Epigenetics and DNA Methylation - Genes Beyond Sequence
- Epigenetics: Heritable gene expression changes; no DNA sequence alteration. Regulates gene activity.
- Key Mechanisms:
- DNA Methylation: Methyl group ($CH_3$) addition to DNA, primarily at CpG sites.
- Histone Modification: Acetylation, methylation, phosphorylation of histone tails.
- Non-coding RNAs (ncRNAs): e.g., microRNAs (miRNAs), long non-coding RNAs (lncRNAs).
- DNA Methylation Focus:
- Enzymes: DNA Methyltransferases (DNMTs).
- Effect: Usually gene silencing (transcriptional repression).
- CpG islands (promoters): methylation often leads to ↓ gene expression.
- Reversible: Demethylation (e.g., by TET enzymes) can reactivate genes.
- Significance: Vital for normal development, cellular differentiation, genomic imprinting, X-chromosome inactivation, and diseases (e.g., cancer).
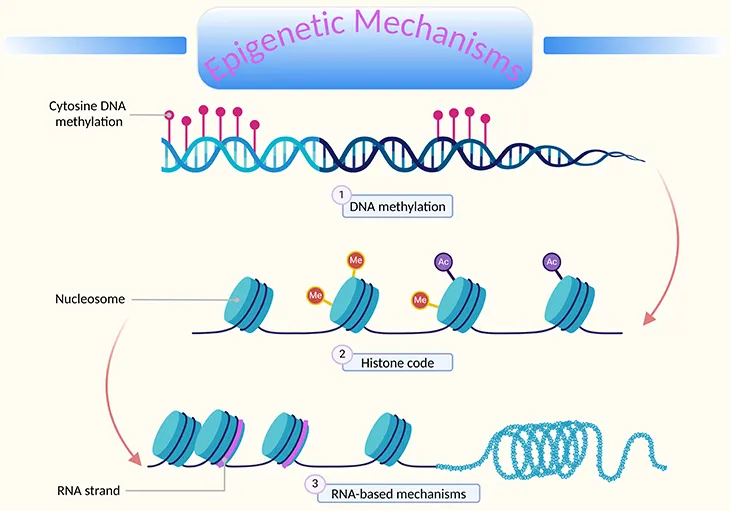
⭐ Epigenetic changes, unlike genetic mutations, do not alter the DNA sequence itself but can be stably inherited through cell divisions.
Epigenetics and DNA Methylation - Silencing Secrets
- Epigenetics: Heritable changes in gene expression without altering DNA sequence. Environmentally influenced.
- DNA Methylation: Key epigenetic modification.
- Mechanism: Addition of a methyl group ($-CH_3$) to DNA, primarily at cytosine bases.
- Enzymes: DNA Methyltransferases (DNMTs).
- Methyl donor: S-adenosylmethionine (SAM).
- Primary Site: CpG dinucleotides (Cytosine-phosphate-Guanine).
- Clusters of CpGs = CpG islands, often in promoter regions.
- Effect: Typically leads to gene silencing.
- Blocks transcription factor binding.
- Recruits methyl-CpG-binding domain proteins (MBDs) $\rightarrow$ chromatin condensation.
- Demethylation: Active process involving TET (Ten-Eleven Translocation) enzymes.
- Mechanism: Addition of a methyl group ($-CH_3$) to DNA, primarily at cytosine bases.
- Clinical Relevance: Implicated in cancer (tumor suppressor gene silencing), imprinting disorders (Prader-Willi, Angelman), developmental processes.
⭐ DNA methylation predominantly occurs at CpG dinucleotides, and methylation of CpG islands in promoter regions is typically associated with gene silencing.
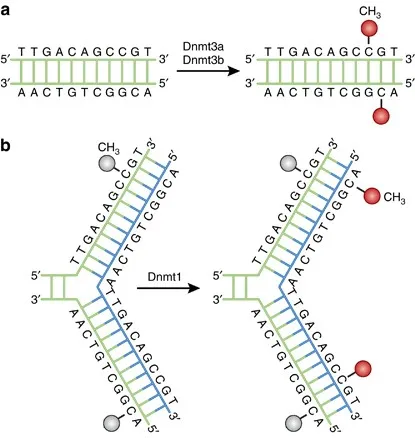
Epigenetics and DNA Methylation - Chromatin's Chat
- Histone Modifications: Reversible chemical changes to histone proteins, altering chromatin structure & gene activity.
- Key Modifications:
- Acetylation:
- By HATs (Histone Acetyltransferases).
- Loosens chromatin (euchromatin) → ↑ transcription. 📌 "A"cetylation = "A"ctive.
- Deacetylation:
- By HDACs (Histone Deacetylases).
- Compacts chromatin (heterochromatin) → ↓ transcription.
- Methylation:
- By HMTs (Histone Methyltransferases).
- Context-dependent: can activate (e.g., H3K4me3) or repress (e.g., H3K9me3, H3K27me3).
