Transcription Overview - Blueprinting Life's Code
- Core Process: Selective synthesis of RNA from a DNA template strand.
- Key Enzyme: DNA-dependent RNA Polymerase (RNAP); no primer needed.
- Synthesis Direction: New RNA chain grows exclusively 5' → 3'.
- DNA Template Strand: Antisense strand, read by RNAP in the 3' → 5' direction.
- DNA Coding Strand: Sense strand; its sequence matches RNA (Uracil replaces Thymine).
- Fundamental Stages: Initiation at promoter, Elongation, Termination at signal.
⭐ Transcription is the primary regulatory point for gene expression.
 oka
oka
Prokaryotic Transcription - Bacterial RNA Show
- Enzyme: DNA-dependent RNA Polymerase (RNAP).
- Holoenzyme: $\alpha_2\beta\beta'\omega\sigma$ (sigma for initiation).
- Core enzyme: $\alpha_2\beta\beta'\omega$ (for elongation).
- Promoter Elements: Recognized by $\sigma$ factor.
- -10 sequence (Pribnow box: TATAAT).
- -35 sequence (TTGACA).
- Process: No primer needed.
- Inhibitor: Rifampicin (binds $\beta$ subunit). 📌 RNA Pol inhibited by Rifampicin.
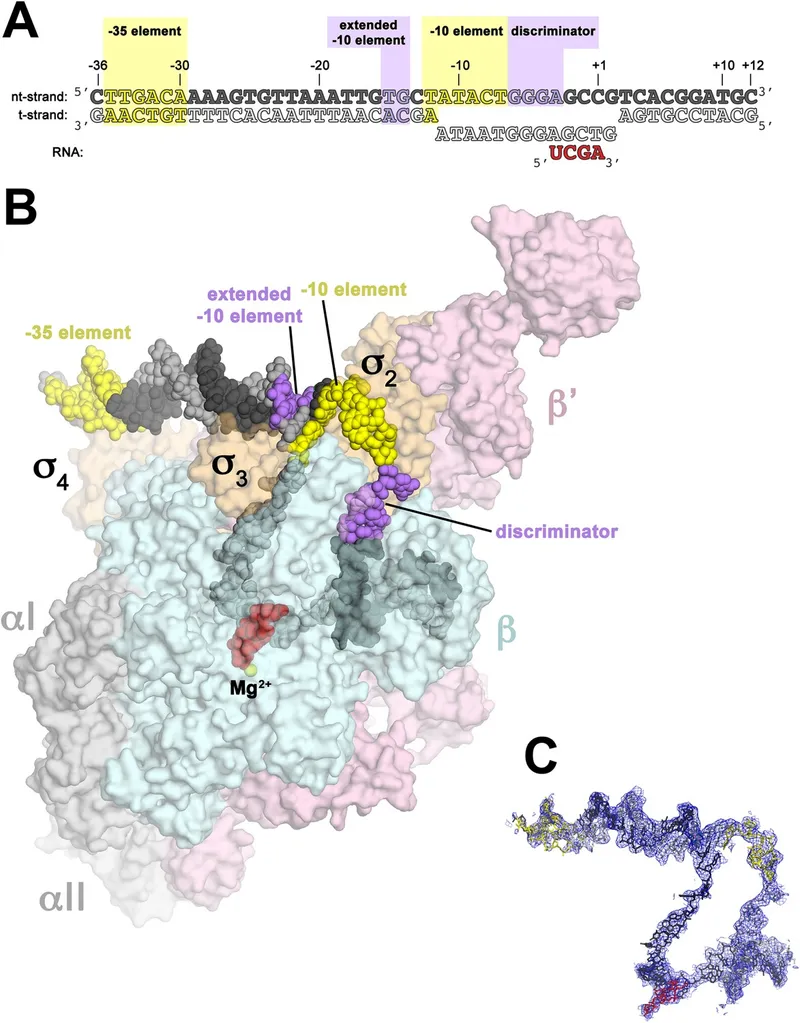
⭐ The $\sigma$ factor is crucial for promoter specificity in prokaryotic transcription, dissociating after initiation.
