RNA Processing and Splicing: Overview - Transcript Makeover
- Eukaryotic pre-mRNA undergoes extensive post-transcriptional modifications in the nucleus to become mature mRNA.
- Essential for mRNA stability, nuclear export, and efficient translation.
- Key modifications ("transcript makeover"):
- 5' Capping: Addition of $m^7G$ (7-methylguanosine) cap.
- Splicing: Intron removal, exon ligation.
- 3' Polyadenylation: Addition of poly(A) tail.
- Prokaryotes: Minimal processing; transcription-translation often coupled.
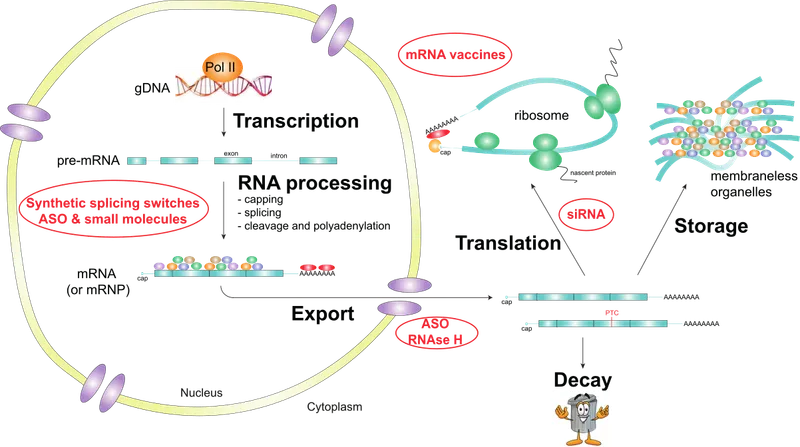
⭐ Most human genes contain introns; precise splicing is vital for correct protein synthesis.
RNA Processing and Splicing: 5' Cap & Poly‑A Tail - RNA's Hat & Tail
- 5' Cap ("Hat"):
- Structure: Modified guanine (7-methylguanosine, m7G) linked via unusual 5'-5' triphosphate bridge.
- Enzymes: RNA triphosphatase, guanylyltransferase, methyltransferase.
- Functions: Protects mRNA from 5' exonuclease degradation; essential for nuclear export; promotes translation initiation (recognized by eIF4E).
- Poly-A Tail ("Tail"):
- Structure: Chain of ~200-250 adenine nucleotides at 3' end.
- Enzyme: Poly(A) polymerase (PAP), template-independent.
- Signal: AAUAAA (polyadenylation signal) upstream of cleavage site.
- Functions: Protects from 3' exonuclease degradation; enhances translation efficiency (PABP binding); aids nuclear export; regulates mRNA stability.
⭐ Poly(A) polymerase adds the poly-A tail post-transcriptionally without requiring a DNA template for the adenine residues a crucial distinction from DNA/RNA polymerases during transcription.
RNA Processing and Splicing: The Spliceosome Show - Introns Out!
- Splicing: Crucial post-transcriptional modification; precisely removes introns (non-coding regions) and ligates exons (coding regions) from pre-mRNA.
- The Spliceosome: Dynamic, large RNA-protein complex that orchestrates splicing.
- Components: snRNPs (small nuclear ribonucleoproteins) - U1, U2, U4, U5, U6 - plus other proteins.
- U1 snRNP binds the 5' splice site.
- U2 snRNP binds the branch point A.
- Consensus Sequences:
- 5' splice site (donor): GU sequence.
- 3' splice site (acceptor): AG sequence.
- Branch Point: Adenine (A) within the intron, upstream of 3' site.
- Components: snRNPs (small nuclear ribonucleoproteins) - U1, U2, U4, U5, U6 - plus other proteins.
- Mechanism (Two Transesterification Reactions):
- The 2'-OH of branch point A attacks the 5' splice site → forms a lariat (loop) structure.
- The freed 3'-OH of the 5' exon attacks the 3' splice site → exons are joined; lariat intron is released.
- Alternative Splicing: Allows a single gene to produce multiple distinct mRNA (and thus protein) variants. Significantly expands proteomic diversity.
⭐ Mutations in splice sites or splicing factors are a common cause of genetic diseases (e.g., some β-thalassemias, spinal muscular atrophy, cystic fibrosis variants).
RNA Processing and Splicing: Splice Variants & Errors - Code Remix & Glitches
- Alternative Splicing: "Code Remix" - one gene, multiple protein isoforms.
- Mechanism: Differential inclusion/exclusion of exons.
- Impact: ↑ Proteome diversity; tissue-specific functions (e.g., tropomyosin).
- RNA Editing: Further sequence modification post-transcription.
- Types: A-to-I (by ADARs), C-to-U (by APOBECs).
- Example: ApoB mRNA editing (ApoB100 vs. ApoB48).
- Splicing Errors: "Code Glitches" - mutations disrupting normal splicing.
- Targets: Splice sites (📌 GU-AG rule), enhancers, silencers.
- Outcomes: Exon skipping, intron retention, cryptic splice site activation.
- Diseases: β-thalassemia, Spinal Muscular Atrophy (SMA), Duchenne Muscular Dystrophy.
⭐ A significant proportion, estimated around 15-50%, of human genetic disease mutations affect pre-mRNA splicing.

High‑Yield Points - ⚡ Biggest Takeaways
- hnRNA is the primary transcript processed into mature mRNA.
- 5' capping with 7-methylguanosine protects mRNA and aids ribosome binding.
- 3' polyadenylation adds a poly(A) tail, increasing mRNA stability and export.
- Splicing, by spliceosomes (snRNPs), removes introns and joins exons.
- GU-AG rule: Introns typically start with GU (5' site) and end with AG (3' site).
- Alternative splicing generates diverse protein isoforms from one gene.
- RNA editing (e.g., APOBEC) post-transcriptionally alters RNA sequence.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more


