Protein Structure and Function

On this page
🏗️ Protein Architecture: The Molecular Blueprint Mastery
A master architect reviews 2,847 structural blueprints in a single skyscraper project, ensuring each beam placement achieves millimeter precision across 40+ floors. Similarly, protein structure demands understanding how 20 amino acids create over 100,000 distinct proteins through precise architectural principles that determine every biological function.
Protein structure represents the most sophisticated molecular architecture in biology, where sequence dictates shape, and shape determines destiny. Every enzyme's catalytic power, every antibody's specificity, and every hormone's signaling capacity emerges from structural precision that rivals the finest engineering marvels.
📌 Remember: PSTQ - Primary (sequence), Secondary (local folds), Tertiary (3D shape), Quaternary (subunit assembly) - Each level builds upon the previous with exponential complexity increase
The structural hierarchy follows thermodynamic principles where primary structure (amino acid sequence) contains all information needed for proper folding, yet requires molecular chaperones in 30-50% of cases to achieve native conformation. This relationship between sequence and structure forms the foundation for understanding protein function, dysfunction, and therapeutic intervention.
-
Primary Structure Foundation
- Linear amino acid sequence connected by peptide bonds
- Contains 100% of folding information (Anfinsen's principle)
- Single amino acid changes can alter function by >90%
- Sickle cell: Glu→Val at position 6 in β-globin
- Phenylketonuria: Phe→Ser at position 408 in phenylalanine hydroxylase
- Cystic fibrosis: ΔPhe508 deletion in CFTR protein
-
Secondary Structure Patterns
- α-helices: 3.6 amino acids per turn, 1.5 Å rise per residue
- β-sheets: Extended conformation with 3.5 Å spacing between residues
- Turns and loops: Connect secondary elements, comprise 30-40% of protein structure
- β-turns: 4 amino acid segments with 180° direction change
- Random coils: Flexible regions allowing conformational changes
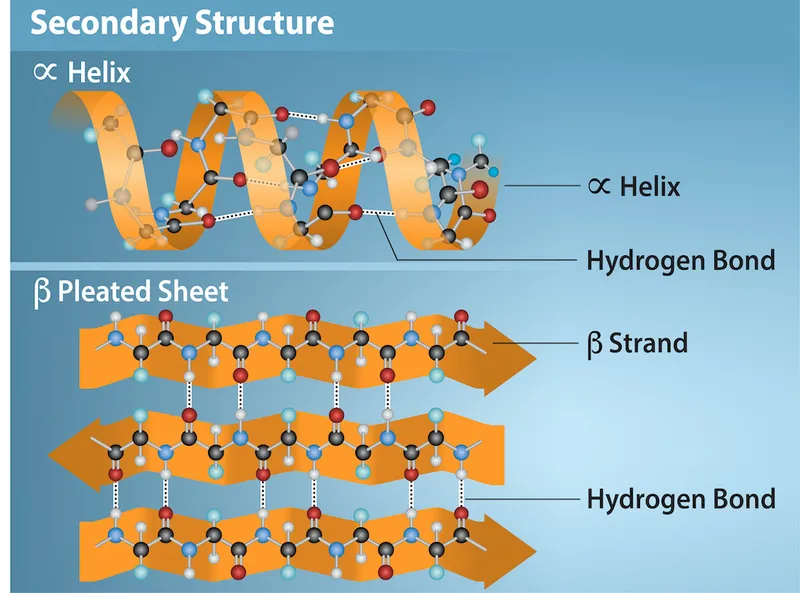
⭐ Clinical Pearl: Prion diseases result from α-helix to β-sheet conversion in PrP protein, demonstrating how secondary structure changes cause fatal neurodegenerative disorders with 100% mortality rate
| Structure Level | Stabilizing Forces | Energy (kcal/mol) | Clinical Examples | Disruption Effects |
|---|---|---|---|---|
| Primary | Covalent peptide bonds | 80-120 | Sickle cell anemia | Complete loss of function |
| Secondary | Hydrogen bonds | 2-5 per bond | Alzheimer's amyloid | Aggregation, toxicity |
| Tertiary | Multiple weak forces | 1-15 total | Cystic fibrosis | Misfolding, degradation |
| Quaternary | Subunit interactions | 5-20 | Hemoglobinopathies | Altered cooperativity |
| Supersecondary | Motif combinations | 10-30 | Immunodeficiencies | Domain dysfunction |
Understanding protein architecture transforms clinical reasoning by revealing how single nucleotide polymorphisms can cascade through structural levels to produce disease phenotypes. This knowledge predicts drug binding sites, explains resistance mechanisms, and guides therapeutic design strategies.
