Molecular Biology and Genomics

On this page
🧬 The Molecular Command Center: Decoding Life's Blueprint
You'll journey from DNA's double helix to the cutting-edge genomic tools reshaping clinical practice, mastering how cells decode genetic instructions, regulate gene expression, and translate information into functional proteins. This lesson builds your understanding of transcription, RNA processing, translation, and epigenetic control-the molecular machinery underlying every physiological process and disease state. You'll then explore how next-generation sequencing, CRISPR, and precision medicine technologies translate these principles into diagnostic power and targeted therapies that are revolutionizing patient care today.
Understanding molecular biology and genomics transforms medical practice from symptom management to precision intervention. Every genetic variant, every transcriptional change, and every epigenetic modification creates the molecular foundation for disease susceptibility, drug responses, and therapeutic outcomes. Master these molecular mechanisms, and you possess the analytical framework to predict disease patterns, optimize treatments, and interpret the genomic data that increasingly drives clinical decisions.
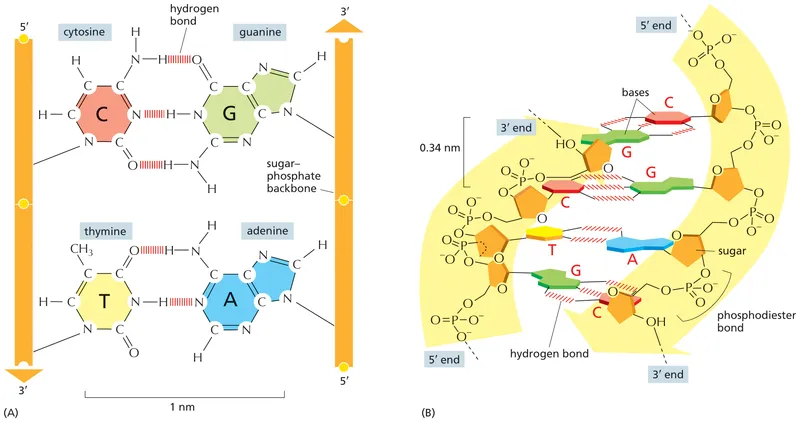
The human genome contains 20,000-25,000 protein-coding genes distributed across 23 chromosome pairs, yet protein-coding sequences represent only 1.5% of the total genome. The remaining 98.5% includes regulatory elements, non-coding RNAs, and repetitive sequences that control gene expression with extraordinary precision. Each cell contains 6 billion base pairs of DNA that, if stretched end-to-end, would span 2 meters - yet fits within a nucleus measuring only 10 micrometers in diameter.
📌 Remember: ATGC - Adenine pairs with Thymine (2 hydrogen bonds), Guanine pairs with Cytosine (3 hydrogen bonds). The stronger GC bonds explain why GC-rich regions have higher melting temperatures and greater stability.
Central dogma flow demonstrates the information transfer pathway: DNA → RNA → Protein. However, modern molecular biology reveals this as a dynamic network rather than linear progression. MicroRNAs regulate 30% of human genes post-transcriptionally, long non-coding RNAs modulate chromatin structure, and epigenetic modifications create heritable changes without altering DNA sequence.
