Gene Regulation Basics - The Control Panel
- Gene Regulation: Cellular processes that control the rate and manner of gene expression. Ensures proteins are produced at the right time and amount.
- Control Points: Primarily at transcription initiation; also post-transcriptional, translational, and post-translational levels.
- Key Elements & Proteins:
- Cis-acting elements: DNA sequences (e.g., promoters, enhancers, silencers, operators) on the same DNA molecule.
- Trans-acting factors: Proteins (e.g., transcription factors, activators, repressors) that bind to cis-elements.
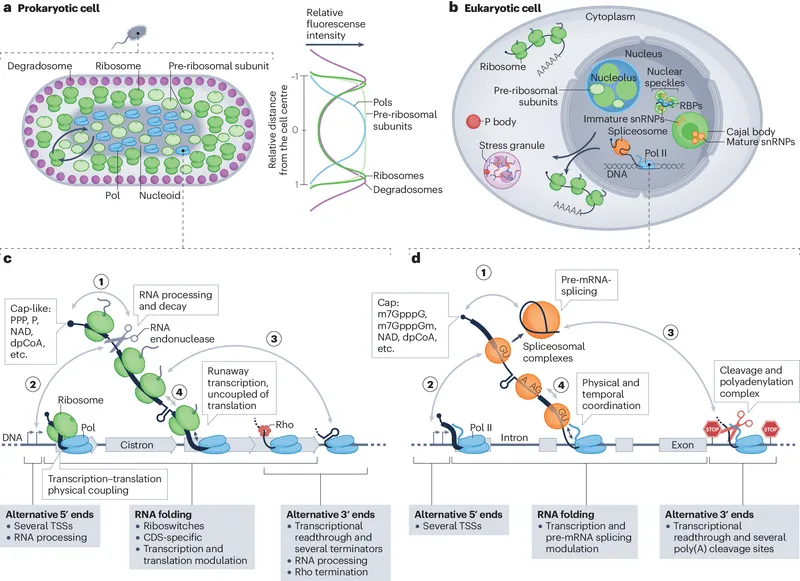
- Prokaryotes vs. Eukaryotes:
- Prokaryotes: Simpler; often involves operons (e.g., Lac, Trp operons) for coordinated gene expression.
- Eukaryotes: More complex; involves chromatin remodeling (acetylation/deacetylation, methylation), multiple transcription factors, and RNA processing.
⭐ In eukaryotes, enhancers can be located thousands of base pairs away from the promoter and can function in either orientation.
Prokaryotic Regulation - Operon Operators
- Operon: DNA unit; multiple genes under a single promoter & operator.
- Operator (O):
- DNA sequence; repressor protein binding site.
- Repressor binding to operator blocks RNA polymerase, halting transcription (negative control).
- Lac Operon (Inducible):
- No inducer (allolactose): LacI repressor binds operator → transcription OFF.
- Inducer present: LacI releases → transcription ON.
- Trp Operon (Repressible):
- Co-repressor (tryptophan) present: TrpR + tryptophan binds operator → transcription OFF.
- No co-repressor: TrpR inactive, operator free → transcription ON.
⭐ Operator sequences are often palindromic, facilitating binding of dimeric repressor proteins.
Eukaryotic Regulation - Complex Choreography
- Levels of Control: Chromatin structure, transcription, RNA processing & transport, translation, post-translation.
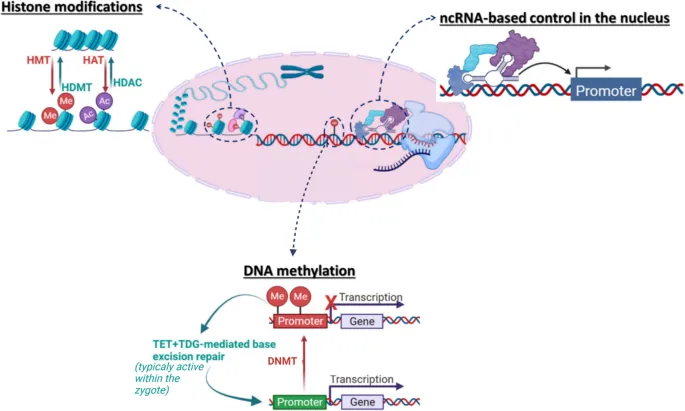
- Chromatin Remodeling:
- Histone Acetylation (HATs): ↑Expression (euchromatin).
- Histone Deacetylation (HDACs): ↓Expression (heterochromatin).
- DNA Methylation (CpG islands): Gene silencing.
- Transcriptional Initiation:
- Promoters (TATA box, CAAT box), Enhancers (distal), Silencers.
- Transcription Factors (TFs): General (basal) & specific (regulatory).
- DNA-binding motifs: Zinc finger, Leucine zipper, Helix-loop-helix.
- Mediator complex: Bridges TFs & RNA Polymerase II.
- Post-Transcriptional Control:
- RNA Processing: 5' capping, 3' polyadenylation (poly-A tail), splicing.
- Alternative splicing → protein diversity.
- mRNA transport from nucleus to cytoplasm.
- RNA Processing: 5' capping, 3' polyadenylation (poly-A tail), splicing.
- Translational & Post-Translational Control:
- mRNA stability (e.g., AU-rich elements in 3'UTR).
- RNA interference (miRNA, siRNA): mRNA degradation or translation repression.
- Protein modification (e.g., phosphorylation, ubiquitination).
⭐ Many eukaryotic genes are regulated by enhancers, DNA sequences that can be thousands of base pairs away from the promoter and significantly increase transcription rates when bound by activator proteins.
Advanced & Applied - Tweaks & Troubles
- RNA Interference (RNAi): Post-transcriptional gene silencing.
- siRNA/miRNA: Guide RISC to target mRNA for cleavage/repression.
- Processed by Dicer; therapeutic potential (e.g., antiviral, cancer).
- Epigenetics: Heritable gene expression changes; no DNA sequence alteration.
- DNA Methylation: CpG islands; silences genes (DNMTs).
- Histone Modification:
- Acetylation (HATs): ↑ expression (euchromatin).
- Deacetylation (HDACs): ↓ expression (heterochromatin).
- Methylation: Variable effects.
- Genomic Imprinting: Parent-specific expression (e.g., Prader-Willi, Angelman).
- Clinical Links & Therapies:
- Cancer: Aberrant DNA methylation/histone mods silence Tumor Suppressor Genes (TSGs).
- Syndromes: Imprinting disorders (Prader-Willi, Angelman).
- Drugs: Azacitidine (DNMT inhibitor), Vorinostat (HDAC inhibitor).
⭐ Hypermethylation of CpG islands in TSG promoters is a hallmark of many cancers, leading to gene silencing. (📌 Methylation Mutes)
High‑Yield Points - ⚡ Biggest Takeaways
- Operon model (e.g., lac, trp) is central to prokaryotic regulation; lac operon is inducible, trp operon repressible.
- Eukaryotic control uses transcription factors, enhancers, silencers, and promoters.
- Chromatin remodeling (e.g., histone acetylation ↑, methylation) critically impacts gene accessibility.
- RNA interference (RNAi) by miRNAs and siRNAs causes post-transcriptional gene silencing.
- Alternative splicing generates diverse protein isoforms from a single eukaryotic gene.
- Many hormones regulate genes via nuclear receptors acting as transcription factors.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more