🏗️ Protein Architecture: The Molecular Blueprint Mastery
⚙️ Folding Dynamics: The Conformational Energy Landscape
A Formula 1 engineer calculates 15,000+ aerodynamic variables during a 2-second pit stop, optimizing performance through microsecond precision. Protein folding navigates an equally complex energy landscape, sampling millions of conformations in microseconds to find the single native state among astronomical possibilities.
Protein folding represents one of biology's most elegant solutions to the Levinthal paradox - how proteins find their native conformation in seconds to minutes rather than the billions of years required for random search. The folding process follows specific pathways guided by kinetic intermediates and thermodynamic stability.
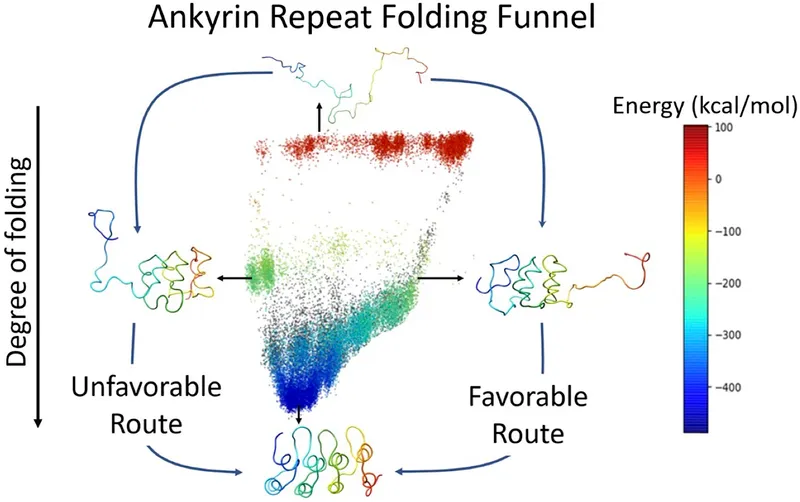
📌 Remember: HIKE - Hydrophobic collapse (first), Intermediate formation, Kinetic traps avoided, Energy minimum reached - Folding follows hierarchical assembly with hydrophobic forces driving initial collapse in <1 millisecond
The folding energy landscape resembles a funnel where entropy decreases and enthalpy becomes favorable as the protein approaches its native state. Molecular chaperones smooth this landscape by preventing kinetic traps and aggregation, essential for 25-30% of cellular proteins.
-
Folding Kinetics and Thermodynamics
- Folding rates: 10^-6 to 10^3 seconds depending on protein size
- Stability margins: 5-15 kcal/mol between native and unfolded states
- Cooperativity: Most proteins show two-state folding transitions
- Small proteins (<100 residues): Single-step folding
- Large proteins (>150 residues): Multi-step with intermediates
- Multi-domain proteins: Independent domain folding
-
Chaperone-Assisted Folding
- Hsp70 system: Binds hydrophobic segments, prevents aggregation
- GroEL/GroES: Anfinsen cage for isolated folding of 30-60 kDa proteins
- Hsp90: Late-stage folding assistance for signaling proteins
- ATP consumption: 2-4 molecules per folding cycle
- Success rate: 70-90% for chaperone-dependent proteins
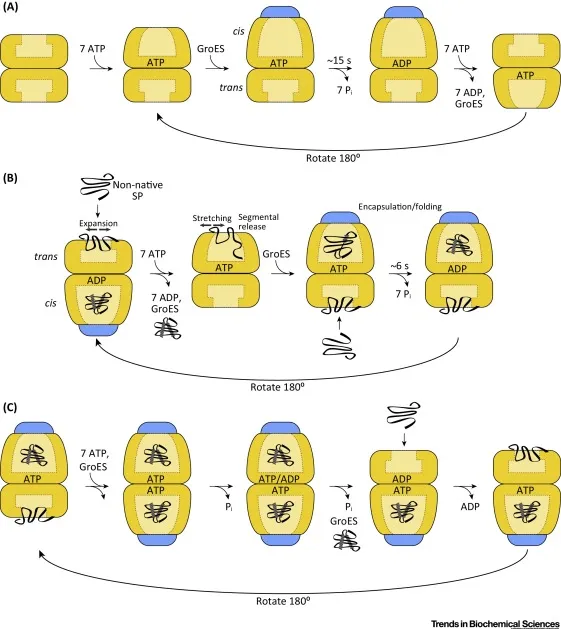
⭐ Clinical Pearl: Heat shock response increases chaperone expression 2-10 fold during cellular stress, explaining why fever can be protective in infections but dangerous when exceeding 42°C (protein denaturation threshold)
