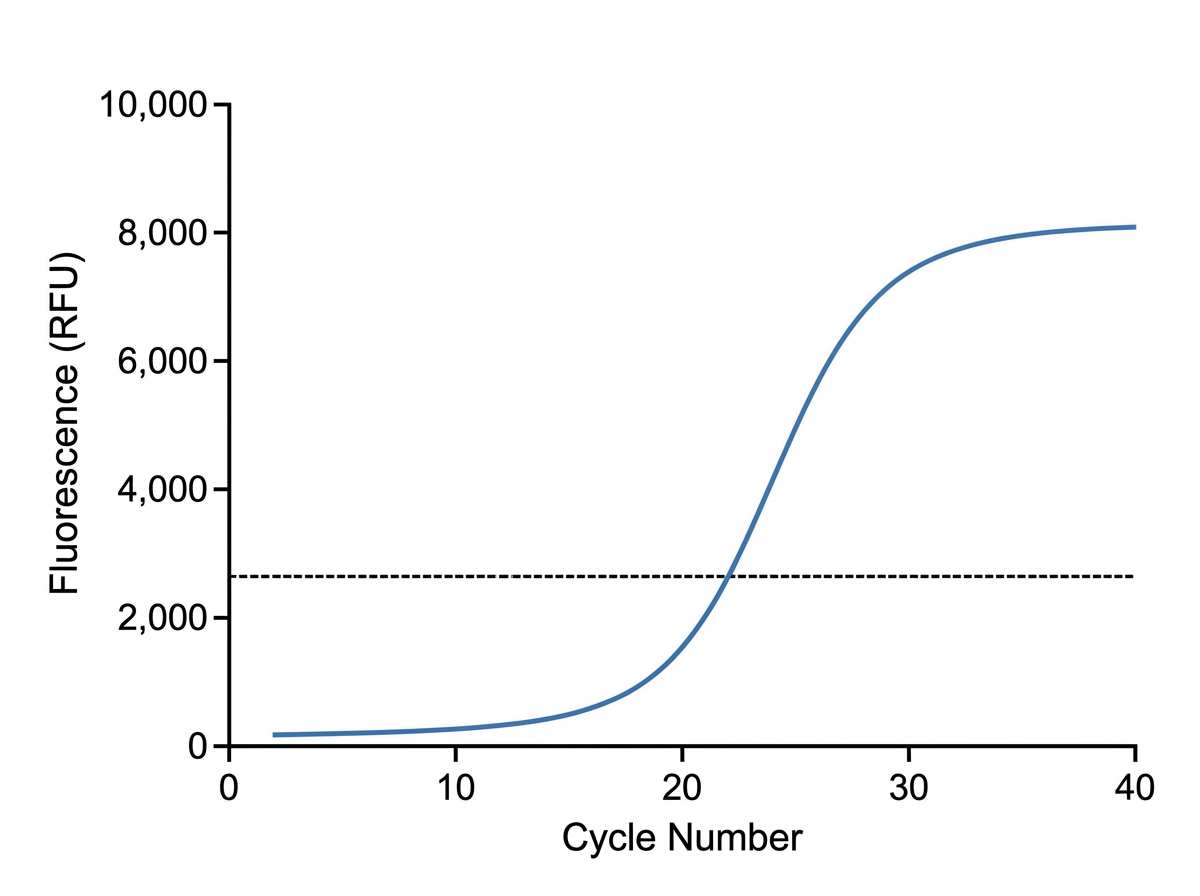
The graph below depicts a polymerase chain reaction. What type of PCR is shown?
In mass spectrometry, peptides are studied by projection of what?
Which of the following compounds is luminous?
Who invented Polymerase Chain Reaction (PCR)?
Introduction of DNA into cells with the help of electricity is known as?
Cells can be separated based on specific antigen receptors using which technique?
The number of gram equivalents in 1 litre of a solution is known as:
What is the method used to locate the isoelectric point of a protein?
Electrophoresis done under pH gradient is called?
If a radioimmunoassay is properly conducted and the amount of radioactive hormone bound to antibody is low, what would this result indicate?
Practice by Chapter
Spectrophotometry and Colorimetry
Practice Questions
Chromatography Techniques
Practice Questions
Electrophoresis and Applications
Practice Questions
Centrifugation and Ultracentrifugation
Practice Questions
Radioisotope Techniques
Practice Questions
Enzyme-Linked Immunosorbent Assay (ELISA)
Practice Questions
Polymerase Chain Reaction (PCR)
Practice Questions
Blotting Techniques: Southern, Northern, Western
Practice Questions
Mass Spectrometry in Biochemistry
Practice Questions
Recombinant DNA Technology
Practice Questions
DNA Sequencing
Practice Questions
Proteomics and Metabolomics
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
