DNA Structure - Helix Blueprint
- DNA: Deoxyribonucleotide polymer; genetic blueprint.
- Components: Deoxyribose, phosphate, N-bases (A,T,G,C).
- Purines: Adenine (A), Guanine (G) (📌 Pure As Gold).
- Pyrimidines: Cytosine (C), Thymine (T) (📌 CUT Py).
- Nucleoside: Base + Sugar. Nucleotide: Base + Sugar + Phosphate.
- 5'-3' Phosphodiester bonds form backbone.
- Watson-Crick B-DNA: Right-handed double helix.
- Antiparallel strands.
- Complementary base pairing (Chargaff's rules):
- A=T (2 H-bonds)
- G≡C (3 H-bonds)
- Purines (A+G) = Pyrimidines (T+C)
- Dimensions (B-DNA):
- ~10.5 base pairs/turn.
- Pitch: 3.4 nm/turn.
- Diameter: 2 nm.
- Major & minor grooves: Sites for protein binding.
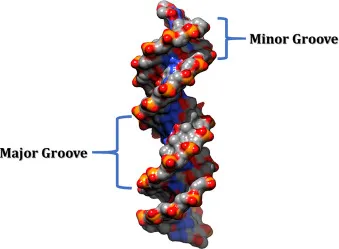
⭐ B-DNA: right-handed helix; 10.5 bp/turn, pitch 3.4 nm. Z-DNA: left-handed.
DNA Packaging - Chromatin Coils
- DNA (~2m) compacts into nucleus (~10µm) through hierarchical coiling.
- Nucleosome ("beads-on-a-string"): Fundamental unit.
- DNA (146 bp) wraps ~1.65 times around a histone octamer (2 each of H2A, H2B, H3, H4).
- H1 histone (linker histone) binds to linker DNA and the nucleosome, aiding compaction.
- Forms a 10-11 nm fiber.
- Solenoid:
- Helical coiling of 6 nucleosomes per turn.
- Forms a 30 nm fiber.
- Higher-order packaging: 30 nm fiber forms loops (300 nm), then scaffolds (700 nm), ultimately condensing into a metaphase chromosome (1400 nm).
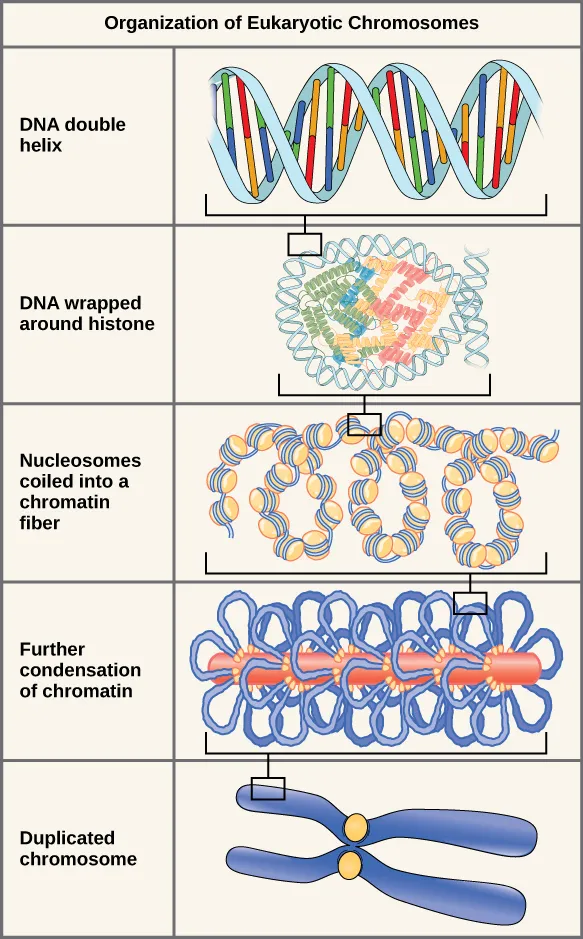
⭐ Histones are rich in basic amino acids (Lysine and Arginine), giving them a net positive charge that facilitates binding to the negatively charged DNA backbone.