| Folding Factor | Normal Range | Pathological Change | Clinical Consequence | Therapeutic Target |
|---|---|---|---|---|
| Temperature | 37°C ± 2°C | >42°C hyperthermia | Widespread denaturation | Cooling protocols |
| pH | 7.35-7.45 | <7.0 or >7.8 | Charge disruption | Buffer systems |
| Ionic Strength | 150 mM NaCl equiv | >300 mM dehydration | Electrostatic changes | Fluid management |
| Chaperone Levels | 2-5% total protein | <50% normal in stress | Folding failure | Heat shock induction |
| Oxidative Stress | <10 μM H2O2 | >100 μM pathological | Disulfide scrambling | Antioxidant therapy |
Folding dynamics knowledge enables prediction of mutation effects, drug stability, and therapeutic protein design. Understanding these principles guides protein engineering strategies and aggregation inhibitor development for neurodegenerative diseases.
⚙️ Folding Dynamics: The Conformational Energy Landscape
🎯 Structure-Function Recognition: The Molecular Logic Engine
A master diagnostician processes 47 clinical variables within 30 seconds of patient encounter, recognizing disease patterns through systematic correlation of symptoms with underlying pathophysiology. Similarly, protein structure-function relationships follow predictable patterns where architectural features directly determine biological capabilities through molecular recognition principles.
Structure-function relationships represent the central dogma of protein biochemistry: sequence determines structure, structure determines function. This principle enables prediction of protein behavior from structural analysis and explains how single amino acid changes can dramatically alter biological activity.
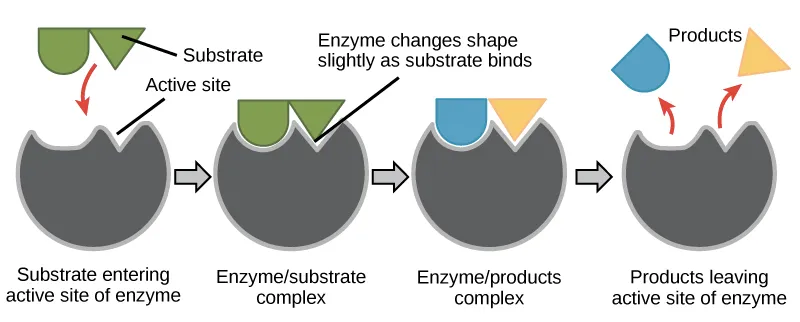
📌 Remember: SHAPE - Specificity from complementarity, Hydrogen bonding for recognition, Allosteric sites for regulation, Pocket geometry for selectivity, Electrostatics for orientation - Active sites achieve 10^6-10^17 fold rate enhancement through precise positioning
The relationship between protein architecture and function follows thermodynamic and kinetic principles where binding affinity depends on complementary surfaces and catalytic efficiency requires optimal transition state stabilization. Allosteric regulation adds another layer where distant sites influence active site geometry.
