Genetic Code & Ribosomes - Code Crackers Crew
- Genetic Code: Triplet, non-overlapping, commaless, degenerate, nearly universal.
- 64 codons: 61 sense (code for amino acids), 3 nonsense (stop).
- Start codon: AUG (Methionine; rarely GUG).
- Stop codons: UAA, UAG, UGA. 📌 Mnemonic: "U Are Away, U Are Gone, U Go Away".
- Ribosomes: rRNA + protein; site of translation.
- Prokaryotic: 70S (50S + 30S subunits).
- Eukaryotic: 80S (60S + 40S subunits).
- A, P, E sites for tRNA binding.
- mRNA: Carries genetic information from DNA as codons.
- tRNA: Adaptor molecule; anticodon pairs with mRNA codon, carries specific amino acid.
⭐ The peptidyl transferase activity, forming peptide bonds, is a ribozyme function of the large ribosomal subunit's rRNA (23S in prokaryotes, 28S in eukaryotes).
Translation Steps - Protein Assembly Line
- Initiation:
-
- Small subunit (30S/40S) + mRNA + initiator tRNA (fMet/Met) + IFs → complex.
-
- Prok: Shine-Dalgarno. Euk: 5' cap, Kozak seq.
-
- Large subunit (50S/60S) joins; GTP used.
-
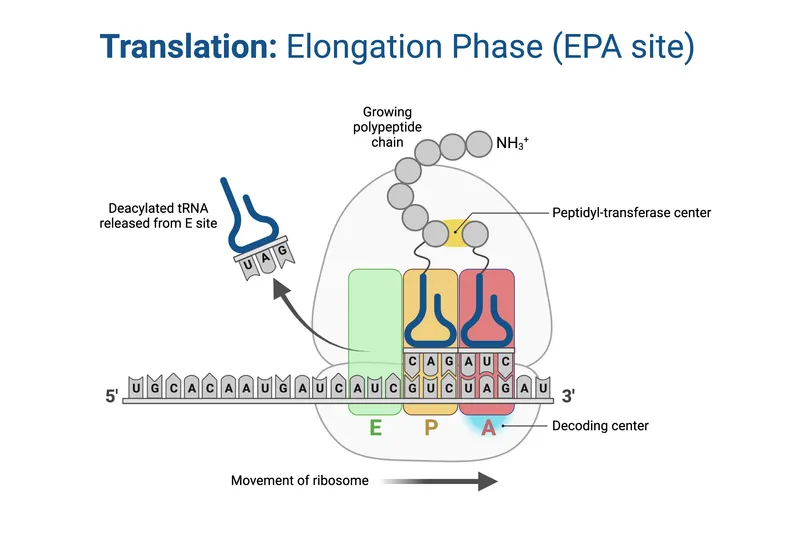
- Elongation: (📌 APE sites: Aminoacyl, Peptidyl, Exit)
-
- Codon Recognition: aa-tRNA to A-site (EF-Tu/eEF1A, GTP).
-
- Peptide Bond: Peptidyl transferase (rRNA ribozyme) links AAs. Peptide to A-tRNA.
-
- Translocation: Ribosome moves 1 codon (EF-G/eEF2, GTP). P→E, A→P.
-
- Termination:
-
- Stop codon (UAA, UAG, UGA) in A-site.
-
- Release Factors (RFs/eRFs) bind.
-
- Polypeptide released; complex dissociates (GTP).
-
⭐ Peptidyl transferase is a ribozyme (catalytic rRNA: 23S in prokaryotes, 28S in eukaryotes).
Translation Control & Inhibitors - Quality & Sabotage
- Control:
- Initiation: Key regulatory step. eIF2-P (stress, ↓ global), eIF4E (cap-binding, ↑ cancer). IRES (cap-independent translation).
- mRNA stability (e.g., AU-rich elements affecting decay).