Eukaryotic Transcription - Complex RNA Symphony
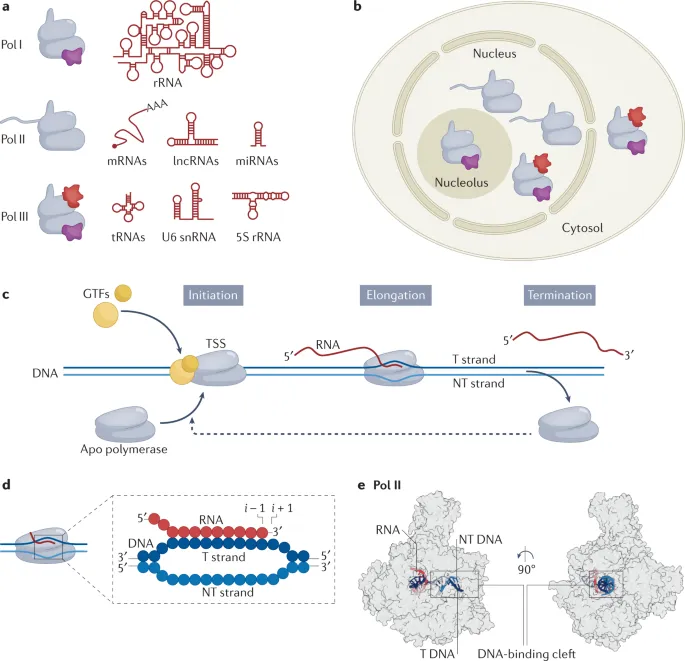
- RNA Polymerases (RNAPs) & Location:
- RNAP I: rRNA (28S, 18S, 5.8S) - Nucleolus.
- RNAP II: mRNA, snRNA, miRNA - Nucleoplasm. Highly sensitive to α-amanitin.
- RNAP III: tRNA, 5S rRNA, U6 snRNA - Nucleoplasm.
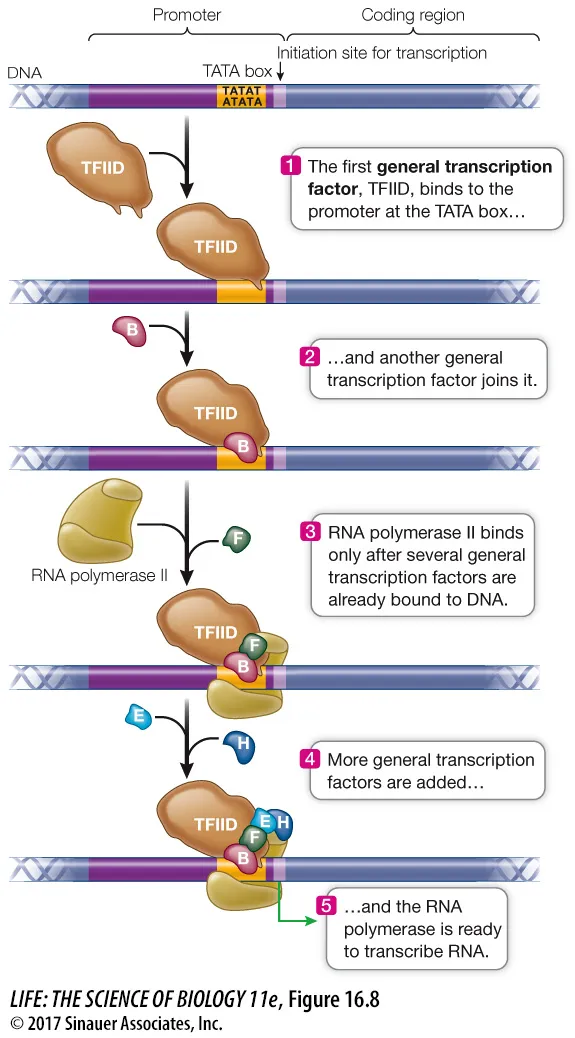
- Promoter Elements: Core (TATA box, Inr, DPE, BRE) & Proximal (CAAT box, GC box).
