Mutation and Mutagenesis - Genetic Code Glitches
- Mutation: Heritable change in DNA.
- Types:
- Point Mutations (single base affected):
- Silent: Codes for same amino acid (AA); no change in protein.
- Missense: Codes for different AA; may alter protein function.
- Nonsense: Codes for STOP codon (UAG, UAA, UGA); premature termination.
- Frameshift Mutations: Insertion or deletion of bases not in multiples of 3; alters reading frame downstream, often creating non-functional protein.
- Point Mutations (single base affected):
- Mutagens (Causes):
- Spontaneous: e.g., DNA replication errors, deamination.
- Induced:
- Physical: UV radiation (forms pyrimidine dimers, e.g., thymine dimers), Ionizing radiation (X-rays, gamma rays - cause DNA strand breaks).
- Chemical: Base analogs (e.g., 5-bromouracil), Intercalating agents (e.g., ethidium bromide, acridine orange), Alkylating agents.
and frameshift mutation)
⭐ Nonsense mutations (UAG, UAA, UGA) result in premature termination of protein synthesis, often leading to truncated, non-functional proteins. 📌 STop A Garbage, STop All Activity, STop Going Anywhere (UAG, UAA, UGA are stop codons).
Mutation and Mutagenesis - Agents of Alteration
Mutagens: Agents causing DNA damage, increasing mutation rates.
Table: Classification of Mutagens
| Type | Class | Examples & Key Effect(s) |
|---|---|---|
| Physical | Ionizing Radiation | X-rays, γ-rays: Breaks, base damage. |
| Non-ionizing | UV (260nm): Pyrimidine dimers. | |
| Chemical | Base Analogs | 5-Bromouracil (5-BU), 2-Aminopurine (2-AP): Mispairing. |
| Alkylating Agents | EMS, MNNG: Alkylation → mispairing. | |
| Intercalating Agents | Ethidium Br, Acridines: Frameshifts. | |
| Deaminating Agents | Nitrous acid ($HNO_2$): C→U, A→HX (mispairing). | |
| Hydroxylating Agents | Hydroxylamine ($NH_2OH$): C→N-OH-C (pairs A). |
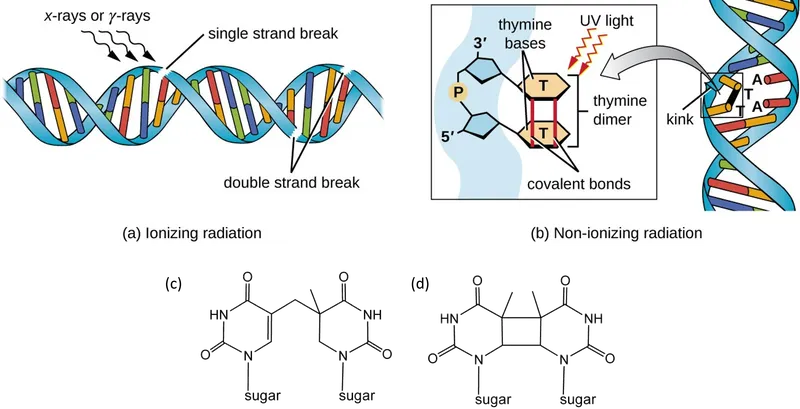
Mutation and Mutagenesis - Cellular Fix-It Crew
- Cells employ diverse DNA repair systems to maintain genomic stability against spontaneous or induced damage.
- Major Repair Pathways:
- Direct Reversal:
- Photoreactivation: Light-dependent repair of pyrimidine dimers by photolyase.
- Excision Repair:
- Base Excision Repair (BER): Targets non-bulky lesions (e.g., uracil, alkylated bases).
- Nucleotide Excision Repair (NER): Removes bulky adducts & pyrimidine dimers (e.g., UV damage).
- Mismatch Repair (MMR): Corrects replication errors (base mismatches, small indels).
- Double-Strand Break (DSB) Repair:
- Homologous Recombination (HR): High-fidelity, uses sister chromatid.
