Infectious Diseases

On this page
🦠 Infectious Disease Mastery: Your Clinical Command Center
Infectious diseases remain among medicine's most dynamic challenges, where pathogens evolve, immune systems mount complex defenses, and clinical decisions can mean the difference between recovery and catastrophe. You'll master how to recognize infection patterns, interpret diagnostic data strategically, deploy antimicrobials with precision, and anticipate resistance before it undermines treatment. This lesson builds your clinical command from immune mechanisms through laboratory interpretation to therapeutic decision-making, equipping you with frameworks that transform uncertainty into confident action at the bedside.

Infectious diseases represent the dynamic battlefield between microbial invaders and human host defenses. Understanding this complex interplay requires mastering four core domains: pathogen characteristics, host immune responses, clinical manifestations, and therapeutic interventions. Each domain interconnects to create the clinical presentations you'll encounter in practice.
📌 Remember: SHIP - Susceptible host, Host factors, Infectious agent, Pathway of transmission. These four elements must align for infection to occur, with >85% of healthcare-associated infections preventable through understanding transmission pathways.
The infectious disease spectrum spans from localized superficial infections affecting single organ systems to systemic sepsis with >40% mortality in severe cases. Recognition patterns depend on understanding pathogen virulence factors, host immune status, and the time-dependent progression from colonization to invasive disease.
Pathogen Classification Framework
- Bacterial Pathogens
- Gram-positive cocci: S. aureus (25-30% of skin/soft tissue infections)
- Gram-negative rods: E. coli (80% of uncomplicated UTIs)
- Enterobacteriaceae: >50% of hospital-acquired infections
- Non-fermenters: P. aeruginosa with 15-25% mortality in bacteremia
- Viral Pathogens
- DNA viruses: HSV reactivation in 60-90% of immunocompromised patients
- RNA viruses: Influenza with 3-5 million severe cases annually
- Respiratory syncytial virus: >125,000 hospitalizations yearly
- SARS-CoV-2: Case fatality rate 0.5-3% depending on demographics
- Fungal Pathogens
- Yeasts: Candida species causing >400,000 life-threatening infections
- Molds: Aspergillus with >50% mortality in invasive disease
⭐ Clinical Pearl: Pathogen identification within 6-8 hours using rapid molecular diagnostics reduces mortality by 15-20% compared to traditional culture methods requiring 24-48 hours.
| Pathogen Type | Incubation Period | Diagnostic Window | Mortality Risk | Key Virulence Factor |
|---|---|---|---|---|
| Gram+ Bacteria | 1-3 days | 6-24 hours | 10-25% | Toxin production |
| Gram- Bacteria | 1-7 days | 6-24 hours | 15-40% | Endotoxin/biofilms |
| DNA Viruses | 2-14 days | 1-3 days | 5-15% | Latency/reactivation |
| RNA Viruses | 1-10 days | 1-2 days | 1-10% | Rapid mutation |
| Yeasts | 1-5 days | 12-48 hours | 20-50% | Biofilm formation |
| Molds | 5-21 days | 24-72 hours | 30-70% | Tissue invasion |
Connect these foundational pathogen characteristics through host immune response patterns to understand how clinical syndromes develop and progress.
🦠 Infectious Disease Mastery: Your Clinical Command Center
⚔️ Host Defense Architecture: The Immune Battleground
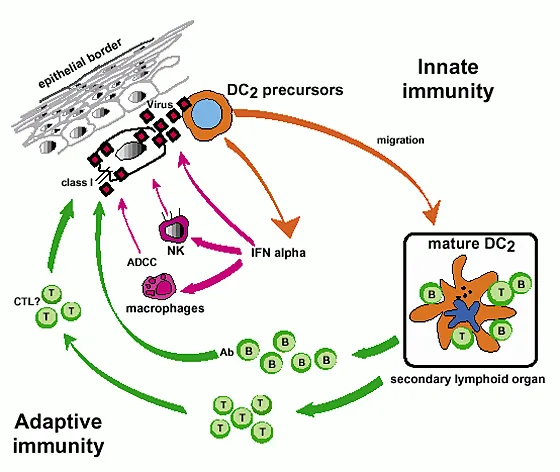
Host defense operates through three integrated tiers: physical barriers, innate immunity, and adaptive immunity. Each tier activates within specific timeframes, creating overlapping protection with >99.9% effectiveness against most environmental pathogens in healthy individuals.
📌 Remember: PINA - Physical barriers (0-4 hours), Innate immunity (4-96 hours), Neutrophil response (6-24 hours), Adaptive immunity (96+ hours). This temporal sequence determines clinical presentation timing and therapeutic windows.
