DNA Fingerprinting Principles - Identity Unlocked
- Concept: A technique to identify individuals at the molecular level using unique DNA patterns.
- Pioneer: Sir Alec Jeffreys in 1984, revolutionizing forensic science.
- Basis - DNA Polymorphism: Variations in DNA sequences among individuals.
- These variations are primarily in non-coding regions (introns).
- VNTRs (Variable Number of Tandem Repeats): Also known as minisatellites. Comprise repeat units of 10-100 base pairs. Highly polymorphic.
- STRs (Short Tandem Repeats): Also known as microsatellites. Simpler, with 2-6 base pair repeat units. Currently more common in forensics.
- Inheritance: DNA patterns are inherited from biological parents (50% from each).
- Uniqueness: Each person's DNA fingerprint is unique, except for identical (monozygotic) twins.
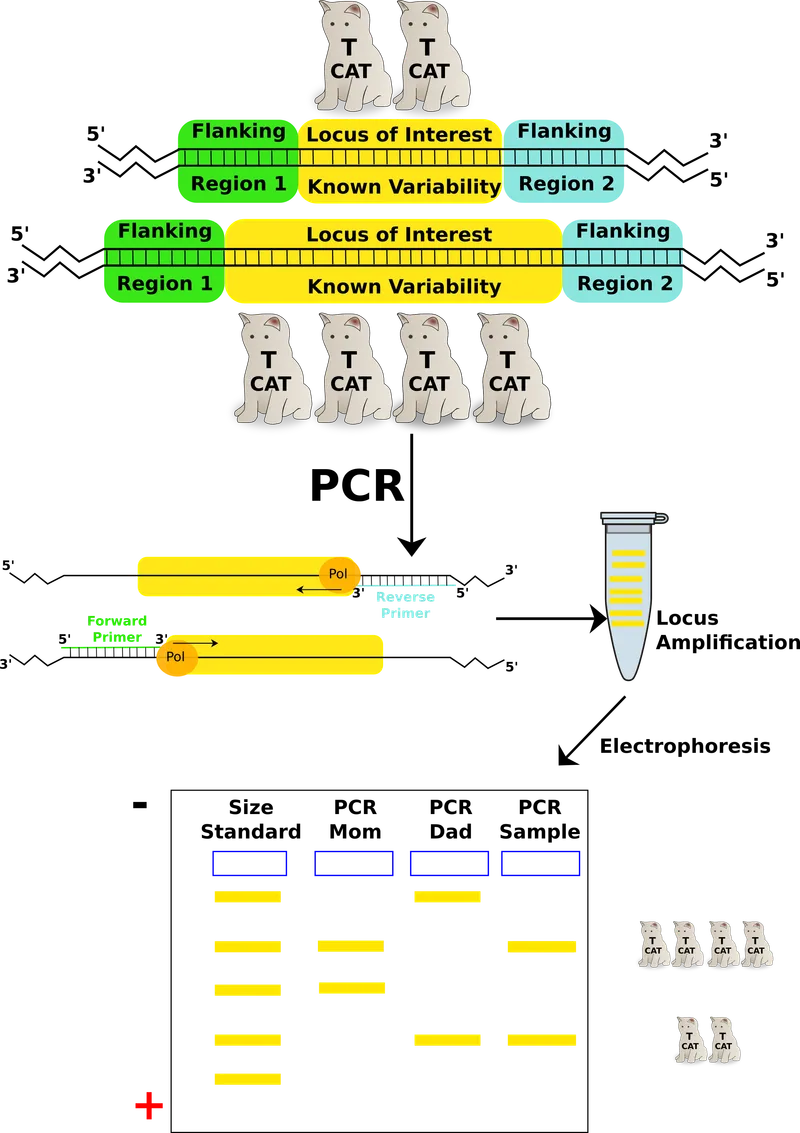
⭐ The original DNA fingerprinting method developed by Sir Alec Jeffreys utilized multi-locus probes targeting VNTRs, creating a complex, barcode-like pattern.
