Enzyme Basics & Structure - Protein Powerhouses
- Enzymes: Biological catalysts (mostly proteins) that ↑ reaction rates; not consumed.
- Properties: High catalytic power, specificity (substrate-specific), regulated activity.
- IUBMB Classification (6 Classes): 📌 Mnemonic: Oh These Huge Lads Are Little (OTHLIL).
Class Reaction Type 1. Oxidoreductases Redox reactions 2. Transferases Group transfer 3. Hydrolases Hydrolysis 4. Lyases Group removal/addition (non-hydrolytic) 5. Isomerases Isomerization 6. Ligases Joining molecules (ATP-dependent) - Components:
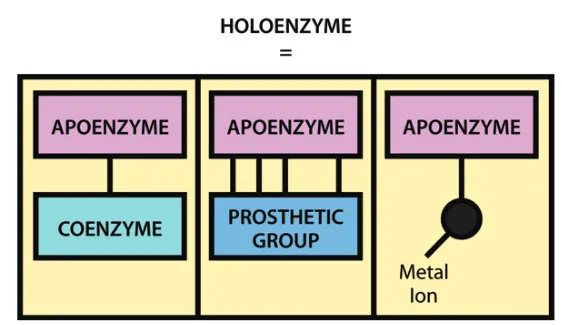
- Apoenzyme: Inactive protein part.
- Cofactor: Non-protein part for activity (inorganic ions like $Mg^{2+}$, $Zn^{2+}$).
- Coenzyme: Organic cofactor (often vitamin-derived).
- Prosthetic Group: Tightly bound coenzyme/cofactor.
- Holoenzyme: Active enzyme; $Apoenzyme + Cofactor \rightleftharpoons Holoenzyme$.
- Active Site: Specific substrate-binding region; contains binding & catalytic sites.
⭐ Ribozymes: RNA with enzymatic activity, not all enzymes are proteins.
Enzyme Action & Kinetics - Reaction Accelerators
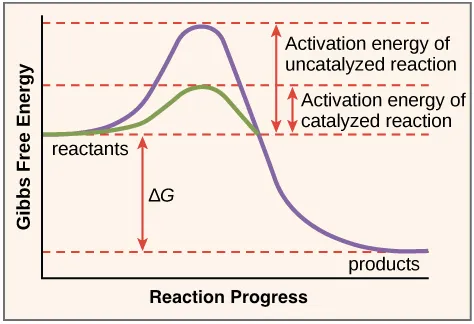
- Mechanism: Enzymes accelerate reactions by lowering activation energy ($E_a$) via transition state stabilization.
- Models:
- Lock-and-Key: Rigid active site precisely fits substrate.
- Induced-Fit: Substrate binding induces enzyme conformational change for optimal fit.
- Kinetics:
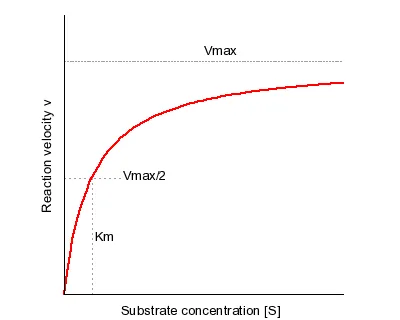
- Michaelis-Menten Equation: $V_0 = \frac{V_{max}[S]}{K_m + [S]}$
- $K_m$ (Michaelis Constant): [S] at $\frac{1}{2}V_{max}$. Inverse measure of affinity (↓$K_m$ = ↑affinity).
- 📌 $K_m$: Koncentration for half Max.
⭐ $K_m$ is numerically equal to the substrate concentration at which reaction velocity is $\frac{1}{2}V_{max}$.
- $V_{max}$ (Maximum Velocity): Rate at enzyme saturation; proportional to [Enzyme].
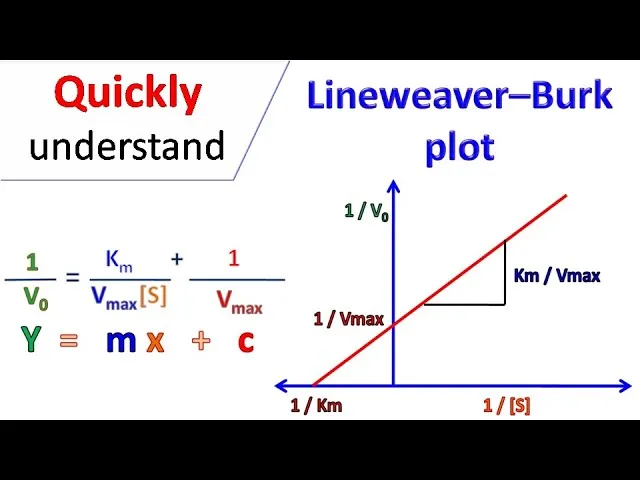
- Lineweaver-Burk Plot: $\frac{1}{V_0} = (\frac{K_m}{V_{max}})\frac{1}{[S]} + \frac{1}{V_{max}}$
- X-intercept: $-\frac{1}{K_m}$; Y-intercept: $\frac{1}{V_{max}}$; Slope: $\frac{K_m}{V_{max}}$.
- Michaelis-Menten Equation: $V_0 = \frac{V_{max}[S]}{K_m + [S]}$
- Factors Affecting Activity:
- [Substrate]: ↑[S] → ↑$V_0$ until $V_{max}$.
- [Enzyme]: ↑[Enzyme] → ↑$V_{max}$.
- Temperature: Optimal ~37°C (human); extremes denature.
- pH: Optimal pH specific; extremes denature.