Immune Response Mechanisms
- Physical Barrier Defense
- Skin integrity: <2% infection rate with intact barriers
- Mucosal surfaces: >90% pathogen clearance via ciliary action
- Gastric acid: pH <2 eliminates >99% ingested bacteria
- Respiratory mucus: >10^6 bacteria/day cleared normally
- Innate Immune Activation
- Neutrophil recruitment: Peak response 6-12 hours post-exposure
- Macrophage activation: >100-fold increase in phagocytic capacity
- Complement cascade: <30 seconds for C5a generation
- Cytokine release: TNF-α peaks 2-4 hours, IL-6 peaks 4-8 hours
- Adaptive Immune Response
- T-cell activation: 72-96 hours for clonal expansion
- Antibody production: Primary response 7-14 days, secondary 2-5 days
- Memory formation: >90% protection against reinfection
- Cross-reactivity: 10-30% effectiveness against variant strains
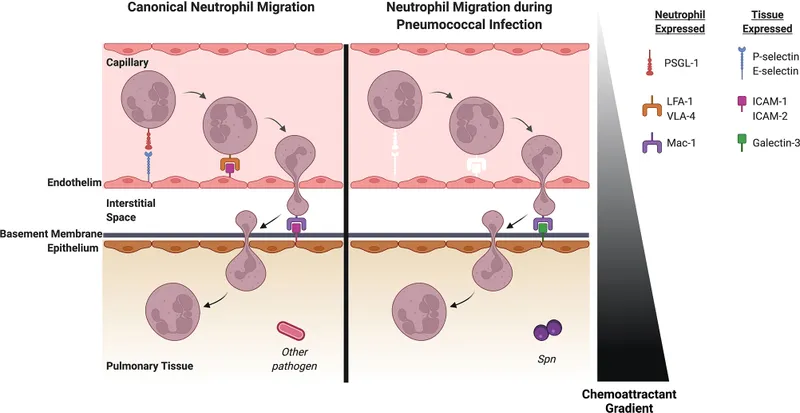
⭐ Clinical Pearl: Neutrophil count <500 cells/μL increases infection risk >100-fold, with >80% of febrile neutropenic patients developing serious bacterial infections within 48-72 hours without empirical antibiotics.
| Immune Component | Response Time | Peak Activity | Duration | Clinical Marker |
|---|---|---|---|---|
| Complement | <1 minute | 5-15 minutes | 2-4 hours | C3, C4 levels |
| Neutrophils | 2-6 hours | 6-24 hours | 2-5 days | ANC, left shift |
| Monocytes | 6-12 hours | 24-72 hours | 5-14 days | Monocyte count |
| T-cells | 72-96 hours | 7-14 days | Weeks-years | CD4/CD8 ratio |
| B-cells | 5-7 days | 14-21 days | Months-years | Immunoglobulins |
| Memory cells | 2-5 days | 5-10 days | Years-lifetime | Specific antibodies |
Connect immune response timing through clinical syndrome recognition to understand how different pathogens exploit specific host vulnerabilities.
⚔️ Host Defense Architecture: The Immune Battleground
🎯 Clinical Pattern Recognition: The Diagnostic Framework
📌 Remember: FEVER - Focal signs, Epidemiologic clues, Vital sign patterns, Exposure history, Risk factors. This systematic approach identifies >85% of infectious syndromes within first 10 minutes of clinical assessment.
Syndrome-Based Recognition Patterns
- Respiratory Tract Infections
- Upper respiratory: >90% viral etiology, self-limited 7-10 days
- Community-acquired pneumonia: S. pneumoniae 30-40%, atypicals 15-25%
- Severity scoring: PSI >130 or CURB-65 ≥3 indicates hospitalization
- Mortality risk: <1% outpatient, 5-15% hospitalized, >30% ICU
- Urinary Tract Infections
- Uncomplicated cystitis: E. coli 75-85%, S. saprophyticus 10-15%
- Complicated UTI: Enterobacteriaceae 60%, Enterococcus 15%, P. aeruginosa 10%
- Bacteremia risk: <5% cystitis, 15-25% pyelonephritis
- Resistance patterns: >25% E. coli resistance to fluoroquinolones
- Skin and Soft Tissue Infections
- Cellulitis: S. pyogenes 40%, S. aureus 35%, mixed 25%
- Necrotizing fasciitis: Mortality >30% without surgical debridement
- LRINEC score: >6 suggests necrotizing infection
- Time-critical: <6 hours to surgery improves survival
⭐ Clinical Pearl: Fever pattern analysis provides diagnostic clues: continuous fever suggests gram-negative bacteremia, intermittent fever indicates abscess or malaria, relapsing fever suggests Borrelia or Brucella infection.
