Microbiome Analysis Techniques - Gut Feeling Groundwork
- Sample Collection: The Foundation
- Types: Stool (common for gut), biopsies (mucosal), saliva, skin swabs.
- Crucial: Standardized protocols, aseptic technique (avoid contamination).
- Host metadata: Record diet, medications (antibiotics!), health status for context.
- Preservation: Maintaining Fidelity
- Goal: Prevent microbial changes & nucleic acid degradation.
- Immediate stabilization:
- Freezing: Snap-freeze (liquid $N_2$), store -80°C.
- Chemical stabilizers: RNAlater, OMNIgene•GUT kits, ethanol (field-friendly).
- Storage: Long-term at -80°C (gold standard).
- Transport: Strict cold chain (dry ice).
- 📌 Mnemonic: Sample Carefully, Stabilize Swiftly, Store Securely! (SCSS)
⭐ Fresh stool samples (<24h at room temp, or immediately frozen/stabilized) are vital for accurate gut microbiome in vivo representation.
Microbiome Analysis Techniques - Decoding Diversity
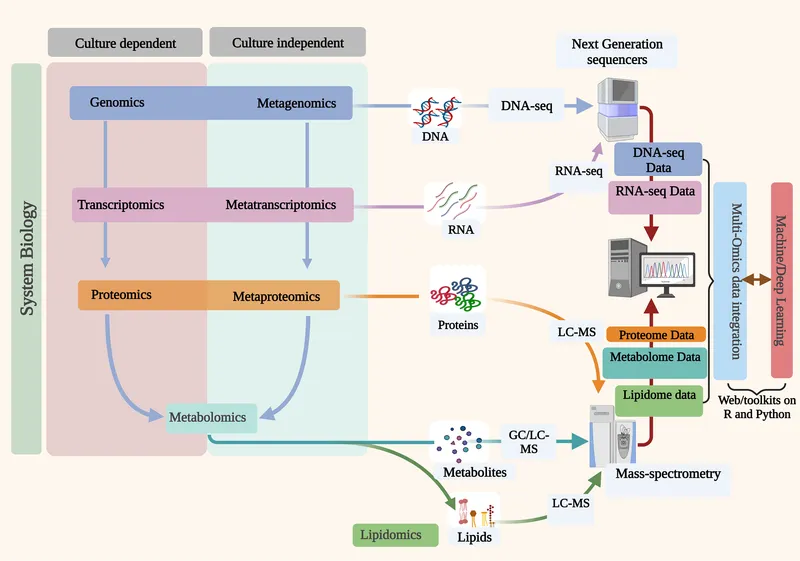
- Culture-Independent Methods: For unculturable microbes (Viable But Not Culturable - VBNC). 📌 Amplicons Spot, Shotgun Scans (Functions).
- Amplicon Sequencing (Targeted): Cost-effective for "who is there?".
- 16S rRNA gene: For bacteria & archaea. Targets hypervariable regions (e.g., V3-V4).
- ITS (Internal Transcribed Spacer): For fungi.
- Process: DNA extraction → PCR of marker gene → Sequencing → OTU/ASV clustering → Taxonomic assignment.
- Diversity Analysis:
- Alpha diversity (within-sample): Richness (Chao1), evenness (Shannon, Simpson).
- Beta diversity (between-samples): Compares composition (Bray-Curtis, Jaccard, UniFrac). Visualized by PCoA.
- Shotgun Metagenomics (WGS): Unbiased sequencing of all microbial DNA.
- Reveals:
- Taxonomic composition (bacteria, archaea, viruses, fungi, protists).
- Functional potential (metabolic pathways, virulence factors, resistance genes).
- Higher resolution & broader scope than amplicon sequencing; more expensive.
- Reveals:
- Functional 'Omics (What are they doing?):
- Metatranscriptomics (RNA): Active gene expression.
- Metaproteomics (Proteins): Expressed proteins, direct functional evidence.
- Metabolomics (Metabolites): Small molecule products, host-microbe chemical crosstalk.
- Amplicon Sequencing (Targeted): Cost-effective for "who is there?".
⭐ Alpha diversity (e.g., Shannon) indicates richness/evenness within a sample. Beta diversity (e.g., Bray-Curtis) compares composition differences between samples.