Enzyme Inhibition & Regulation - Catalyst Controllers
- Reversible Inhibition: Comparison of types:
Type Binding Site $K_m$ $V_{max}$ Lineweaver-Burk Plot Mnemonic 📌 Competitive Active site ↑ ↔ Intersect on Y-axis Competitive: $K_m$↑, $V_{max}$↔ (Kome Up, $V_{max}$ stays) Non-competitive Allosteric site ↔ ↓ Intersect on X-axis Non-competitive: $K_m$↔, $V_{max}$↓ ($K_m$ stays, $V_{max}$ decays) Uncompetitive ES complex only ↓ ↓ Parallel lines Uncompetitive: $K_m$↓, $V_{max}$↓ (UNique: both UNder) 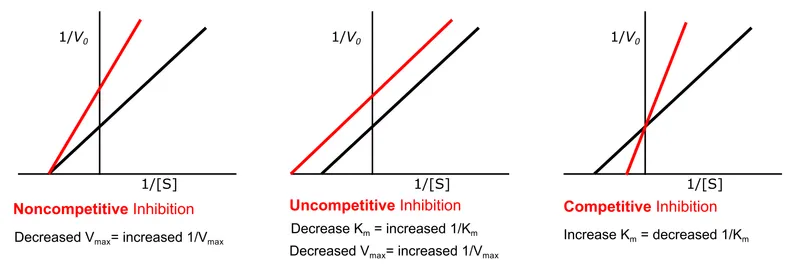
- Irreversible Inhibition: Covalent modification of enzyme, often "suicide" inhibitors (e.g., penicillin, aspirin, organophosphates).
- Allosteric Regulation: Modulators bind to allosteric site (not active site), altering enzyme activity (positive/negative). Shows cooperativity, sigmoidal kinetics.
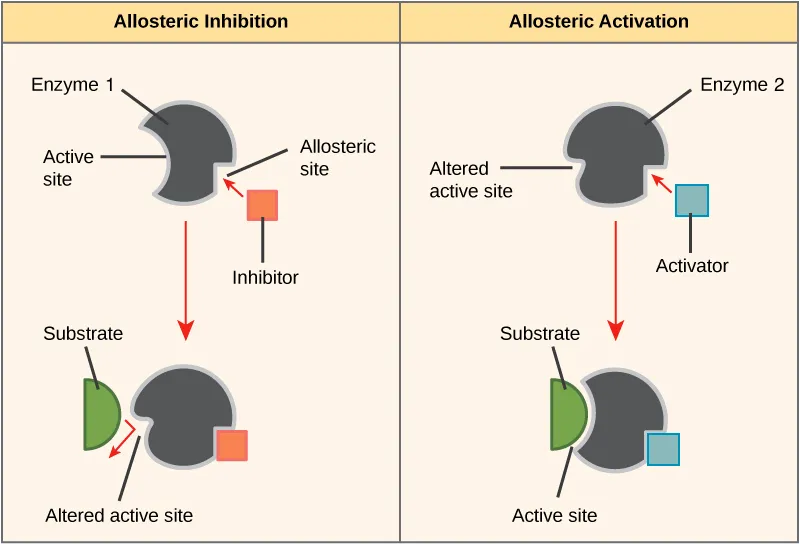
- Covalent Modification: Activity regulated by adding/removing chemical groups.
- Phosphorylation/dephosphorylation (kinases/phosphatases).
- Zymogen activation: inactive precursor (e.g., pepsinogen) cleaved to active enzyme (e.g., pepsin).
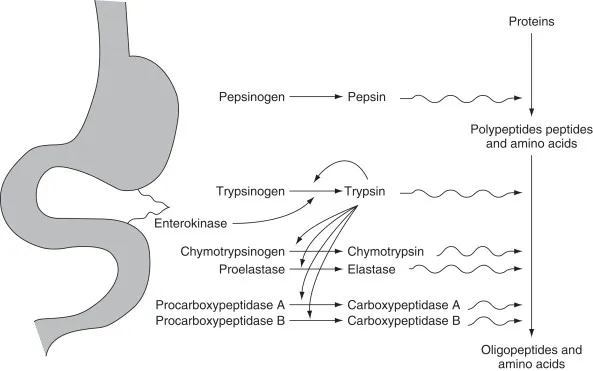
- Feedback Inhibition: End product of a metabolic pathway inhibits an early enzyme in that pathway, regulating its own synthesis.
- Clinical Inhibitors (Examples):
- Statins (e.g., atorvastatin): Competitive inhibitors of HMG-CoA reductase (cholesterol synthesis).
- ACE inhibitors (e.g., captopril): Treat hypertension.
- Allopurinol: Inhibits xanthine oxidase (gout treatment).
⭐ Methanol poisoning is treated with ethanol, which acts as a competitive inhibitor for alcohol dehydrogenase, preventing methanol's conversion to toxic formaldehyde.
High‑Yield Points - ⚡ Biggest Takeaways
- Enzymes: Biological catalysts that lower activation energy, not consumed in reactions.
- Active site: Binds substrate; specificity by Lock & Key or Induced Fit models.
- Michaelis-Menten kinetics: Km indicates substrate affinity (↑Km = ↓affinity); Vmax is maximum velocity.
- Competitive inhibitors: Bind active site, ↑Km, Vmax unchanged.
- Non-competitive inhibitors: Bind allosteric site, ↓Vmax, Km unchanged.
- Allosteric regulation: Modulators bind non-active sites, altering activity.
- Coenzymes (vitamins) & cofactors (metal ions) are often vital.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more