| Clinical Syndrome | Most Common Pathogen | Diagnostic Test | Time to Result | Empirical Therapy |
|---|---|---|---|---|
| CAP (outpatient) | S. pneumoniae (35%) | Chest X-ray | <1 hour | Amoxicillin |
| CAP (hospitalized) | S. pneumoniae (30%) | Blood cultures | 24-48 hours | Ceftriaxone + azithromycin |
| Acute cystitis | E. coli (80%) | Urinalysis | <1 hour | Nitrofurantoin |
| Cellulitis | S. pyogenes (40%) | Clinical diagnosis | Immediate | Cephalexin |
| Meningitis | S. pneumoniae (50%) | CSF analysis | 2-4 hours | Ceftriaxone + vancomycin |
| Sepsis | Mixed (variable) | Blood cultures | 24-48 hours | Piperacillin-tazobactam |
Connect pattern recognition skills through systematic diagnostic approaches to understand how laboratory and imaging studies confirm clinical suspicions.
🎯 Clinical Pattern Recognition: The Diagnostic Framework
🔬 Diagnostic Precision: Laboratory Intelligence Network

Diagnostic testing strategy follows three-tiered approach: rapid screening tests, confirmatory identification, and antimicrobial susceptibility testing. Each tier provides specific information within defined timeframes, enabling progressive therapeutic refinement from empirical to targeted therapy.
📌 Remember: RAPID - Rapid antigen/PCR (15 minutes-2 hours), Automated blood cultures (6-24 hours), Pathogen identification (12-24 hours), Identification confirmation (24-48 hours), Drug susceptibility (24-72 hours). This timeline guides therapeutic decision-making.
Modern Diagnostic Technologies
- Molecular Diagnostics
- PCR-based assays: Results in 1-4 hours, sensitivity >95%
- Multiplex panels: >20 pathogens simultaneously, >90% concordance
- Respiratory panel: Identifies 15-25 viruses/bacteria in <2 hours
- GI panel: >20 enteric pathogens with >95% sensitivity
- Rapid Antigen Detection
- Point-of-care testing: Results in 15-30 minutes
- Influenza rapid tests: Sensitivity 50-70%, specificity >95%
- Strep throat: Sensitivity 85-95%, specificity >98%
- COVID-19 antigen: Sensitivity 85% in symptomatic patients
- Automated Blood Culture Systems
- Time to positivity: Mean 12-18 hours for common bacteria
- MALDI-TOF identification: Results in 2-4 hours post-positivity
- Antimicrobial susceptibility: Additional 18-24 hours
- Resistance gene detection: PCR results in 1-2 hours
⭐ Clinical Pearl: Procalcitonin levels distinguish bacterial from viral infections: <0.25 ng/mL suggests viral, >0.5 ng/mL indicates bacterial infection with >80% accuracy. Levels >2.0 ng/mL correlate with severe bacterial infection requiring immediate antibiotics.
| Diagnostic Method | Time to Result | Sensitivity | Specificity | Cost Factor | Clinical Application |
|---|---|---|---|---|---|
| Rapid antigen | 15-30 minutes | 70-90% | 95-99% | Low | Point-of-care screening |
| PCR (single) | 1-4 hours | 95-99% | 98-99% | Moderate | Targeted pathogen detection |
| Multiplex PCR | 2-6 hours | 90-95% | 95-98% | High | Syndrome-based testing |
| Blood culture | 24-72 hours | 85-95% | >99% | Moderate | Bacteremia/fungemia |
| MALDI-TOF | 2-4 hours* | 95-99% | 98-99% | Low* | Rapid organism ID |
| Susceptibility | 18-48 hours* | 95-98% | 98-99% | Low* | Antimicrobial guidance |
💡 Master This: Diagnostic stewardship requires understanding test limitations and appropriate utilization. Multiplex PCR panels detect colonizing organisms alongside pathogens, requiring clinical correlation. Positive predictive value depends on disease prevalence - low-prevalence settings increase false-positive rates.
Connect diagnostic precision through evidence-based treatment algorithms to understand how laboratory results guide antimicrobial selection and monitoring.