| Molecular Component | Size Range | Function | Clinical Significance | Therapeutic Targets |
|---|---|---|---|---|
| Genomic DNA | 3.2 billion bp | Information storage | Genetic testing, GWAS | Gene therapy, CRISPR |
| mRNA | 500-15,000 nt | Protein template | Expression profiling | Antisense, siRNA |
| miRNA | 18-25 nt | Gene silencing | Biomarkers | miRNA mimics/inhibitors |
| Proteins | 50-30,000 aa | Cellular functions | Drug targets | Small molecules, biologics |
| Chromatin | Variable | Gene regulation | Epigenetic diseases | HDAC inhibitors |
- Genomic Organization Hierarchy
- Nuclear genome: 3.2 billion base pairs, 23 chromosome pairs
- Mitochondrial genome: 16,569 base pairs, 37 genes
- Maternal inheritance pattern
- 1,000-10,000 copies per cell
- Critical for oxidative metabolism
- Chromatin packaging: 147 base pairs per nucleosome
- 6 levels of DNA compaction
- 10,000-fold condensation during mitosis
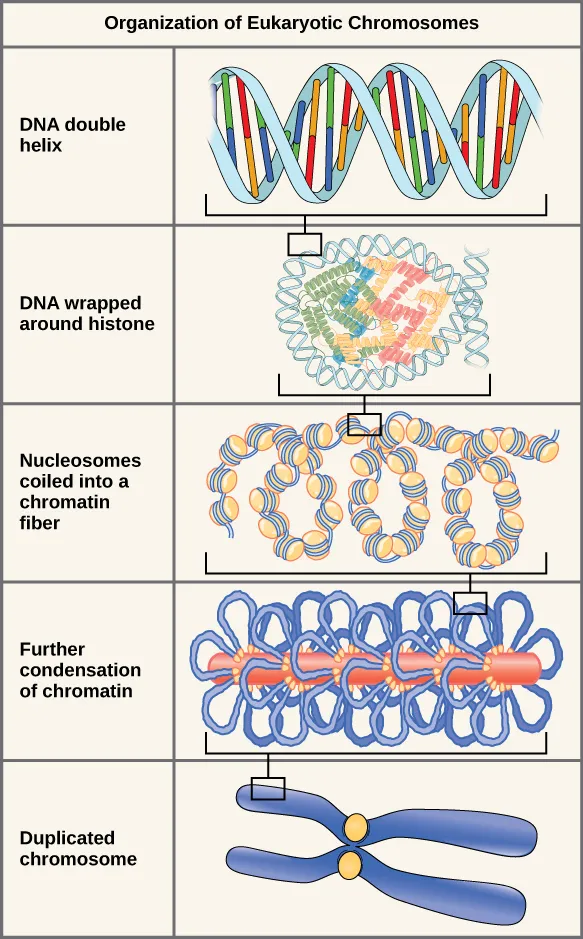
💡 Master This: Genomic imprinting affects 150+ human genes, where expression depends on parent-of-origin. Disruption causes diseases like Prader-Willi syndrome (paternal deletion) and Angelman syndrome (maternal deletion) - demonstrating how identical genetic changes produce different phenotypes based on inheritance pattern.
The molecular complexity extends beyond static DNA sequences to dynamic regulatory networks. Transcription factors bind 10,000+ regulatory sites per cell type, creating tissue-specific expression patterns. Alternative splicing generates 100,000+ protein isoforms from 20,000 genes, while post-translational modifications create additional functional diversity through 200+ modification types.
Connect this molecular foundation through transcriptional control mechanisms to understand how 4 base pairs generate the complexity of human biology and disease.
🧬 The Molecular Command Center: Decoding Life's Blueprint
⚙️ The Transcriptional Orchestra: Gene Expression Command Systems
RNA Polymerase II transcribes all protein-coding genes and most non-coding RNAs, requiring 6 general transcription factors (TFIIA, TFIIB, TFIID, TFIIE, TFIIF, TFIIH) for initiation. The pre-initiation complex assembles at core promoter elements, including the TATA box (located 25-30 base pairs upstream of transcription start site) and initiator elements. TFIIH possesses both helicase activity (unwinding DNA) and kinase activity (phosphorylating RNA polymerase II C-terminal domain).
📌 Remember: TFIID-TFIIA-TFIIB-TFIIF-RNA Pol II-TFIIE-TFIIH assembly sequence: "The Transcription Factory Includes Diverse Talented Individuals Building Fantastic Proteins Every Time, Including Heavy-duty Help"
Enhancer elements regulate transcription from distances up to 1 million base pairs through chromatin looping mechanisms. Mediator complex (26 subunits) bridges enhancer-bound activators with promoter-associated machinery, creating 3D chromatin architecture that brings distant regulatory elements into proximity. Super-enhancers span 10-50 kilobases and contain dense clusters of transcription factor binding sites, driving high-level expression of cell identity genes.
| Regulatory Element | Distance from TSS | Function | Binding Factors | Clinical Relevance |
|---|---|---|---|---|
| Core Promoter | -50 to +50 bp | Initiation | General TFs | Promoter mutations |
| Proximal Promoter | -500 to -50 bp | Fine-tuning | Specific TFs | Regulatory SNPs |
| Enhancers | 1 kb to 1 Mb | Activation | Activators | GWAS associations |
| Silencers | Variable | Repression | Repressors | Tumor suppressors |
| Insulators | Variable | Boundary | CTCF | Imprinting disorders |
- Chromatin States and Histone Modifications
- H3K4me3: Active promoters (transcription start sites)
- H3K27ac: Active enhancers (tissue-specific regulation)
- H3K36me3: Gene bodies (actively transcribed regions)
- Prevents cryptic initiation
- Recruits DNA repair machinery
- Maintains transcriptional fidelity
- H3K27me3: Polycomb repression (developmental silencing)
- Bivalent domains in stem cells
- CpG island association
- Cancer stem cell maintenance
💡 Master This: Transcriptional memory allows cells to respond faster to repeated stimuli through chromatin bookmarking and poised RNA polymerase II. This mechanism underlies trained immunity, stress responses, and drug tolerance - explaining why previous exposures influence future cellular responses even after stimulus removal.