Forensic DNA Techniques - Lab Detectives
-
Core Principle: Analyzing unique DNA variations (polymorphisms) for individual identification.
-
Key Markers: Specific DNA regions for profiling.
- STRs (Short Tandem Repeats): Highly variable, short, repeated DNA sequences in non-coding regions. Gold standard. 📌 PCR & STRs: Partners in Crime (Solving!)
- Typically 13-20 core STR loci (e.g., CODIS) used for high discrimination.
- mtDNA (Mitochondrial DNA): Maternally inherited. For degraded samples (old bones, hair shafts) due to high copy number.
- Y-STRs: Paternally inherited (Y-chromosome). For male DNA in mixed samples (sexual assaults).
- STRs (Short Tandem Repeats): Highly variable, short, repeated DNA sequences in non-coding regions. Gold standard. 📌 PCR & STRs: Partners in Crime (Solving!)
-
Workflow:
-
Techniques:
- PCR (Polymerase Chain Reaction): Amplifies minute DNA quantities.
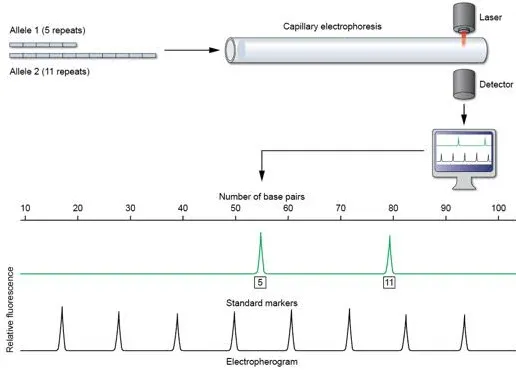
- Capillary Electrophoresis (CE): Separates amplified STR fragments by size.
- RFLP (Restriction Fragment Length Polymorphism): Older method, requires larger DNA amounts, less common.
- SNP (Single Nucleotide Polymorphism) Typing: Emerging for ancestry/phenotype.
⭐ STRs are the workhorse of modern DNA forensics, with 13-20 core loci typically analyzed to generate a highly discriminating DNA profile.
Applications & Legalities - Justice Jigsaw
- Core Forensic Uses:
- Crime Scene Analysis: Matching biological evidence (blood, semen, hair, saliva, skin cells) to suspects or victims.
- Identification:
- Victims in mass disasters (e.g., plane crashes, natural calamities).
- Missing persons.
- Exonerating the innocent.
- Beyond Crime Scenes:
- Paternity/Maternity Testing: Resolving parentage disputes.
- Immigration: Verifying claimed biological relationships.
- Wildlife Forensics: Identifying species, combating poaching & illegal trade.
- Legal Standing (India):
- Admissibility: Indian Evidence Act, 1872 (Sec. 45 - expert opinion; Sec. 112 - paternity).
- Sample Collection: CrPC Sec. 53 & 53A empower medical exam & DNA collection from accused.
- Chain of Custody: Critical for evidence integrity; meticulous record from collection to court.
- Ethical & Database Considerations:
- DNA Technology Bill: Proposes DNA Data Banks (e.g., NDNAD).
- Key Concerns:
- Privacy violations & potential data misuse.
- Informed consent for sample collection, storage & database inclusion.
- Familial searching ethics & risks of genetic surveillance.
- Accuracy & challenges in interpreting complex/degraded DNA samples.
⭐ Locus of Amelogenin gene (AMELX, AMELY) is commonly used for sex determination in forensic DNA typing.
High‑Yield Points - ⚡ Biggest Takeaways
- DNA fingerprinting relies on Variable Number Tandem Repeats (VNTRs), unique to each individual.
- PCR is crucial for amplifying small DNA samples in forensic cases.
- Short Tandem Repeats (STRs) are the current standard, used in databases like CODIS.
- Key applications: paternity testing, crime scene analysis, and identification.
- RFLP is an older method; Southern blotting is a related detection technique.
- The Amelogenin gene helps in sex determination from DNA samples.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