🔬 Diagnostic Precision: Laboratory Intelligence Network
💊 Therapeutic Precision: The Antimicrobial Arsenal
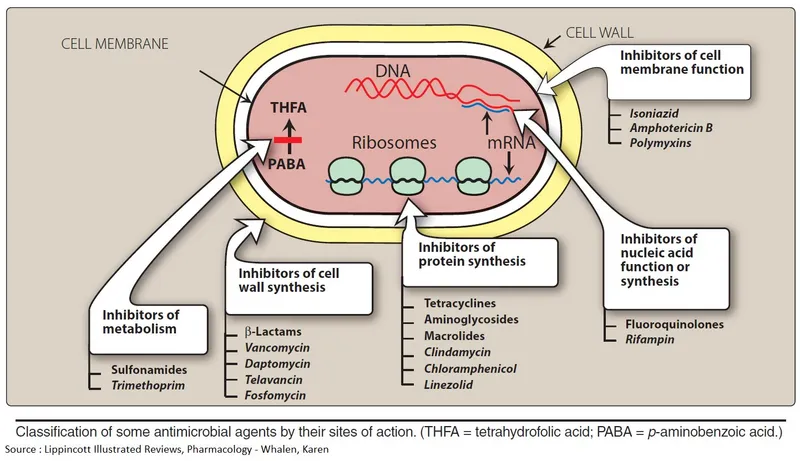
Antimicrobial selection follows four core principles: spectrum appropriateness, pharmacokinetic optimization, resistance prevention, and adverse effect minimization. Master clinicians integrate these principles to achieve maximum therapeutic benefit while preserving antimicrobial effectiveness for future patients.
📌 Remember: SMART - Spectrum (narrow when possible), Minimum inhibitory concentration, Allergy history, Resistance patterns, Tissue penetration. This framework guides >95% of antimicrobial decisions with optimal outcomes.
Antimicrobial Classification and Selection
- Beta-lactam Antibiotics
- Penicillins: Time-dependent killing, >50% time above MIC required
- Cephalosporins: Generation-specific spectrum, 1st-4th generation progression
- Ceftriaxone: CNS penetration 10-15%, half-life 8 hours
- Cefepime: Anti-pseudomonal activity, >90% susceptibility in most regions
- Fluoroquinolones
- Concentration-dependent killing: AUC/MIC >125 for optimal efficacy
- Resistance development: >25% E. coli resistance in many regions
- Levofloxacin: Respiratory penetration >90%, QTc prolongation risk
- Ciprofloxacin: Urinary concentration >100x serum, tendon rupture <0.1%
- Glycopeptides and Lipopeptides
- Vancomycin: Trough levels 15-20 mg/L for serious MRSA infections
- Daptomycin: Dose 6-12 mg/kg, CPK monitoring for myopathy
- Linezolid: 100% oral bioavailability, thrombocytopenia >10 days
- Tedizolid: Once-daily dosing, reduced hematologic toxicity

⭐ Clinical Pearl: Antimicrobial de-escalation within 48-72 hours based on culture results reduces resistance development by >30% and decreases adverse effects by >25% without compromising clinical outcomes.
| Antimicrobial Class | Mechanism | PK/PD Target | Common Resistance | Monitoring Parameter |
|---|---|---|---|---|
| Beta-lactams | Cell wall synthesis | Time > MIC (50%) | Beta-lactamases | Renal function |
| Fluoroquinolones | DNA gyrase | AUC/MIC (>125) | Target mutations | QTc interval |
| Aminoglycosides | Protein synthesis | Peak/MIC (8-10x) | Enzymatic modification | Creatinine, hearing |
| Glycopeptides | Cell wall synthesis | AUC/MIC (>400) | Target alteration | Trough levels |
| Macrolides | Protein synthesis | Time > MIC (40%) | Efflux pumps | Liver enzymes |
| Tetracyclines | Protein synthesis | AUC/MIC (>25) | Efflux/ribosomal | Photosensitivity |
Connect therapeutic precision through advanced integration concepts to understand how antimicrobial resistance patterns and emerging threats shape clinical practice.
💊 Therapeutic Precision: The Antimicrobial Arsenal
🌐 Resistance Networks: The Evolutionary Battlefield

Resistance development follows predictable evolutionary pathways driven by selective pressure, horizontal gene transfer, and clonal expansion. Understanding these mechanisms enables proactive resistance prevention and guides empirical therapy selection in high-resistance environments.
📌 Remember: ESCAPE pathogens - Enterococcus faecium, Staphylococcus aureus, Clostridium difficile, Acinetobacter baumannii, Pseudomonas aeruginosa, Enterobacteriaceae. These organisms account for >70% of healthcare-associated infections and demonstrate multiple resistance mechanisms.