Alternative promoters generate tissue-specific isoforms for 50% of human genes, creating functional diversity through different N-terminal domains or subcellular localizations. Bidirectional promoters (10% of genes) coordinate expression of gene pairs, often involved in DNA repair, cell cycle control, and metabolic pathways. TATA-less promoters (80% of genes) use CpG islands, initiator elements, and downstream promoter elements for transcription initiation.
Understanding transcriptional control mechanisms through post-transcriptional processing reveals how primary transcripts undergo extensive modifications to generate mature, functional mRNAs.
⚙️ The Transcriptional Orchestra: Gene Expression Command Systems
🎬 The RNA Processing Studio: From Transcript to Message
Spliceosome assembly occurs through stepwise recruitment of small nuclear ribonucleoproteins (snRNPs). U1 snRNP recognizes the 5' splice site (GU dinucleotide), while U2AF proteins bind the polypyrimidine tract and 3' splice site (AG dinucleotide). U2 snRNP associates with the branch point adenosine (located 20-50 nucleotides upstream of 3' splice site), followed by U4/U6•U5 tri-snRNP recruitment and dynamic rearrangements that create the catalytic core.
📌 Remember: Splice site consensus sequences: 5' splice site = GU-rich, 3' splice site = AG preceded by polypyrimidine tract. "GU-AG rule" governs 99.5% of human introns, with rare AU-AC introns (0.1%) spliced by U12-dependent spliceosome.
Alternative splicing affects 95% of multi-exon genes, generating 3-5 isoforms per gene on average. Exon skipping (40% of events) creates protein variants with deleted domains, while intron retention (25% of events) often introduces premature stop codons triggering nonsense-mediated decay. Tissue-specific splicing factors like NOVA, PTBP1, and RBFOX create cell-type-specific isoform patterns essential for neuronal function, muscle development, and immune responses.
| Splicing Pattern | Frequency | Mechanism | Functional Impact | Disease Examples |
|---|---|---|---|---|
| Exon Skipping | 40% | Weak splice sites | Domain deletion | Duchenne MD |
| Intron Retention | 25% | Incomplete removal | Premature termination | Thalassemia |
| Alternative 5'/3' | 20% | Competing sites | Length variation | CFTR mutations |
| Mutually Exclusive | 10% | Regulatory networks | Isoform switching | DSCAM diversity |
| Alternative Polyadenylation | 70% | Multiple poly(A) sites | 3'UTR variation | Cancer progression |
- 5' Cap Structure and Function
- 7-methylguanosine cap added co-transcriptionally
- Cap-binding complex (CBC) facilitates nuclear export
- eIF4E recognition enables ribosome recruitment
- Rate-limiting step for translation initiation
- mTOR pathway target for growth control
- Oncogene activation mechanism
- Cap methylation by RNMT/RAM complex
- N7-methylguanosine (first methylation)
- 2'-O-methylation of first transcribed nucleotide
- Innate immune evasion mechanism
💡 Master This: Nonsense-mediated decay (NMD) eliminates aberrant mRNAs containing premature termination codons through exon junction complex recognition. UPF1, UPF2, and UPF3 proteins detect downstream exon-exon junctions after translation termination, triggering mRNA degradation that prevents truncated protein accumulation.
3' end processing involves cleavage and polyadenylation at AAUAAA hexamer sequences located 10-30 nucleotides upstream of cleavage sites. Cleavage and polyadenylation specificity factor (CPSF) recognizes the polyadenylation signal, while cleavage stimulation factor (CstF) binds GU-rich or U-rich downstream elements. Poly(A) tail length (200-250 adenines) correlates with mRNA stability and translational efficiency.
Alternative polyadenylation affects 70% of human genes, creating isoforms with different 3'UTR lengths. Shorter 3'UTRs (common in cancer and proliferating cells) escape microRNA regulation and RNA-binding protein control, leading to increased protein expression. Global 3'UTR shortening represents a hallmark of cellular transformation and oncogene activation.
Understanding RNA processing mechanisms through translational control reveals how mature mRNAs undergo ribosomal decoding and protein synthesis with extraordinary fidelity and regulatory precision.