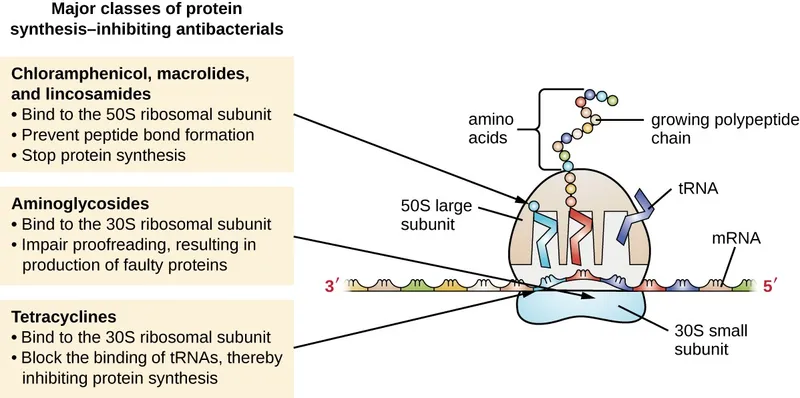
- Prokaryotic Inhibitors (70S): 📌 "Buy AT 30, CELL at 50"
- 30S: Aminoglycosides (misreading mRNA), Tetracyclines (block A-site tRNA entry).
- 50S: Macrolides (translocation), Chloramphenicol (peptidyltransferase), Clindamycin (peptide bond), Linezolid (prevents 70S initiation complex).
- Eukaryotic Inhibitors (80S):
- Diphtheria toxin: ADP-ribosylates eEF-2, halting elongation.
- Ricin (plant toxin): Cleaves rRNA in 60S subunit.
- Quality/Sabotage:
- NMD: Degrades mRNA with PTCs (Premature Termination Codons), preventing truncated proteins.
- Viral shutoff: Poliovirus protease cleaves eIF4G.
⭐ Diphtheria toxin inactivates eukaryotic elongation factor 2 (eEF-2) via ADP-ribosylation, stopping protein synthesis.
Post-Translational Modifications - Protein Finishing Touches
- Post-synthesis changes; vital for protein folding, stability, localization, activity.
- Common Modifications:
- Phosphorylation: Ser/Thr/Tyr. Kinases add, phosphatases remove. Regulates signaling, enzyme activity.
- Glycosylation: N-linked (Asn, ER), O-linked (Ser/Thr, Golgi). Folding, targeting, recognition.
⭐ Mannose-6-phosphate (M6P) targets enzymes to lysosomes; M6P pathway defect causes I-cell disease.
- Ubiquitination: Lys. Tags proteins for proteasomal degradation.
- Hydroxylation: Pro, Lys in collagen. Needs Vit C (📌 Collagen C needs Vit C). Deficiency: Scurvy.
- Carboxylation: Glu in clotting factors. Needs Vit K (📌 K for Koagulation).
- Acetylation/Methylation: Lys, Arg in histones. Regulates gene expression.
- Proteolytic Cleavage: Zymogen activation (trypsinogen), signal peptide removal.

High‑Yield Points - ⚡ Biggest Takeaways
- Wobble hypothesis: Inosine in tRNA anticodon allows binding multiple codons, explaining code degeneracy.
- Antibiotics: Streptomycin (30S) causes mRNA misreading; Chloramphenicol (50S) inhibits peptidyl transferase.
- Puromycin: Aminoacyl-tRNA analog, causes premature termination in prokaryotes and eukaryotes.
- Protein targeting: Signal Recognition Particle (SRP) directs proteins to ER for N-linked glycosylation; O-linked glycosylation occurs in Golgi.
- Modifications: Ubiquitination targets proteins for proteasomal degradation; SUMOylation alters function/localization.
- Protein folding: Assisted by chaperones (e.g., Hsp70); misfolding is linked to diseases like Alzheimer's.
- Kozak sequence (eukaryotes) or Shine-Dalgarno sequence (prokaryotes) initiates translation at the start codon.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

