PCR Basics - The DNA Photocopier
PCR: Rapid in-vitro method to amplify specific DNA segments exponentially.
- Principle: DNA replication.
- Key Components:
- DNA Template
- Primers (Fwd/Rev)
- Taq Polymerase
- dNTPs
- MgCl₂
- Buffer
- Steps (in Thermal Cycler) 📌 DAE: Don't Annoy Elephants!
- Denaturation: 94-96°C (DNA strands separate)
- Annealing: 50-65°C (Primers bind template)
- Extension: 72°C (Taq synthesizes new DNA)
- Amplification: $2^n$ copies.
⭐ Taq polymerase from Thermus aquaticus is heat-stable, vital for repeated high-temperature denaturation in PCR.
PCR Variants - Twists on a Theme
| Variant | Key Feature | Primary Use(s) |
|---|---|---|
| RT-PCR | RNA template → cDNA (via Reverse Transcriptase). | RNA viruses (HIV, Flu), gene expression analysis. |
| qPCR | Real-time DNA quantification using fluorescent probes. | Viral load, gene expression levels, pathogen quantification. |
| Nested PCR | Two primer sets (outer & inner); ↑ specificity/sensitivity. | Low copy DNA, diagnostics (TB, viral infections). |
| Multiplex PCR | Multiple DNA targets amplified in one reaction. | Pathogen panel testing, forensics, SNP genotyping. |
| ARMS-PCR | Detects known SNPs/mutations using allele-specific primers. | Genetic disease diagnosis, mutation detection (e.g., cancer). |
⭐ qPCR allows for the quantification of initial DNA/RNA amounts in real-time using fluorescent detection.
Bug Hunting with PCR - Microbe Detective
- Applications in Infectious Diseases:
- Viral: Detects HIV (viral load), HBV, HCV (genotyping), HPV, SARS-CoV-2.
- Bacterial: Identifies M. tuberculosis (GeneXpert for Rifampicin resistance), Chlamydia trachomatis.
- Fungal: Diagnoses Candida spp., Aspergillus spp. infections.
- Parasitic: Screens for Malaria (Plasmodium spp.), Leishmaniasis.
- Advantages over Traditional Methods:
- ↑ High sensitivity & specificity.
- Rapid results, often within hours.
- Crucial for non-culturable or slow-growing organisms.

⭐ PCR, like GeneXpert for TB, significantly cuts diagnostic time for Mycobacterium tuberculosis versus culture, speeding up treatment initiation.
Genes & Cancer PCR - Genetic Insights
- Genetic Disorders Diagnosis:
- Inherited diseases: Cystic Fibrosis (CFTR gene mutations), Sickle Cell Anemia (HBB gene), Thalassemias (HBA/HBB genes).
- Prenatal diagnosis using fetal DNA (amniocentesis, CVS).
- HLA typing for precise tissue matching in transplantation.
- Oncology Applications:
- Detecting oncogenes (e.g., BCR-ABL fusion in CML).
- Identifying tumor suppressor gene mutations (e.g., p53 alterations).
- Minimal Residual Disease (MRD) detection in leukemia/lymphoma.
- Microsatellite Instability (MSI) analysis (Lynch syndrome, colorectal cancer).
- Pharmacogenomics: KRAS mutations predicting therapy response.
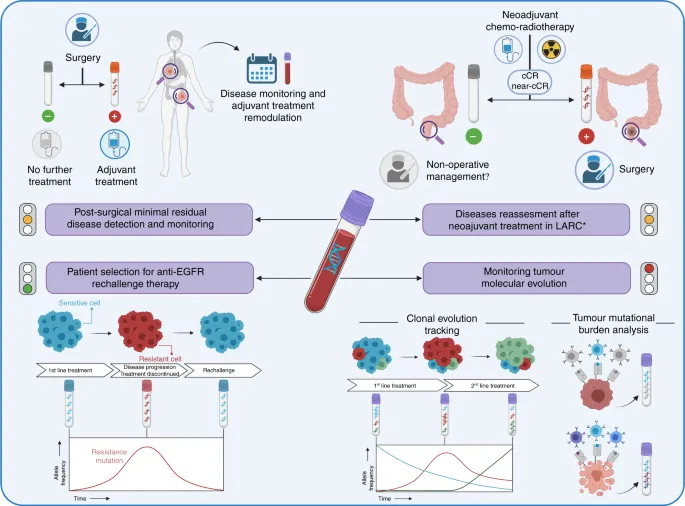
⭐ RT-PCR for BCR-ABL fusion transcript is the gold standard for diagnosing Chronic Myeloid Leukemia (CML) and monitoring treatment response.
PCR in Forensics & More - Beyond Medicine
- Forensic Applications: Vital for human identification.
- DNA fingerprinting (STRs - Short Tandem Repeats)
- Paternity testing
- Disaster victim identification
- Research Applications: Key in molecular biology.
- Gene cloning, sequencing
- Site-directed mutagenesis
- Gene expression studies (RT-qPCR)
- Other Diverse Uses: Beyond clinical settings.
- Evolutionary biology studies
- Food industry quality control (GMOs, pathogens)
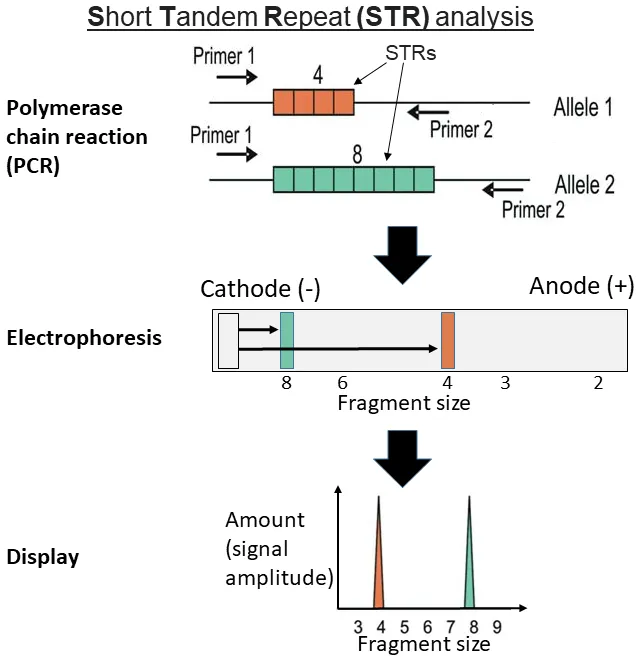
⭐ PCR amplification of Short Tandem Repeats (STRs) is the cornerstone of modern forensic DNA fingerprinting due to high variability among individuals.
High‑Yield Points - ⚡ Biggest Takeaways
- PCR rapidly detects infectious agents: HIV, HBV, HCV, HPV, TB.
- Crucial for diagnosing inherited genetic disorders like cystic fibrosis.
- Vital in cancer for oncogene/tumor suppressor gene analysis and MRD detection.
- Standard for DNA fingerprinting in forensics and paternity disputes.
- Essential for HLA typing prior to organ transplantation.
- Monitors treatment response, e.g., viral load in HIV/HCV.
- Facilitates prenatal diagnosis of chromosomal/genetic abnormalities.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