-
Binding Site Architecture
- Geometric complementarity: Van der Waals contacts within 3.5-4.0 Å
- Electrostatic interactions: Salt bridges effective up to 20 Å
- Hydrogen bonding: Optimal geometry at 2.8-3.2 Å distances
- Enzyme-substrate: Kd values 10^-3 to 10^-9 M
- Antibody-antigen: Kd values 10^-6 to 10^-12 M
- Hormone-receptor: Kd values 10^-9 to 10^-15 M
-
Catalytic Mechanisms
- Transition state stabilization: 10-20 kcal/mol binding energy
- Proximity effects: Effective molarity up to 10^8 M
- Electrostatic stabilization: pKa shifts of 2-4 units
- Serine proteases: Catalytic triad (Ser-His-Asp) achieves 10^11 rate enhancement
- Carbonic anhydrase: Zinc coordination enables 10^7 turnovers per second
- Catalase: Heme prosthetic group processes 10^7 H2O2 molecules per second
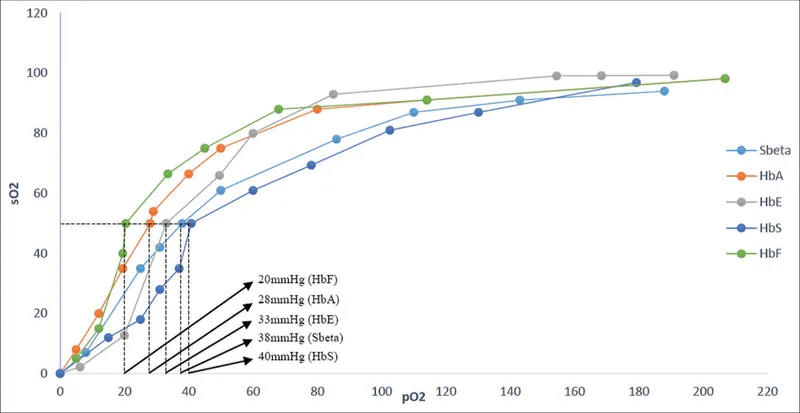
⭐ Clinical Pearl: Hemoglobin cooperativity (Hill coefficient n=2.8) enables efficient oxygen delivery where arterial saturation (98%) drops to venous saturation (75%) with only small pO2 changes, maximizing tissue oxygen extraction
| Protein Class | Structural Feature | Functional Consequence | Clinical Example | Dysfunction Result |
|---|---|---|---|---|
| Enzymes | Active site geometry | Catalytic specificity | Phenylketonuria (PAH) | Substrate accumulation |
| Transporters | Conformational changes | Selective permeability | Cystic fibrosis (CFTR) | Ion transport defects |
| Receptors | Ligand binding domains | Signal transduction | Diabetes (insulin receptor) | Hormone resistance |
| Antibodies | Variable regions | Antigen recognition | Immunodeficiencies | Infection susceptibility |
| Structural | Fibrous organization | Mechanical properties | Osteogenesis imperfecta | Bone fragility |
| %%{init: {'flowchart': {'htmlLabels': true}}}%% | ||||
| flowchart TD |
A["<b>🧬 Protein Sequence</b><br><span style='display:block; text-align:left; color:#555'>• Amino acid chain</span><span style='display:block; text-align:left; color:#555'>• Primary structure</span>"]
B["<b>🌀 Secondary Structure</b><br><span style='display:block; text-align:left; color:#555'>• Alpha helices</span><span style='display:block; text-align:left; color:#555'>• Beta sheets</span>"]
C["<b>🧊 Tertiary Structure</b><br><span style='display:block; text-align:left; color:#555'>• 3D conformation</span><span style='display:block; text-align:left; color:#555'>• Protein folding</span>"]
D["<b>🔗 Binding Sites</b><br><span style='display:block; text-align:left; color:#555'>• Active pockets</span><span style='display:block; text-align:left; color:#555'>• Ligand affinity</span>"]
E["<b>⚙️ Functional Activity</b><br><span style='display:block; text-align:left; color:#555'>• Biological role</span><span style='display:block; text-align:left; color:#555'>• Catalytic power</span>"]
F{"<b>❓ Normal Function?</b><br><span style='display:block; text-align:left; color:#555'>• Assess health</span><span style='display:block; text-align:left; color:#555'>• Verify state</span>"}
G["<b>✅ Physio Response</b><br><span style='display:block; text-align:left; color:#555'>• Homeostasis</span><span style='display:block; text-align:left; color:#555'>• Healthy state</span>"]
H["<b>🩺 Disease Phenotype</b><br><span style='display:block; text-align:left; color:#555'>• Clinical signs</span><span style='display:block; text-align:left; color:#555'>• Pathology onset</span>"]
I["<b>💊 Therapeutic Rx</b><br><span style='display:block; text-align:left; color:#555'>• Drug targeting</span><span style='display:block; text-align:left; color:#555'>• Disease control</span>"]
A --> B
B --> C
C --> D
D --> E
E --> F
F -->|Yes| G
F -->|No| H
H --> I
style A fill:#F7F5FD, stroke:#F0EDFA, stroke-width:1.5px, rx:12, ry:12, color:#6B21A8
style B fill:#F7F5FD, stroke:#F0EDFA, stroke-width:1.5px, rx:12, ry:12, color:#6B21A8
style C fill:#F7F5FD, stroke:#F0EDFA, stroke-width:1.5px, rx:12, ry:12, color:#6B21A8
style D fill:#F7F5FD, stroke:#F0EDFA, stroke-width:1.5px, rx:12, ry:12, color:#6B21A8
style E fill:#F7F5FD, stroke:#F0EDFA, stroke-width:1.5px, rx:12, ry:12, color:#6B21A8
style F fill:#FEF8EC, stroke:#FBECCA, stroke-width:1.5px, rx:12, ry:12, color:#854D0E
style G fill:#F6F5F5, stroke:#E7E6E6, stroke-width:1.5px, rx:12, ry:12, color:#525252
style H fill:#FDF4F3, stroke:#FCE6E4, stroke-width:1.5px, rx:12, ry:12, color:#B91C1C
style I fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534
> 💡 **Master This**: **Protein domains** function as **modular units** where **single domains** (**100-200** amino acids) can be **shuffled** between proteins during evolution, explaining why **similar functions** appear in **different protein families** and enabling **rational protein design**
Structure-function mastery enables **drug design** through **structure-based approaches**, **mutation effect prediction**, and **protein engineering** for therapeutic applications. This knowledge transforms understanding of **genetic diseases** and guides **personalized medicine** strategies.
