Pharmacogenomics: Foundations - Genes & Drugs Dance
- Pharmacogenomics (PGx): How the entire genome influences drug response, guiding therapy.
- Pharmacogenetics: How specific gene variations affect individual drug response.
- Primary Goals:
- Enhance therapeutic drug efficacy (↑).
- Reduce adverse drug toxicity (↓).
- Key Genetic Variations:
- SNPs (Single Nucleotide Polymorphisms): Common single base-pair changes.
- Indels (Insertions/Deletions): Small DNA base-pair additions/removals.
- CNVs (Copy Number Variations): Altered number of copies of DNA segments.

⭐ Drug response traits can be Monogenic (e.g., TPMT variants and azathioprine toxicity) or Polygenic (e.g., warfarin dosing influenced by CYP2C9, VKORC1).
Pharmacogenomics: CYP Superstars - Metabolism Mavericks
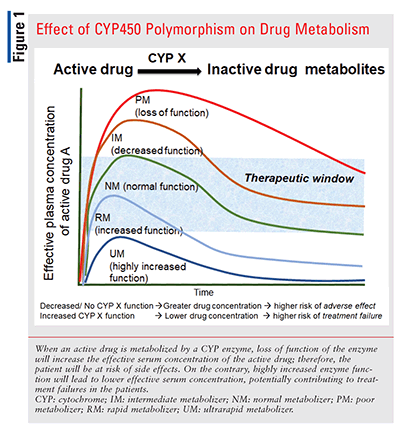
Cytochrome P450 (CYP) enzymes metabolize drugs. Genetic variants create phenotypes:
- UM: Ultrarapid Metabolizer
- EM: Extensive Metabolizer (normal)
- IM: Intermediate Metabolizer
- PM: Poor Metabolizer
Key CYP Enzymes:
| Enzyme | Drug Examples | Clinical Impact (Phenotype Dependent) |
|---|---|---|
| CYP2D6 | Codeine, Tamoxifen | UM: ↑ Codeine toxicity. PM: ↓ Codeine efficacy. |
| CYP2C19 | Clopidogrel | PM: ↓ Clopidogrel efficacy (↑ thrombosis). UM: ↑ bleeding. |
| CYP2C9 | Warfarin | PM: ↑ Warfarin sensitivity (↑ bleeding). Dose ↓ needed. |
| CYP3A4/5 | Tacrolimus, Statins | Metabolizes ~50% drugs. Variability impacts exposure. |
| %%{init: {'flowchart': {'htmlLabels': true}}}%% | ||
| flowchart TD |
Start["💊 Drug
• Pharmacotherapy• Administration"]
Genotype["🧬 CYP Genotype
• Genetic testing• Allele analysis"]
UM["⚡ UM: Fast
• Ultra-metabolizer• High enzyme activity"]
EM["✅ EM: Normal
• Extensive metab• Standard dosing"]
IM["⚠️ IM: Reduced
• Intermediate metab• Lowered activity"]
PM["🐢 PM: Slow
• Poor metabolizer• Absent activity"]
UM_Res["🚨 UM Effects
• Prodrug: ⬆️ Toxicity• Drug: ⬇️ Efficacy"]
PM_Res["🚨 PM Effects
• Prodrug: ⬇️ Efficacy• Drug: ⬆️ Toxicity"]
Start --> Genotype Genotype --> UM Genotype --> EM Genotype --> IM Genotype --> PM UM --> UM_Res PM --> PM_Res
style Start fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534 style Genotype fill:#FFF7ED, stroke:#FFEED5, stroke-width:1.5px, rx:12, ry:12, color:#C2410C style UM fill:#FEF8EC, stroke:#FBECCA, stroke-width:1.5px, rx:12, ry:12, color:#854D0E style EM fill:#F6F5F5, stroke:#E7E6E6, stroke-width:1.5px, rx:12, ry:12, color:#525252 style IM fill:#FEF8EC, stroke:#FBECCA, stroke-width:1.5px, rx:12, ry:12, color:#854D0E style PM fill:#FEF8EC, stroke:#FBECCA, stroke-width:1.5px, rx:12, ry:12, color:#854D0E style UM_Res fill:#FDF4F3, stroke:#FCE6E4, stroke-width:1.5px, rx:12, ry:12, color:#B91C1C style PM_Res fill:#FDF4F3, stroke:#FCE6E4, stroke-width:1.5px, rx:12, ry:12, color:#B91C1C
> ⭐ **CYP2D6 & Codeine:** Ultrarapid metabolizers (UMs) rapidly convert codeine to morphine, risking overdose (e.g., respiratory depression). Poor metabolizers (PMs) get poor analgesia.
