Biochemical Techniques — MCQs

On this page
Million-Nasse's reaction is specific for which amino acid?
In a centrifugal distribution, which type of protein is precipitated first?
L-J chart is used for monitoring which of the following parameters?
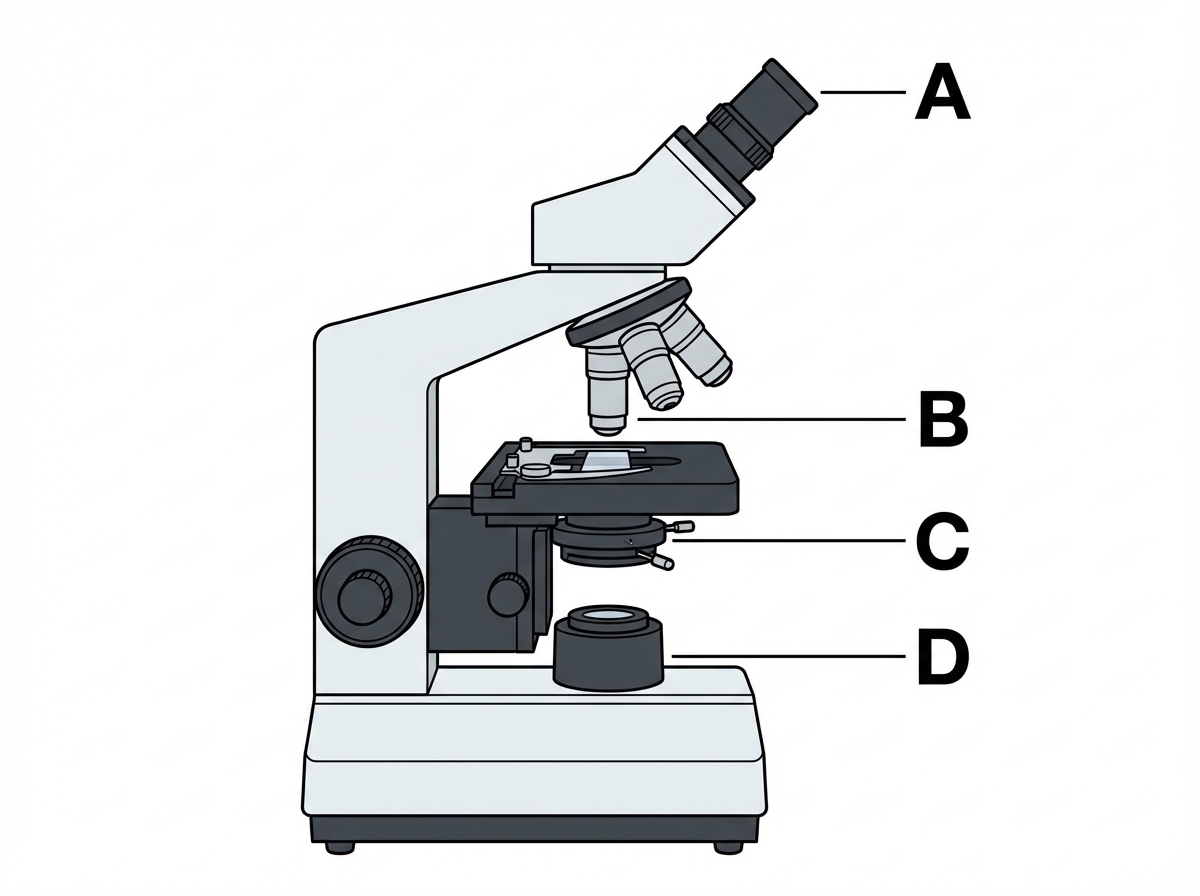
Which of the following labels corresponds to the condenser of the microscope?
Real-time PCR is primarily used for what purpose?
What specialized type of microscope enables quantitative measurement of the chemical constituents of cells?
What is essential for the Polymerase Chain Reaction (PCR)?
Which of the following statements regarding the microarray technique is false?
Which of the following methods most accurately estimates blood creatinine level?
Protein is purified using ammonium sulfate by which method?
Practice by Chapter
Spectrophotometry and Colorimetry
Practice Questions
Chromatography Techniques
Practice Questions
Electrophoresis and Applications
Practice Questions
Centrifugation and Ultracentrifugation
Practice Questions
Radioisotope Techniques
Practice Questions
Enzyme-Linked Immunosorbent Assay (ELISA)
Practice Questions
Polymerase Chain Reaction (PCR)
Practice Questions
Blotting Techniques: Southern, Northern, Western
Practice Questions
Mass Spectrometry in Biochemistry
Practice Questions
Recombinant DNA Technology
Practice Questions
DNA Sequencing
Practice Questions
Proteomics and Metabolomics
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