- Non-Homologous End Joining (NHEJ): Error-prone, direct ligation.
- Direct Reversal:
⭐ The SOS response in bacteria is an inducible, error-prone DNA repair system activated by extensive DNA damage, increasing mutation rates but allowing survival.
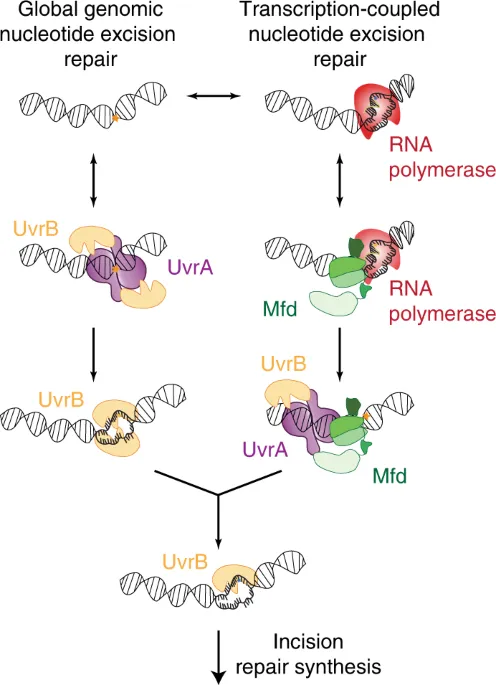
Mutation and Mutagenesis - Effects & Detection
-
Effects of Mutations:
- Phenotypic:
- Silent: No amino acid (AA) change.
- Missense: Different AA; conservative or non-conservative.
- Nonsense: Premature STOP codon; truncated protein.
- Frameshift: Insertion/deletion (not multiple of 3); altered reading frame.
- Functional: Loss-of-function (null, hypomorph), Gain-of-function (hypermorph, neomorph), Lethal, Conditional (e.g., temperature-sensitive), Antibiotic resistance.
- Phenotypic:
-
Detection of Mutations:
- Phenotypic Screening: Replica plating (auxotrophs, antibiotic resistance), specific indicator media.
- Genotypic Screening: PCR-based methods (e.g., RFLP, ASO probes), DNA sequencing.
-
Detection of Mutagens (Ames Test):
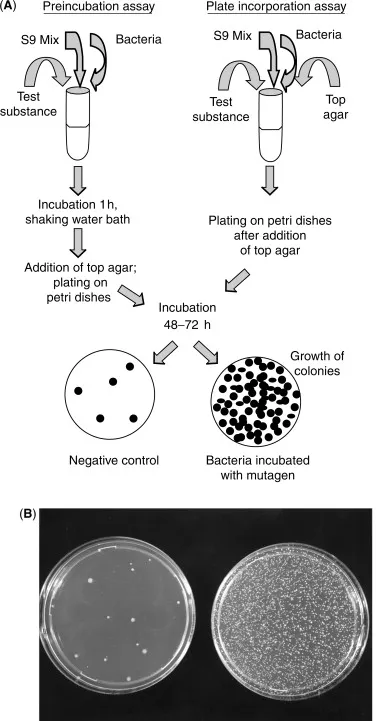
⭐ The Ames test utilizes histidine auxotrophs of Salmonella typhimurium to screen for chemical mutagens by detecting reversion to prototrophy, often with liver extract (S9) to simulate metabolic activation.
High‑Yield Points - ⚡ Biggest Takeaways
- Point mutations: silent (no change), missense (amino acid change), nonsense (premature STOP codon).
- Frameshift mutations: insertions/deletions not multiple of 3, alters downstream reading frame.
- Mutagens: Physical (UV radiation → pyrimidine dimers); Chemical (base analogs, intercalating agents).
- Ames test: detects chemical mutagens via reversion in auxotrophic bacteria (e.g., Salmonella His-).
- DNA repair: NER (Nucleotide Excision Repair) fixes pyrimidine dimers; MMR (Mismatch Repair) corrects replication errors.
- Transposons ("jumping genes"): cause insertional mutagenesis and can alter gene expression.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