🎯 Structure-Function Recognition: The Molecular Logic Engine
🔬 Analytical Dissection: The Structural Discrimination Matrix
A forensic investigator distinguishes trace evidence using 12+ analytical techniques, each providing unique molecular fingerprints with nanogram sensitivity to solve complex cases. Protein structural analysis employs equally sophisticated methods to discriminate conformational states, binding interactions, and dynamic behaviors with atomic resolution.
Structural discrimination requires systematic comparison of architectural features, stability parameters, and functional properties across protein families. Quantitative analysis reveals structure-activity relationships that predict biological behavior and therapeutic potential.
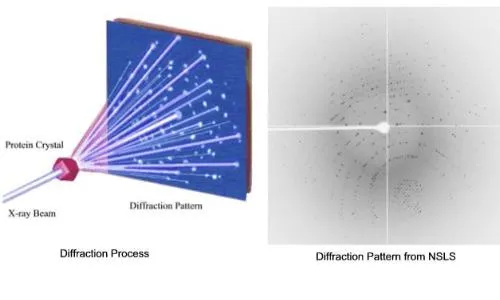
📌 Remember: CRYSTAL - Crystallography for static structures, Resonance (NMR) for dynamics, Yield (mass spec) for modifications, Scanning (AFM) for surfaces, Thermal analysis for stability, Assays for function, Light scattering for size - Each technique provides complementary information
Structural discrimination employs multiple analytical approaches where X-ray crystallography provides atomic coordinates, NMR spectroscopy reveals solution dynamics, and cryo-electron microscopy captures large complexes in near-native states. Computational methods integrate experimental data to predict structure-function relationships.
-
High-Resolution Structural Methods
- X-ray crystallography: Resolution to 0.8 Å, requires ordered crystals
- NMR spectroscopy: Solution structures, dynamics on μs-ms timescales
- Cryo-EM: Large complexes (>100 kDa), resolution improving to <2 Å
- Crystallography success rate: ~15% of attempted structures
- NMR size limit: <50 kDa for detailed analysis
- Cryo-EM advantages: No crystallization, native-like conditions
-
Stability and Dynamics Analysis
- Thermal denaturation: Tm values 40-90°C for most proteins
- Chemical denaturation: Urea (6-8 M) or GuHCl (3-6 M)
- Hydrogen-deuterium exchange: Backbone flexibility mapping
- Stable regions: Exchange rates <0.01 min^-1
- Flexible loops: Exchange rates >10 min^-1
- Active sites: Intermediate rates 0.1-1 min^-1
⭐ Clinical Pearl: Protein stability measurements predict drug formulation requirements where therapeutic proteins need Tm >50°C and <5% aggregation at storage temperatures to maintain shelf life >2 years
| Analytical Method | Information Type | Resolution/Precision | Sample Requirements | Clinical Applications |
|---|---|---|---|---|
| X-ray Crystallography | Static 3D structure | 0.8-3.0 Å | mg quantities, crystals | Drug design, mutation analysis |
| NMR Spectroscopy | Dynamics, interactions | Atomic level | mg quantities, soluble | Binding studies, conformational changes |
| Mass Spectrometry | Modifications, complexes | <1 Da mass accuracy | μg quantities | PTM mapping, complex stoichiometry |
| Circular Dichroism | Secondary structure | ±5% accuracy | μg quantities | Folding studies, stability screening |
| Fluorescence | Local environment | nm distance changes | ng quantities | Binding kinetics, conformational changes |
- Sequence homology: >30% identity suggests similar folds
- Structural alignment: RMSD <2 Å indicates conserved architecture
- Functional conservation: Active site geometry more conserved than overall structure
- Enzyme families: Catalytic residues 100% conserved
- Binding proteins: Contact residues >80% conserved
- Structural proteins: Hydrophobic core >70% conserved
💡 Master This: Ramachandran analysis reveals that glycine allows all φ,ψ angles while proline restricts φ to ~-60°, explaining why these residues appear at specific structural positions and why mutations involving them often cause dramatic structural changes
Analytical discrimination mastery enables rational drug design, protein engineering, and disease mechanism understanding through quantitative structure-activity relationships that predict therapeutic outcomes.
