Translation: Components - The Protein Factory Crew
- Ribosomes: Site of protein synthesis.
- Prokaryotes: 70S (50S+30S); Eukaryotes: 80S (60S+40S).
- Sites: A (aminoacyl), P (peptidyl), E (exit).
- mRNA: Carries genetic code as codons (AUG start; UAA, UAG, UGA stop).
- tRNA: Anticodon pairs with mRNA codon; 3'-CCA attaches specific amino acid.
- Aminoacyl-tRNA Synthetases: "Charge" tRNAs; crucial proofreading.
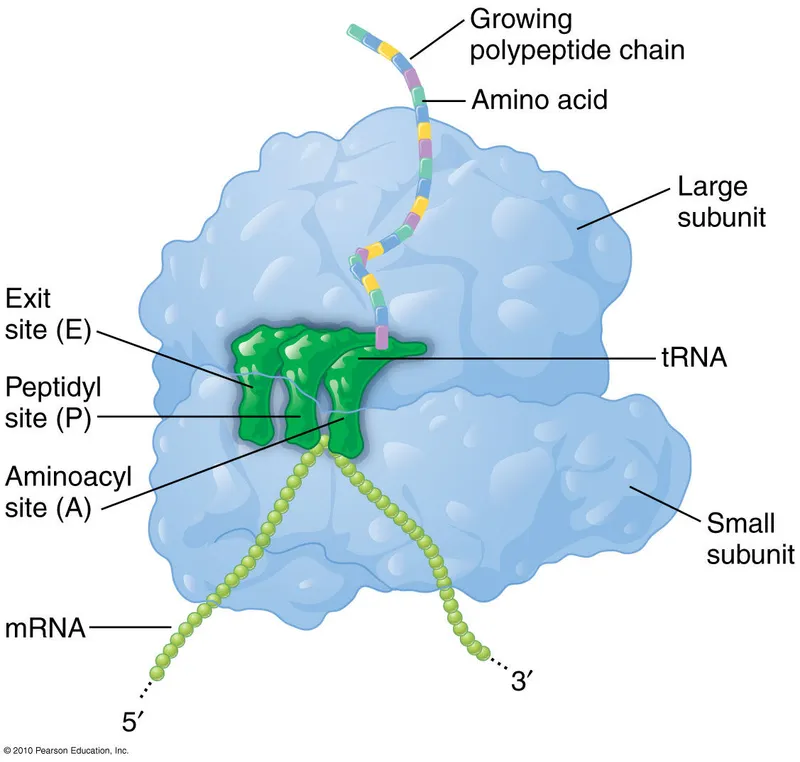
⭐ Ribosomal RNA (rRNA) in the large subunit (e.g., 23S in prokaryotes, 28S in eukaryotes) possesses peptidyl transferase activity, making it a ribozyme.
Translation: Initiation - Ready, Set, Synthesize!
Prokaryotic Initiation (forms 70S complex):
- mRNA's Shine-Dalgarno sequence (📌 S-D for Small ribosome) aligns with 16S rRNA (30S subunit).
- Initiation Factors: IF1, IF2-GTP (binds fMet-tRNA), IF3.
- Initiator tRNA: fMet-tRNA directly to P-site.
Eukaryotic Initiation (forms 80S complex):
- 40S subunit (with Met-tRNA, eIF2-GTP) binds mRNA's 5' cap, scans to Kozak sequence (📌 Kozak for euKaryotes) around AUG.
- Initiation Factors: Multiple eIFs (e.g., eIF4E binds cap).
⭐ In prokaryotes, the Shine-Dalgarno sequence on mRNA base-pairs with a complementary sequence on the 16S rRNA of the 30S ribosomal subunit to position it correctly for initiation.