Microbiome Analysis Techniques - Interpreting Interactions
- Goal: Decipher community structure, function, microbial interactions (synergism/antagonism), and host-microbe interplay from processed data.
- Core Analyses:
- Diversity Metrics:
- Alpha (α): Intra-sample richness (Chao1) & evenness (Shannon). High α = ↑diversity.
- Beta (β): Inter-sample dissimilarity (Bray-Curtis, UniFrac). Visualized: PCoA, NMDS.
- Comparative Analysis:
- Differential Abundance: Identifies taxa (OTUs/ASVs) varying between groups (LEfSe, DESeq2). Key for biomarkers.
- Functional Insights:
- Functional Profiling: Predicts gene functions & pathways (PICRUSt2 - 16S; HUMAnN3 - metagenomes).
- Interaction & Dynamics:
- Network Analysis: Infers co-occurrence/co-exclusion, suggesting microbial interactions.
- Longitudinal Studies: Tracks community shifts over time.
- Multi-omics Integration: Combines with host data (transcriptomics, metabolomics) for systems view.
- Diversity Metrics:
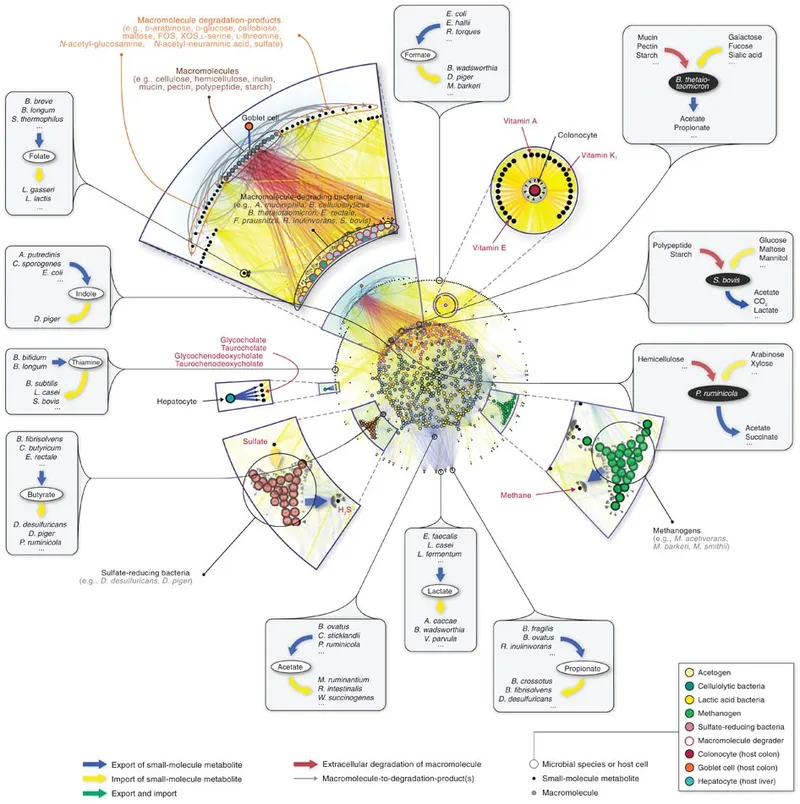
⭐ UniFrac distance is a key beta diversity metric that incorporates phylogenetic information, crucial for comparing relatedness of microbial communities across different samples or conditions based on evolutionary divergence between taxa present in them.
High‑Yield Points - ⚡ Biggest Takeaways
- 16S rRNA sequencing: Key for taxonomic profiling (who is there?), targets variable gene regions.
- Shotgun metagenomics: Sequences all genomic DNA, reveals functional potential & species-level identification.
- Metatranscriptomics (RNA-Seq): Shows active gene expression and community function.
- Metaproteomics: Identifies proteins, confirming functional activity of the microbiome.
- Metabolomics: Analyzes small molecule metabolites, indicating microbial metabolic output and host interactions.
- Alpha diversity: Measures species richness/evenness within a single sample (e.g., Shannon index).
- Beta diversity: Compares microbial composition between different samples (e.g., Bray-Curtis, PCoA).
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

