RNA Splicing - The Pre-mRNA Haircut
- What: Process in the nucleus where introns (intragenic regions) are removed from pre-mRNA, and exons (expressed regions) are joined.
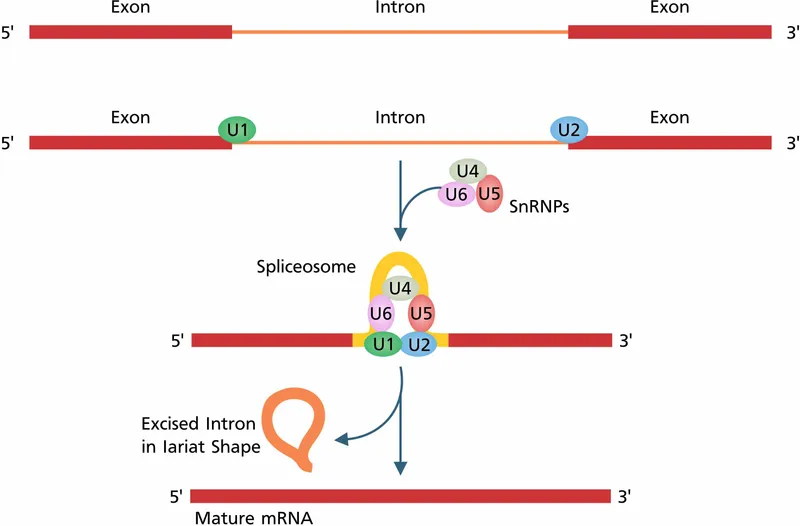
- Who: Catalyzed by the spliceosome, a large complex of small nuclear RNAs (snRNAs) and proteins, forming snRNPs ("snurps").
- How:
- Recognizes specific splice sites.
- 5' site: GU sequence.
- 3' site: AG sequence.
- Forms a lariat-shaped intermediate, which is then excised.
⭐ Autoantibodies to spliceosomal snRNPs (anti-Smith antibodies) are highly specific for Systemic Lupus Erythematosus (SLE).
Splicing Mechanism - Cut, Lariat, Paste
- Spliceosome: A large RNA-protein complex, composed of five snRNPs (U1, U2, U4, U5, U6) and other proteins, that removes introns from a pre-mRNA transcript.
- Key Consensus Sequences:
- 5' Splice Site: GU at the 5' end of the intron.
- 3' Splice Site: AG at the 3' end of the intron.
- Branch Point: An adenine (A) nucleotide ~20-50 bases upstream of the 3' site.
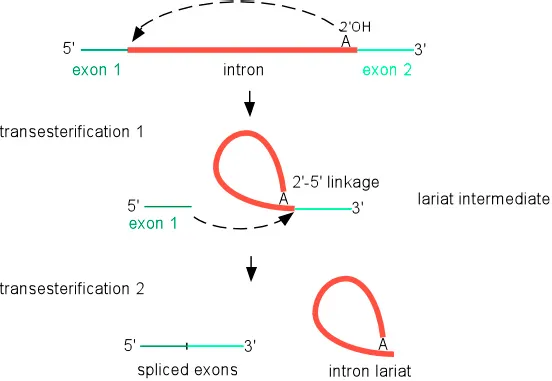
⭐ Splicing proceeds via two sequential transesterification reactions. While the reactions themselves don't consume ATP, ATP hydrolysis is required for spliceosome assembly and rearrangement on the pre-mRNA.
Alternative Splicing - One Gene, Many Proteins
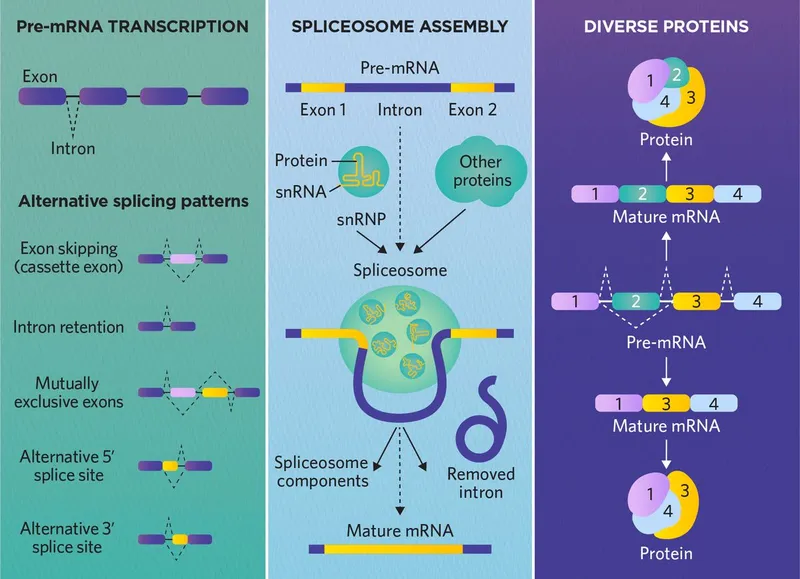
- Core Concept: A single gene's pre-mRNA can be processed in multiple ways to generate different mature mRNAs and, subsequently, different protein isoforms.
- Mechanism:
- Specific exons are selectively included or excluded from the final mRNA.
- Regulated by tissue-specific splicing factors (proteins) that bind to enhancer or silencer sequences on the pre-mRNA.
- Significance: Greatly increases the coding potential of the genome, allowing a single gene to produce a variety of proteins with different functions or properties.
⭐ Tropomyosin and the calcitonin gene are classic examples. In the thyroid, the gene produces calcitonin; in the brain, it produces calcitonin gene-related peptide (CGRP).
Splicing Errors - When the Cut Goes Wrong
- Mutations at splice sites (GU donor, AG acceptor) or branch point sequences disrupt intron removal.
- Leads to:
- Exon skipping: An exon is incorrectly spliced out.
- Intron retention: An intron remains in the mature mRNA.
- Cryptic splice site activation: A nearby, similar sequence is used instead of the correct splice site.
- These errors often cause frameshift mutations and premature stop codons, yielding truncated or unstable proteins.
- Clinical Correlates:
- β-thalassemia
- Gaucher disease
- Myelodysplastic syndromes
- Some forms of Marfan syndrome
⭐ In certain β-thalassemias, a mutation within an intron creates a new, cryptic splice acceptor site. This incorporates part of the intron into the mRNA, causing a frameshift and producing a nonfunctional β-globin protein.
High‑Yield Points - ⚡ Biggest Takeaways
- Splicing removes introns and joins exons via the spliceosome, a complex of snRNPs.
- Recognizes consensus sequences: GU at the 5' donor site and AG at the 3' acceptor site.
- An intermediate lariat loop of the intron is formed and then excised.
- Alternative splicing generates different protein isoforms from a single gene, increasing proteomic diversity.
- Anti-Smith antibodies (specific for SLE) target snRNPs.
- Splicing-site mutations can cause diseases like certain β-thalassemias.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