DNA Replication - Copy Machine
DNA replication is a semi-conservative process, ensuring each daughter DNA molecule has one parental and one newly synthesized strand. It occurs primarily during the S-phase of the cell cycle.
- Key Steps & Enzymes:
- Initiation:
- Origin of Replication (ORI): Specific DNA sequences where replication begins.
- Helicase: Unwinds DNA double helix at replication fork (ATP-dependent).
- Single-Strand Binding Proteins (SSBPs): Stabilize unwound single-stranded DNA (ssDNA).
- Topoisomerases: Relieve supercoiling ahead of fork (e.g., DNA gyrase in prokaryotes).
- Elongation:
- Primase: Synthesizes short RNA primers.
- DNA Polymerase III: Main prokaryotic synthesizing enzyme (5'→3' activity), requires RNA primer.
- Leading strand: Continuous synthesis towards the fork.
- Lagging strand: Discontinuous synthesis (Okazaki fragments) away from the fork.
- DNA Polymerase I: Prokaryotic; removes RNA primers (5'→3' exonuclease activity), fills gaps with DNA.
- Termination:
- DNA Ligase: Joins Okazaki fragments on lagging strand, seals nicks in the phosphodiester backbone.
- Initiation:
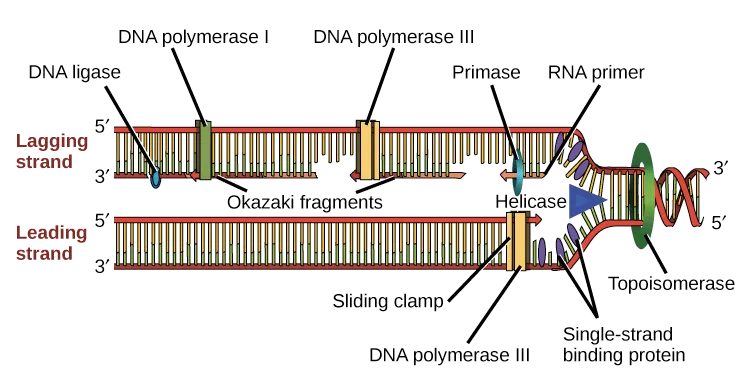
⭐ DNA replication is typically bidirectional from the origin, creating two replication forks. Prokaryotes usually have a single ORI, while eukaryotes possess multiple ORIs to ensure timely replication of larger genomes.
Replication Fidelity & Telomeres - Proof & Protect
- Replication Fidelity (Error Prevention):
- DNA Pol III (prok.) & Pol δ/ε (euk.) 3'→5' exonuclease activity.
- Mismatch Repair (MMR): Corrects residual errors. Proteins: MutS/L/H (prok.), MSH/MLH/PMS (euk.).
- Achieves error rate of $1 \times 10^{-9}$ to $1 \times 10^{-10}$.
- Telomeres (Chromosome End Caps):
- Repetitive DNA (human: TTAGGG) at chromosome ends; prevent gene erosion.
- Counteract "end replication problem" (incomplete lagging strand synthesis).
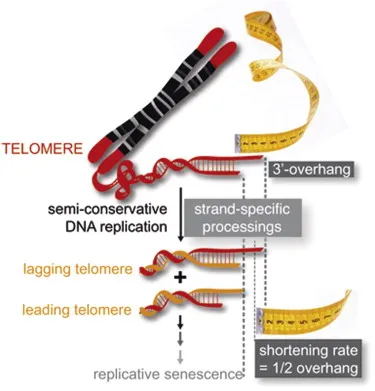
- Telomerase (Maintains Telomeres):
- Ribonucleoprotein enzyme (reverse transcriptase activity).
- Contains RNA template (TERC) & catalytic subunit (TERT).
- Adds telomeric repeats. High in germ, stem, cancer cells. Low in somatic cells (→ Hayflick limit).
- ⭐ > Telomerase inhibition is a therapeutic target in cancer treatment.
High‑Yield Points - ⚡ Biggest Takeaways
Error: Failed to generate content for this concept group.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