Resistance Mechanism Categories
- Enzymatic Inactivation
- Beta-lactamases: >1,000 variants identified, TEM, SHV, CTX-M families
- Extended-spectrum beta-lactamases (ESBLs): >30% E. coli prevalence globally
- Carbapenemases: KPC, NDM, OXA-48 families, >90% mortality untreated
- Aminoglycoside-modifying enzymes: >50 variants, cross-resistance common
- Target Site Modification
- Penicillin-binding protein alterations: MRSA prevalence 20-50% in hospitals
- DNA gyrase mutations: >25% fluoroquinolone resistance in E. coli
- Ribosomal modifications: Macrolide resistance >40% in S. pneumoniae
- Vancomycin resistance: VanA, VanB operons, >95% resistance levels
- Efflux Pump Overexpression
- Multidrug efflux systems: >12 families identified in bacteria
- P. aeruginosa: >4 major pumps, intrinsic resistance to multiple classes
- Acinetobacter: AdeABC system, pan-drug resistance potential
- Enterobacteriaceae: AcrAB-TolC, fluoroquinolone/beta-lactam resistance
⭐ Clinical Pearl: Carbapenem-resistant Enterobacteriaceae (CRE) infections carry >50% mortality rates. Early recognition through rapid carbapenemase detection within 4-6 hours enables appropriate therapy selection and infection control measures.
| Resistance Mechanism | Time to Develop | Genetic Basis | Transferability | Clinical Impact |
|---|---|---|---|---|
| Point mutations | Days-weeks | Chromosomal | Low | Moderate |
| Plasmid acquisition | Hours-days | Extrachromosomal | High | Severe |
| Transposon insertion | Hours-days | Mobile elements | High | Severe |
| Efflux upregulation | Hours-days | Regulatory | Moderate | Moderate |
| Enzyme production | Hours-days | Plasmid/chromosome | High | Severe |
| Target modification | Days-weeks | Chromosomal | Low-moderate | Moderate-severe |
Connect resistance network understanding through clinical mastery frameworks to develop practical tools for immediate clinical application and long-term antimicrobial stewardship.
🌐 Resistance Networks: The Evolutionary Battlefield
🎯 Clinical Mastery Arsenal: Rapid Decision Tools
📌 Remember: MASTER approach - Microorganism identification, Antimicrobial selection, Source control, Timing optimization, Escalation/de-escalation, Resistance prevention. This framework guides >95% of infectious disease management decisions.
Essential Clinical Arsenal
- Rapid Assessment Tools
- qSOFA score: ≥2 points indicates >10-fold increased mortality risk
- SIRS criteria: ≥2 criteria present in >90% of sepsis cases
- Sepsis-3 definition: SOFA score increase ≥2 with suspected infection
- Septic shock: Lactate >2 mmol/L despite adequate fluid resuscitation
- Empirical Therapy Guidelines
- Community-acquired infections: Local resistance patterns guide selection
- Healthcare-associated infections: Broad-spectrum coverage until cultures available
- Immunocompromised patients: Anti-pseudomonal coverage empirically
- Critically ill patients: Combination therapy for severe infections
- Monitoring Parameters
- Clinical response: Fever resolution 48-72 hours, WBC normalization
- Biomarker trends: Procalcitonin decrease >80% indicates effective therapy
- Culture clearance: Negative cultures 48-72 hours post-treatment
- Resistance surveillance: Monthly antibiograms guide empirical choices
⭐ Clinical Pearl: Source control within 6-12 hours is critical for >80% of intra-abdominal infections and >90% of necrotizing soft tissue infections. Delayed source control increases mortality exponentially with each hour of delay.
| Clinical Scenario | Time to Antibiotics | Empirical Choice | Duration | Success Rate |
|---|---|---|---|---|
| Septic shock | <1 hour | Broad-spectrum | 7-10 days | 60-80% |
| Meningitis | <30 minutes | Ceftriaxone + vancomycin | 10-14 days | 85-95% |
| Necrotizing fasciitis | <1 hour | Clindamycin + penicillin | Until debridement | 70-90% |
| Febrile neutropenia | <1 hour | Anti-pseudomonal | Until ANC >500 | 80-90% |
| Healthcare pneumonia | <4 hours | Anti-MRSA + anti-pseudomonal | 7-8 days | 75-85% |
Master these frameworks, and you possess the clinical tools necessary for expert infectious disease management across all practice settings, from community clinics to intensive care units.
🎯 Clinical Mastery Arsenal: Rapid Decision Tools
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app