- General Transcription Factors (GTFs): e.g., TFIID (contains TATA-Binding Protein - TBP), TFIIH (helicase & kinase activity).
- Regulatory Elements: Enhancers & Silencers (distal, position/orientation-independent).
- Chromatin Remodeling: Histone modifications (e.g., acetylation, methylation) essential for access.
- RNA Processing (Co-transcriptional): 5' capping (7-methylguanosine), 3' polyadenylation (poly-A tail), Splicing (introns removed by spliceosome).
⭐ α-Amanitin, a toxin from Amanita phalloides (death cap mushroom), potently inhibits RNA Polymerase II, halting mRNA synthesis. At higher concentrations, it also inhibits RNA Pol III.
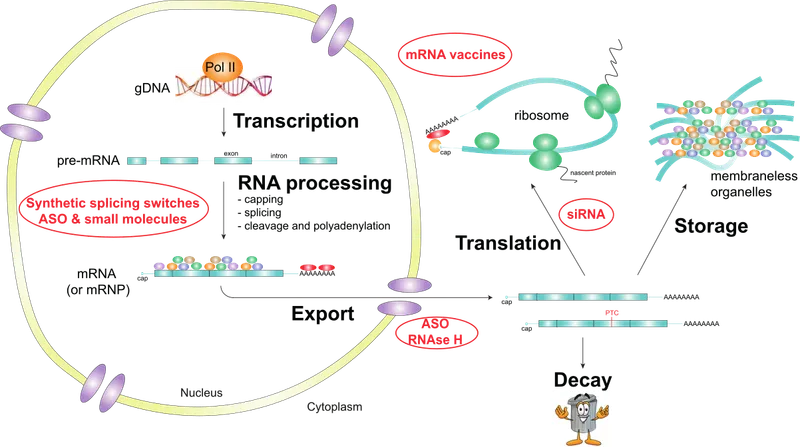
RNA Processing - Post-Transcriptional Tailoring
- Eukaryotic hnRNA (pre-mRNA) tailored to mature mRNA.
- 5' Capping: 7-methylguanosine (7-mG) added; 5'-5' triphosphate linkage.
- Protects, aids ribosome binding.
- Splicing: Introns removed, exons joined by spliceosome (snRNPs: U1, U2, U4, U5, U6).
- Consensus sites: GU (5' splice), AG (3' splice).
- Alternative splicing ↑ protein diversity.
- 3' Polyadenylation: Poly(A) tail (~200-250 A's) by poly(A) polymerase.
- Signal: AAUAAA.
- Enhances stability, export, translation.
⭐ Systemic Lupus Erythematosus (SLE) patients often have autoantibodies against spliceosomal snRNPs (e.g., anti-Sm, anti-RNP).
Transcription Inhibitors - Silencing the Message
- Rifampicin: Binds prokaryotic RNA Pol (β-subunit), blocks initiation. For TB. 📌 "R for Rifampicin, R for RNA Pol".
- Actinomycin D: Intercalates DNA, stalls RNA Pol (prokaryotes & eukaryotes). Anticancer.
- α-Amanitin: Inhibits eukaryotic RNA Pol II (mRNA synthesis). From Amanita phalloides.
⭐ α-Amanitin specifically inhibits RNA Polymerase II, halting mRNA synthesis and leading to severe hepatotoxicity.
- Fidaxomicin: Inhibits bacterial RNA Pol; treats C. difficile infection.
High‑Yield Points - ⚡ Biggest Takeaways
- Eukaryotic RNA Polymerases: Pol I (major rRNAs), Pol II (mRNA, snRNA), Pol III (tRNA, 5S rRNA).
- Single Prokaryotic RNA Polymerase needs sigma (σ) factor for promoter recognition.
- Key promoter elements: Pribnow box (prokaryotes, -10), TATA box (eukaryotes, -25).
- Transcription is 5' → 3', reads template 3' → 5'; no primer is needed.
- Key inhibitors: Rifampicin (prokaryotic RNA Pol), α-amanitin (eukaryotic RNA Pol II).
- mRNA sequence matches coding strand (U for T), complementary to the template DNA strand.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more