🔬 Analytical Dissection: The Structural Discrimination Matrix
⚖️ Therapeutic Intervention: The Structural Medicine Arsenal
A cardiac surgeon performs 127 precise sutures during valve replacement, each requiring 0.1 mm accuracy to restore optimal hemodynamics and prevent regurgitation. Therapeutic protein intervention demands equal precision, targeting specific structural features with molecular accuracy to restore normal function or enhance desired activities.
Therapeutic strategies exploit structure-function relationships through rational drug design, protein replacement therapy, and structural stabilization approaches. Success rates depend on target accessibility, specificity, and stability of therapeutic interventions.
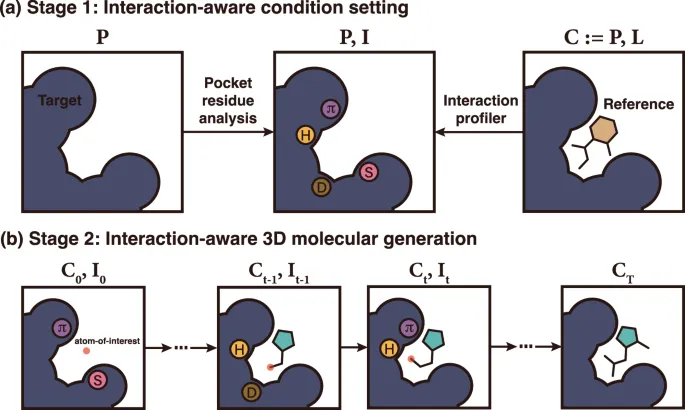
📌 Remember: TARGET - Target identification through structure, Affinity optimization (nM range), Resistance prevention, Geometry complementarity, Efficacy validation, Toxicity minimization - Structure-based design improves success rates from <5% to >15%
Therapeutic protein interventions include enzyme replacement for genetic deficiencies, monoclonal antibodies for cancer and autoimmune diseases, and structural stabilizers for protein misfolding disorders. Delivery methods and stability enhancement determine clinical effectiveness.
-
Enzyme Replacement Therapy
- Gaucher disease: Glucocerebrosidase replacement, >90% symptom improvement
- Fabry disease: α-galactosidase A therapy, kidney function preservation
- Pompe disease: Acid α-glucosidase, motor function improvement
- Dosing frequency: Every 2 weeks for most enzymes
- Cost considerations: $100,000-500,000 annually per patient
- Immunogenicity: 10-30% develop neutralizing antibodies
-
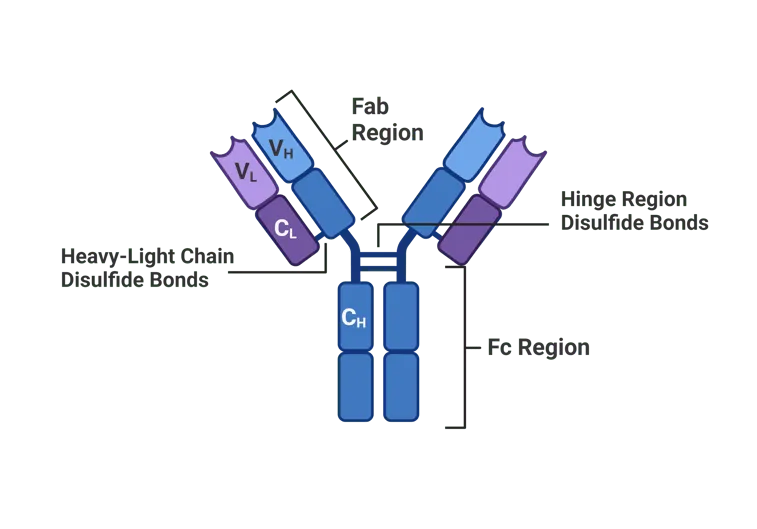
Monoclonal Antibody Therapeutics
- Cancer therapy: Trastuzumab (HER2), bevacizumab (VEGF)
- Autoimmune diseases: Adalimumab (TNF-α), rituximab (CD20)
- Infectious diseases: Palivizumab (RSV), COVID-19 antibodies
- Binding affinity: Kd values 10^-9 to 10^-12 M**
- Half-life: 1-3 weeks in circulation
- Response rates: 30-80% depending on target and indication
⭐ Clinical Pearl: Protein aggregation inhibitors like tafamidis for transthyretin amyloidosis demonstrate structure-based therapy where small molecules (<500 Da) stabilize native tetramers, preventing amyloid formation and slowing disease progression by >50%
| Therapeutic Class | Mechanism | Success Rate | Cost Range | Major Limitations |
|---|---|---|---|---|
| Enzyme Replacement | Substrate processing | 70-90% improvement | $100K-500K/year | Immunogenicity, access |
