DNA Methylation - Silencing the Code
- Mechanism: Covalent addition of a methyl group to cytosine bases within CpG dinucleotides (CpG islands).
- Enzyme: DNA Methyltransferase (DNMT).
- Methyl Donor: S-adenosyl methionine (SAM).
- Function: Gene silencing. Methylated DNA physically obstructs transcription factors and recruits histone deacetylases (HDACs), leading to chromatin condensation (heterochromatin).
- Clinical Relevance:
- Genomic imprinting (e.g., Prader-Willi, Angelman syndromes).
- X-chromosome inactivation (Lyonization).
- Tumor suppressor gene silencing in cancer.
⭐ CpG islands are often located in the promoter regions of genes. Hypermethylation of a tumor suppressor gene's promoter CpG island is a common mechanism for its inactivation in neoplasia.
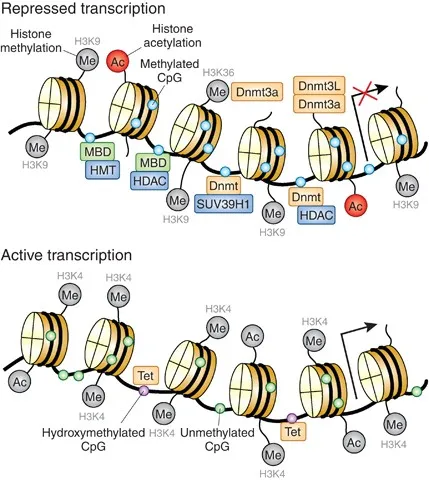
Histone Modification - Spooling the Rules
-
Core Concept: Positively charged lysine residues in histones bind to negatively charged DNA phosphate groups, regulating DNA accessibility.
-
Histone Acetylation:
- Mechanism: Histone Acetyltransferases (HATs) add acetyl groups ($CH_3CO$) to lysine, neutralizing its positive charge.
- Result: ↓ DNA-histone affinity → relaxed chromatin (euchromatin) → ↑ transcription.
- 📌 Histone Acetylation Turns genes Active.
-
Histone Deacetylation:
- Mechanism: Histone Deacetylases (HDACs) remove acetyl groups.
- Result: ↑ DNA-histone affinity → condensed chromatin (heterochromatin) → ↓ transcription.
-
Histone Methylation:
- Mechanism: Histone Methyltransferases (HMTs) add methyl groups ($CH_3$) to lysine or arginine.
- Result: Variable; typically causes reversible gene silencing (heterochromatin), but can also activate transcription depending on the location.
- 📌 Histone Methylation Makes DNA Mute (usually).
⭐ Huntington Disease: Involves abnormal huntingtin protein interaction with HATs and HDACs, leading to histone hypoacetylation and transcriptional repression of neuroprotective genes.
Genomic Imprinting - Parental Grudges
- Process where gene expression is determined by parental origin. Only one allele (maternal or paternal) is expressed, while the other is silenced via methylation.
- Mechanism: Methylation of CpG islands in the promoter region silences the gene. This occurs during gametogenesis.

- Clinical Syndromes (Chromosome 15q11-q13):
- Prader-Willi Syndrome: Paternal allele is deleted or mutated.
- Maternal allele is normally imprinted (silenced).
- Features: Hyperphagia, obesity, intellectual disability, hypogonadism.
- 📌 Prader-Willi: Paternal deletion.
- Angelman Syndrome: Maternal allele is deleted or mutated.
- Paternal allele is normally imprinted (silenced).
- Features: "Happy puppet," seizures, ataxia, severe intellectual disability.
- 📌 Angelman: Maternal deletion.
- Prader-Willi Syndrome: Paternal allele is deleted or mutated.
⭐ Uniparental disomy (UPD), where an individual inherits two copies of a chromosome from one parent and none from the other, is another cause for both Prader-Willi (maternal UPD) and Angelman (paternal UPD) syndromes.
High‑Yield Points - ⚡ Biggest Takeaways
- DNA methylation at CpG islands typically represses transcription.
- Histone acetylation creates euchromatin (relaxed), making DNA accessible and ↑ transcription.
- Histone deacetylation forms heterochromatin (condensed), silencing genes by ↓ transcription.
- Histone methylation can either activate or repress transcription depending on the methylation site.
- Genomic imprinting silences genes based on parental origin, as seen in Prader-Willi and Angelman syndromes.
- Fragile X syndrome is caused by hypermethylation of a CGG repeat expansion, silencing the FMR1 gene.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

