Initiation Essentials - The Key Players
A successful translation launch requires a specific cast of molecules. Key differences exist between prokaryotic and eukaryotic systems, which are crucial for antibiotic and toxin mechanisms.

| Feature | Prokaryotes | Eukaryotes |
|---|---|---|
| Ribosome | 70S (30S + 50S) | 80S (40S + 60S) |
| Initiator tRNA | fMet-tRNA | Met-tRNA |
| mRNA Binding | Shine-Dalgarno seq. | 5' cap & Kozak seq. |
⭐ The Shine-Dalgarno sequence is a purine-rich region upstream of the AUG start codon in prokaryotes that binds to the 16S rRNA of the 30S subunit for alignment.
Prokaryotic Initiation - Shine-Dalgarno's Show
- Core Principle: The 30S ribosomal subunit recognizes and binds to the Shine-Dalgarno sequence on mRNA. This purine-rich region is located upstream of the AUG start codon, ensuring correct reading frame alignment.
- Key Players: Initiation Factors (IFs)
- IF1: Binds to and blocks the A (aminoacyl) site.
- IF3: Prevents the 50S subunit from binding prematurely.
- IF2-GTP: A G-protein that escorts the initiator N-formylmethionyl-tRNA (fMet-tRNA) to the P (peptidyl) site.
- 📌 Mnemonic: 'IF-2 escorts the initiator to the P site.'
⭐ The Shine-Dalgarno sequence (e.g., 5'-AGGAGGU-3') forms base pairs with a complementary sequence on the 16S rRNA of the 30S subunit, anchoring the mRNA correctly.
Eukaryotic Initiation - The Great 5' Cap Scan
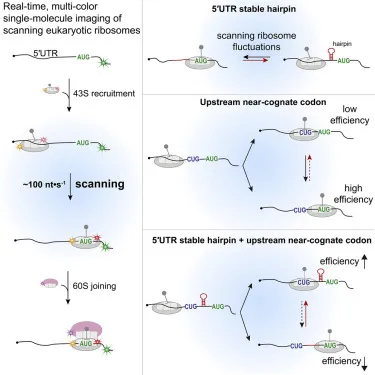
- Initiation requires assembly of the 43S preinitiation complex (40S ribosomal subunit, multiple eukaryotic initiation factors (eIFs), and initiator tRNA charged with methionine, Met-tRNAi).
- The eIF4F complex binds to the 5' cap of the mRNA and recruits the 43S complex.
- The entire assembly then "scans" the mRNA from 5' → 3' to find the AUG start codon, which is typically embedded in a Kozak consensus sequence (gcc)gccRccAUGG.
- eIF2, powered by GTP, is responsible for bringing the Met-tRNAi to the P-site of the 40S subunit.
⭐ Regulation: In response to cellular stress (e.g., amino acid starvation, viral infection), kinases phosphorylate eIF2. This locks it in an inactive GDP-bound state, preventing delivery of Met-tRNAi and globally shutting down translation.
Clinical Inhibitors - Sabotaging the Start
| Inhibitor | Mechanism of Action | Target |
|---|---|---|
| Aminoglycosides (e.g., Streptomycin) | Binds to the 30S subunit, preventing initiation complex formation. | Prokaryotic 30S subunit |
| Causes misreading of mRNA downstream. | ||
| Ricin Toxin (from castor beans) | Depurinates an adenine in the 28S rRNA via N-glycosylase activity. | Eukaryotic 60S subunit |
| This action irreversibly inactivates the ribosome. |
High-Yield Points - ⚡ Biggest Takeaways
- In eukaryotes, the 40S subunit binds the 5' cap and scans for the start codon (AUG).
- In prokaryotes, the 30S subunit binds the Shine-Dalgarno sequence upstream of the start codon.
- The initiator tRNA carries methionine in eukaryotes and formylmethionine (fMet) in prokaryotes.
- Initiation factors (IFs) are crucial for assembling the ribosome and initiator tRNA.
- Energy is supplied by GTP hydrolysis to form the initiation complex.
- The process concludes when the large subunit (60S or 50S) joins, placing the initiator tRNA in the P site.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