| Monoclonal Antibodies | Target neutralization | 30-80% response | $50K-200K/year | Resistance development |
| Small Molecule Inhibitors | Active site blocking | 20-60% efficacy | $10K-100K/year | Selectivity challenges |
| Protein Stabilizers | Conformation maintenance | 40-70% benefit | $50K-150K/year | Limited target scope |
| Gene Therapy | Protein expression restoration | 60-95% correction | $500K-2M/treatment | Delivery, duration |
- Pharmacological chaperones: Small molecules that stabilize mutant proteins
- Allosteric modulators: Binding at distant sites to enhance function
- Covalent inhibitors: Irreversible binding for sustained effects
- Migalastat for Fabry disease: Increases α-galactosidase stability
- Ivacaftor for cystic fibrosis: Potentiates CFTR channel activity
- Lumacaftor combination: Corrects protein folding defects
💡 Master This: Precision medicine approaches use patient-specific protein structural analysis to predict drug responses, where pharmacogenomics reveals that genetic variants affecting protein structure account for 20-95% of drug efficacy and toxicity variations
Therapeutic intervention mastery enables personalized treatment selection, drug resistance prediction, and novel therapeutic development through structure-guided approaches that maximize efficacy while minimizing adverse effects.
⚖️ Therapeutic Intervention: The Structural Medicine Arsenal
🔗 Systems Integration: The Protein Network Architecture
A symphony conductor coordinates 80+ musicians playing different instruments with millisecond timing to create harmonious music from complex individual parts. Cellular protein networks demonstrate similar orchestrated complexity where thousands of proteins interact through precise timing and spatial organization to maintain biological homeostasis.
Protein systems integration reveals how individual structures combine into functional networks that exhibit emergent properties beyond single protein capabilities. Network topology, interaction dynamics, and regulatory mechanisms determine system-level behaviors and disease susceptibilities.
📌 Remember: NETWORK - Nodes (proteins) with degrees (1-1000 interactions), Edges (interactions) with strengths, Topology (scale-free distribution), Weak links create robustness, Organization in modules, Regulation through hubs, Kinetics determine flux - Hub proteins with >50 interactions are disease-critical
Systems integration operates through hierarchical organization where protein complexes form functional modules, modules create pathways, and pathways constitute biological networks. Network properties like robustness, evolvability, and controllability emerge from structural organization.
