RNAi & miRNA - Silencing the Messengers
- A post-transcriptional gene silencing mechanism using non-coding RNAs to regulate gene expression.
- microRNA (miRNA): Endogenous, derived from hairpin loop precursors (~70 nt). Typically causes translational repression through imperfect pairing with the 3' UTR of target mRNA.
- Small interfering RNA (siRNA): Often exogenous (e.g., viral dsRNA, synthetic). Induces mRNA cleavage via perfect or near-perfect base pairing.
⭐ The degree of complementarity dictates the outcome: perfect match (siRNA-like) leads to cleavage, while imperfect match (miRNA-like) leads to repression.
📌 siRNA → Specific interference (cleavage). 📌 miRNA → Mostly inhibitory (repression).
The Machinery - Molecular Scissors & Co.
-
Nuclear Processing (In the Nucleus):
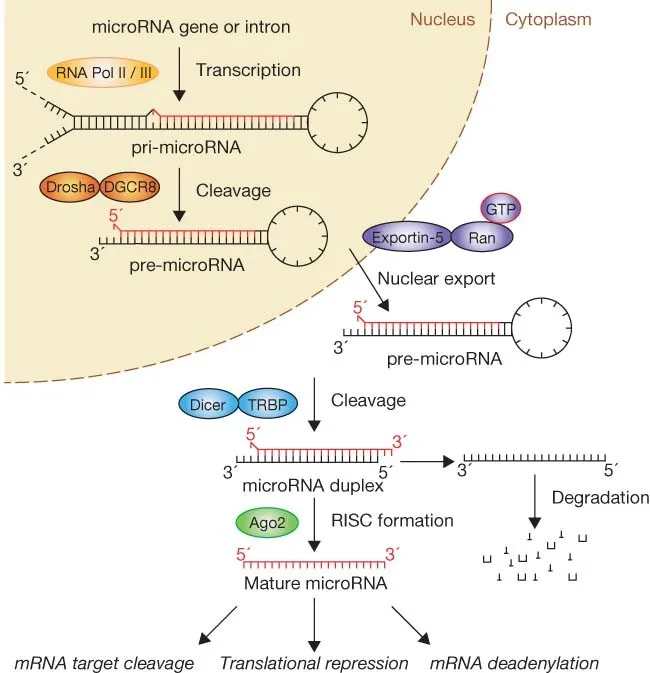
- pri-miRNA (a long primary transcript) is recognized and cleaved by the microprocessor complex, whose key enzyme is Drosha (an RNase III enzyme).
- This creates pre-miRNA, a shorter hairpin-loop structure that is then exported to the cytoplasm.
-
Cytoplasmic Processing:
- The Dicer enzyme (another RNase III) dices the pre-miRNA hairpin into a short, 22-nucleotide double-stranded RNA molecule (miRNA/siRNA).
-
RISC Loading & Silencing:
- One strand of this dsRNA is loaded onto the RNA-Induced Silencing Complex (RISC).
- This single strand guides RISC to its complementary target mRNA.
⭐ The Argonaute protein is the catalytic core of RISC, acting as "molecular scissors" to cleave the target mRNA, leading to gene silencing.
Mechanism of Action - Perfect vs. Imperfect
- Both miRNA and siRNA are loaded into the RNA-Induced Silencing Complex (RISC), which uses one strand as a guide.
- The extent of base pairing between this guide strand and a target mRNA dictates the regulatory outcome.
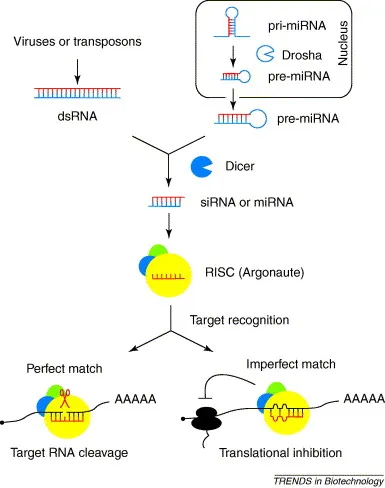
- Perfect Match (siRNA-like): Leads to direct cleavage and degradation of the target mRNA molecule by Argonaute protein (the catalytic component of RISC).
- Imperfect Match (miRNA-like): Physically hinders translation by ribosomes, leading to translational repression. The mRNA is often sequestered to P-bodies for eventual degradation.
⭐ Most endogenous human miRNAs bind with imperfect complementarity to the 3' untranslated region (3' UTR) of their target mRNAs, causing translational repression rather than cleavage.
Clinical Corner - Therapeutic Silencers
- Therapeutic RNA interference (RNAi) uses synthetic small interfering RNAs (siRNAs) to silence pathogenic genes.
- These custom-designed dsRNA molecules are introduced into cells, where they engage the RISC complex to find and cleave a specific target mRNA.
- This prevents the translation of disease-causing proteins.
- Examples:
- Patisiran & Inotersen: Target transthyretin (TTR) mRNA for hereditary amyloidosis (hATTR).
- Inclisiran: Targets PCSK9 mRNA to treat hypercholesterolemia.
⭐ Most siRNA drugs are engineered for targeted delivery to the liver, often by conjugating them to N-acetylgalactosamine (GalNAc), which binds to hepatocyte-specific receptors.
High‑Yield Points - ⚡ Biggest Takeaways
- RNA interference (RNAi) is a potent mechanism of post-transcriptional gene silencing mediated by small non-coding RNAs.
- MicroRNAs (miRNAs) are endogenous regulators, while small interfering RNAs (siRNAs) are often from exogenous sources (e.g., viruses, therapeutics).
- Precursor RNAs are processed by Drosha (nucleus) and Dicer (cytoplasm).
- The mature single-stranded RNA is loaded into the RNA-induced silencing complex (RISC).
- RISC targets mRNA for cleavage (perfect complementarity) or translational repression (imperfect match).
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