Translation: Elongation - Building the Chain
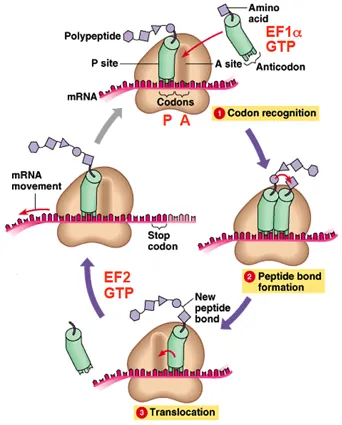
- Polypeptide chain grows via a cycle. 📌 APE sites: Aminoacyl, Peptidyl, Exit.
- Elongation Cycle (GTP-powered):
- 1. Codon Recognition: aa-tRNA + EF-Tu/eEF1A enters A-site.
- 2. Peptide Bond Formation: Peptidyl transferase (rRNA) forms peptide bond; chain moves to A-site tRNA.
- 3. Translocation: EF-G/eEF2 moves ribosome; tRNAs shift (A→P, P→E & exit).
⭐ EF-Tu (prokaryotes) or eEF1A (eukaryotes) is responsible for bringing the charged aminoacyl-tRNA to the A site of the ribosome, a step requiring GTP hydrolysis.
Translation: Termination - It's a Wrap!
- Stop Codons: UAA, UAG, UGA. 📌 (U Are Away, U Go Away, U Are Gone).
- Release Factors (RFs):
- Prokaryotes: RF1, RF2, RF3.
- Eukaryotes: eRF1, eRF3.
- Mechanism:
- RFs bind stop codon at A-site.
- Polypeptide hydrolysis from tRNA.
- Ribosomal subunits, mRNA, tRNA dissociate.
⭐ Stop codons are recognized by protein Release Factors (RFs), not by tRNAs, leading to the termination of translation.
Post-Translational Mods - Protein's Extreme Makeover
- Protein Folding: Chaperones (e.g., Hsp70) ensure correct 3D structure.
- Covalent Modifications:
- Phosphorylation, glycosylation, ubiquitination.
- Acetylation, methylation, proteolytic cleavage.
- Disulfide bond formation.
- Significance: Critical for protein function, stability, and diversity.
⭐ Glycosylation, the addition of sugar moieties to proteins, typically begins in the endoplasmic reticulum and continues in the Golgi apparatus, crucial for protein folding, stability, and targeting.
Protein Synthesis Inhibitors - Antibiotic Hit List
📌 Mnemonic: "Buy AT 30, CCELL at 50"
| Antibiotic | Target Subunit(s) | Mechanism | Organism |
|---|---|---|---|
| Streptomycin | 30S | Misreading mRNA, inhibits initiation (high conc.) | Prokaryotic |
| Tetracyclines | 30S | Blocks A-site | Prokaryotic |
| Chloramphenicol | 50S | Blocks peptidyl transferase | Prokaryotic |
| Macrolides | 50S | Blocks translocation | Prokaryotic |
| Clindamycin | 50S | Blocks translocation | Prokaryotic |
| Linezolid | 50S | Blocks initiation complex formation | Prokaryotic |
| Puromycin | A-site | Premature termination | Both |
| Cycloheximide | 80S (Euk) | Blocks translocation | Eukaryotic |
| Diphtheria Toxin | eEF2 | Inactivates eEF2 (ADP-ribosylation) | Eukaryotic |
High‑Yield Points - ⚡ Biggest Takeaways
- Translation occurs on ribosomes (70S prokaryotes, 80S eukaryotes), decoding mRNA to protein.
- mRNA codons pair with tRNA anticodons carrying specific amino acids.
- Initiation: AUG start codon (Met); Shine-Dalgarno (prok) / Kozak sequence (euk).
- Elongation: Peptidyl transferase (rRNA ribozyme) forms peptide bonds in A, P, E sites.
- Termination: Stop codons (UAA, UAG, UGA) trigger release of polypeptide.
- Key inhibitors: Many antibiotics (tetracyclines, macrolides) target 70S ribosomes.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