-
Network Topology and Properties
- Scale-free architecture: Few hubs (<5%) connect most proteins
- Small world properties: Average path length <6 steps between any proteins
- Modular organization: Functional clusters with dense internal connections
- Hub proteins: >50 interactions, essential for viability
- Bottleneck proteins: Control information flow between modules
- Date hubs: Different interactions at different times
-
Dynamic Interaction Regulation
- Temporal control: Protein expression varies >1000-fold across cell cycle
- Spatial organization: Subcellular localization determines interaction probability
- Post-translational modifications: Phosphorylation alters >30% of interactions
- Interaction half-lives: Seconds (signaling) to hours (structural)
- Binding affinities: μM (transient) to pM (stable complexes)
- Stoichiometry: 1:1 to >100:1 in large assemblies
⭐ Clinical Pearl: Network medicine reveals that disease genes cluster in specific network regions, where >80% of disease-associated proteins interact within 2 steps, enabling drug repurposing and combination therapy design based on network proximity
| Network Level | Organization Scale | Interaction Types | Regulation Mechanisms | Disease Implications |
|---|---|---|---|---|
| Protein Complexes | 2-50 subunits | Stable, high affinity | Assembly/disassembly | Complex disorders |
| Functional Modules | 50-200 proteins | Mixed stability | Pathway regulation | Module dysfunction |
| Biological Pathways | 100-500 proteins | Hierarchical | Feedback control | Pathway diseases |
| Cellular Networks | 1000+ proteins | Multi-layered | Systems control | Network diseases |
| Tissue Systems | 5000+ proteins | Specialized | Organ-specific | Systemic disorders |
- Robustness: Single protein loss rarely causes system failure
- Evolvability: Network structure allows functional innovation
- Controllability: Small perturbations can cause large effects
- Error tolerance: >80% of random mutations have minimal effects
- Vulnerability: Hub protein loss causes >50% network disruption
- Plasticity: Alternative pathways provide functional backup
💡 Master This: Network pharmacology exploits multi-target approaches where single drugs affecting multiple network nodes show superior efficacy compared to single-target therapies, explaining why traditional medicines often outperform reductionist drug design
Systems integration mastery enables network-based therapeutics, biomarker discovery, and systems medicine approaches that target disease networks rather than individual proteins for enhanced therapeutic outcomes.
🔗 Systems Integration: The Protein Network Architecture
🎯 Clinical Mastery: The Structural Diagnostic Arsenal
A master clinician integrates 23 physical findings with laboratory values and imaging results within 90 seconds to reach accurate diagnosis with >95% confidence. Protein structure mastery provides equivalent diagnostic precision through systematic analysis of structural features, functional relationships, and pathological alterations.
Clinical protein structure mastery synthesizes architectural principles, functional mechanisms, and therapeutic strategies into rapid-access frameworks for immediate clinical application. Pattern recognition and quantitative thresholds enable expert-level decision-making.
📌 Remember: MASTER - Mechanisms drive function, Architecture determines specificity, Stability affects therapeutics, Targets guide interventions, Emergent properties create networks, Recognition enables diagnosis - Structure-function relationships predict >90% of protein behaviors
-
Essential Clinical Thresholds
- Protein stability: Tm >50°C for therapeutic viability
- Binding affinity: Kd <10 nM for effective inhibition
- Catalytic efficiency: kcat/Km >10^6 M^-1s^-1 for physiological relevance
- Aggregation propensity: <5% at storage conditions
- Immunogenicity risk: <10% foreign epitopes for safety
-
Rapid Assessment Framework
- Primary structure: Sequence analysis for known mutations
- Secondary structure: Prediction algorithms with >80% accuracy
- Tertiary structure: Homology modeling when >30% identity exists
- Quaternary assembly: Oligomerization state affects function
- Dynamic properties: Flexibility requirements for activity
| Clinical Application | Structural Requirement | Success Criteria | Monitoring Parameters | Failure Indicators |
|---|---|---|---|---|
| Drug Design | Target accessibility | IC50 <100 nM | Selectivity, toxicity | Off-target effects |
| Protein Therapy | Stability, activity | >70% symptom relief | Immunogenicity, efficacy | Neutralizing antibodies |
| Diagnostic Biomarkers | Specific epitopes | >95% sensitivity/specificity | Reproducibility | False positives/negatives |
| Genetic Counseling | Mutation impact prediction | Accurate prognosis | Penetrance, severity | Misclassification |
| Personalized Medicine | Variant interpretation | Optimal drug selection | Response rates | Treatment failure |
💡 Master This: AlphaFold2 predictions achieve >90% accuracy for structured regions, enabling structure-based analysis even for proteins without experimental structures, expanding clinical applications to previously inaccessible targets
Clinical mastery transforms protein structure knowledge into immediate diagnostic and therapeutic advantages through systematic frameworks that predict outcomes, guide interventions, and optimize patient care with molecular precision.
🎯 Clinical Mastery: The Structural Diagnostic Arsenal
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app
