## Pharmacogenomics: Beyond CYPs - Other Key Players
Many non-CYP genes also significantly impact drug response and adverse event profiles.
| Gene | Drug(s) | Key Variant(s) / Allele | Clinical Implication |
|-----------------|---------------------------------------|----------------------------------|------------------------------------------------------------|
| TPMT | Azathioprine, 6-MP, Thioguanine | Low/no activity (e.g., *2, *3A) | ↑ Myelosuppression risk; **dose reduction crucial** |
| UGT1A1 | Irinotecan | UGT1A1*28 (7 TA repeats) | ↑ Neutropenia, diarrhea; **dose reduction if homozygous** |
| SLCO1B1 | Statins (e.g., Simvastatin) | c.521T>C (rs4149056) | ↑ Statin concentration, ↑ myopathy risk |
| VKORC1 | Warfarin | -1639G>A | ↓ VKORC1 expression, **↓ warfarin dose needed** |
| HLA-B | Abacavir | HLA-B*57:01 | ↑ Hypersensitivity (HSR); **contraindicated** |
| HLA-B | Allopurinol | HLA-B*58:01 | ↑ SCARs; **test & avoid if positive** |
| HLA-B | Carbamazepine, Oxcarbazepine | HLA-B*15:02 (esp. Asian) | ↑ SJS/TEN risk; **test & avoid if positive** |> ⭐ TPMT deficiency significantly increases risk of myelosuppression with standard thiopurine doses.
## Pharmacogenomics: PGx in Practice - Clinical Hits
CPIC guidelines inform decisions. Pre-emptive/reactive testing optimizes drug choice/dose, minimizing adverse reactions, maximizing efficacy.
* **Warfarin**: *CYP2C9, VKORC1* variants guide initial dosing.
* **Clopidogrel**: *CYP2C19* LOF alleles ↓ active form; consider alternatives.
* **Abacavir**: **HLA-B\*57:01** testing **mandatory** pre-therapy (prevents severe hypersensitivity).
* **Statins**: *SLCO1B1* variants (OATP1B1) ↑ simvastatin myopathy risk.
* **Irinotecan**: *UGT1A1\*28* allele ↑ risk of severe neutropenia/diarrhea.
* **Thiopurines**: *TPMT* variants ↓ activity, ↑ myelosuppression risk (azathioprine/6-MP).
> ⭐ FDA: Carbamazepine & **HLA-B\*15:02** test in Asians (↑ SJS/TEN risk).
## High‑Yield Points - ⚡ Biggest Takeaways
>
> * **CYP2D6** variants affect **codeine** (ineffective/toxic) and **tamoxifen** efficacy.
> * **CYP2C19** loss-of-function alleles reduce **clopidogrel** activation, increasing thrombotic risk.
> * **TPMT** or **NUDT15** variants ↑ risk of **thiopurine** (azathioprine, 6-MP) toxicity.
> * **UGT1A1\*28** polymorphism (Gilbert's) ↑ **irinotecan** toxicity (neutropenia, diarrhea).
> * **HLA-B\*5701** screening is crucial before **abacavir** to prevent hypersensitivity.
> * **HLA-B\*1502** in Asians predicts **carbamazepine**-induced SJS/TEN.
> * **VKORC1** and **CYP2C9** genotypes guide **warfarin** dosing to minimize bleeding_risk_or_thrombosis_risk_if_dose_is_not_adjusted_appropriately_._
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