- Reversed by HDMs (Histone Demethylases).
- Phosphorylation & Ubiquitination: Also influence chromatin.
- Acetylation:
- "Histone Code": Patterns of modifications recruit specific proteins, dictating gene expression outcomes.
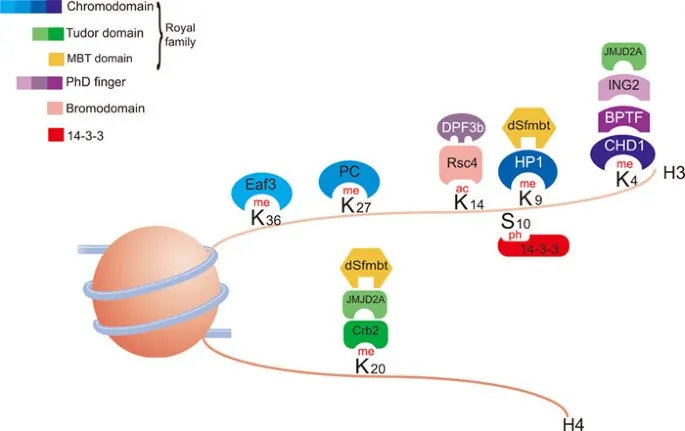
⭐ Histone acetylation, mediated by HATs, generally opens up chromatin and activates transcription, while deacetylation by HDACs has the opposite effect.
Epigenetics and DNA Methylation - Disease & Drugs
- Cancer: Aberrant DNA methylation (e.g., ↑methylation of tumor suppressors, ↓methylation of oncogenes).
- Imprinting Disorders:
- Prader-Willi (PWS), Angelman (AS) syndromes.
- Beckwith-Wiedemann, Russell-Silver syndromes.
- Other Genetic Conditions: Fragile X, Rett, ICF syndromes.
- Therapeutic Agents (Epi-drugs):
- DNMT Inhibitors: Azacitidine, Decitabine (for Myelodysplastic Syndromes (MDS), Acute Myeloid Leukemia (AML)).
- HDAC Inhibitors: Vorinostat, Romidepsin (for Cutaneous T-cell Lymphoma (CTCL), Peripheral T-cell Lymphoma (PTCL)).
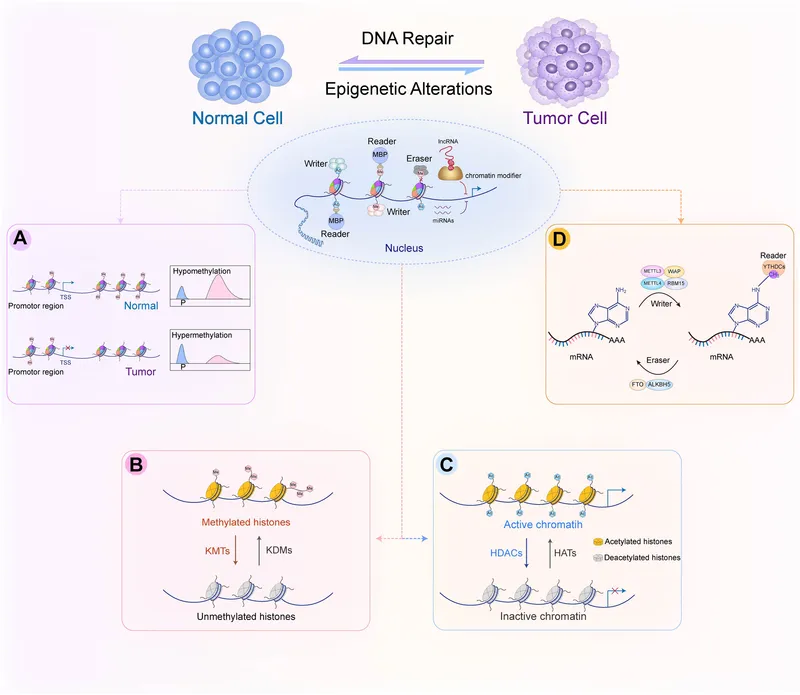
⭐ Genomic imprinting, an epigenetic phenomenon, results in monoallelic gene expression depending on the parent of origin, exemplified by Prader-Willi and Angelman syndromes.
High‑Yield Points - ⚡ Biggest Takeaways
- DNA methylation at CpG islands predominantly leads to heritable gene silencing.
- SAM (S-adenosylmethionine) is the crucial methyl donor for DNA methyltransferases (DNMTs).
- Histone acetylation by HATs activates transcription; HDACs cause repression.
- Histone methylation effects (activation/repression) are highly context-dependent.
- Genomic imprinting dictates parent-of-origin specific gene expression.
- Epigenetic dysregulation drives cancer and syndromes like Prader-Willi/Angelman.
- X-chromosome inactivation (Barr body formation) is a key epigenetic process.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