🎬 The RNA Processing Studio: From Transcript to Message
🏭 The Protein Assembly Line: Translation and Ribosomal Precision
Translation represents the final step of gene expression where genetic information becomes functional proteins. Understanding ribosomal mechanisms reveals how codon usage bias, ribosome profiling patterns, and translational control influence protein folding, cellular stress responses, and disease pathogenesis. Ribosome dysfunction causes ribosomopathies, while translational dysregulation drives cancer, neurodegeneration, and metabolic disorders.
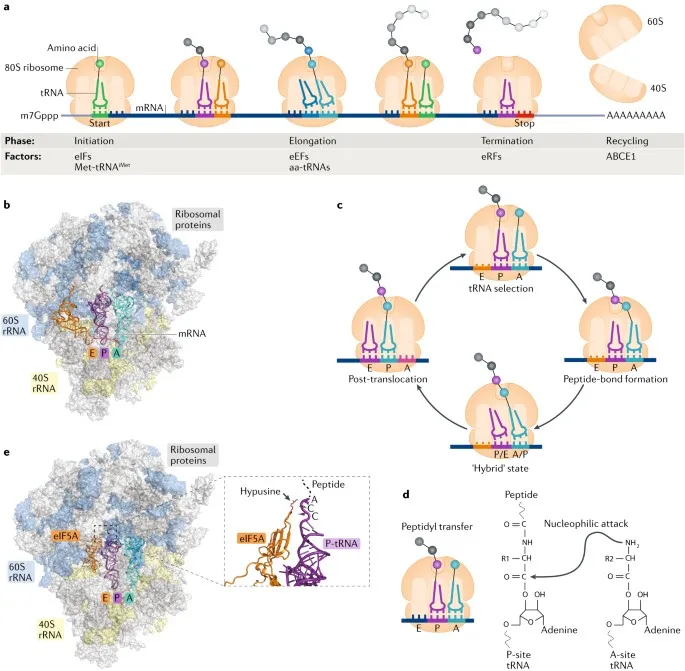
Ribosome structure consists of large (60S) and small (40S) subunits containing 4 rRNA species and 80+ ribosomal proteins. The small subunit contains 18S rRNA and 33 proteins, while the large subunit contains 28S, 5.8S, and 5S rRNAs with 47 proteins. Peptidyl transferase center within 28S rRNA catalyzes peptide bond formation through ribozyme activity, demonstrating RNA's catalytic capabilities in protein synthesis.
📌 Remember: Translation initiation factors: eIF4F (cap recognition), eIF4A (helicase), eIF4E (cap binding), eIF4G (scaffold). "Every Initiation Factor Facilitates Accurate Efficient Growth" - eIF4F-eIF4A-eIF4E-eIF4G coordinate 5' cap recognition and ribosome recruitment.
Genetic code properties include universality (shared across species), degeneracy (61 codons for 20 amino acids), and non-overlapping triplet reading. Wobble base pairing at third codon position allows single tRNAs to recognize multiple codons, while codon usage bias reflects tRNA abundance and translation efficiency. Synonymous codons are not functionally equivalent - optimal codons enhance translation speed and protein folding accuracy.
| Translation Phase | Key Factors | Energy Cost | Fidelity Mechanisms | Clinical Relevance |
|---|---|---|---|---|
| Initiation | eIF4F, eIF2, eIF3 | 1 GTP | Start codon selection | eIF2α phosphorylation |
| Elongation | eEF1A, eEF2 | 2 GTP/codon | Proofreading | Antibiotic targets |
| Termination | eRF1, eRF3 | 1 GTP | Stop codon recognition | Readthrough therapy |
| Recycling | ABCE1, eIF6 | 1 ATP | Subunit dissociation | Ribosome biogenesis |
- Translational Control Mechanisms
- 5'UTR secondary structures: stem-loops reduce ribosome binding
- Internal ribosome entry sites (IRES): cap-independent initiation
- Upstream open reading frames (uORFs): translation attenuation
- Present in 35-49% of human mRNAs
- Stress-responsive regulation
- Disease-associated mutations
- MicroRNA regulation: 3'UTR targeting and translation repression
- 1,000+ human miRNAs target 60% of genes
- Seed sequence complementarity (nucleotides 2-7)
- RISC complex mediated silencing
💡 Master This: Integrated stress response (ISR) coordinates translational control through eIF2α phosphorylation by 4 kinases: PERK (ER stress), PKR (viral infection), GCN2 (amino acid starvation), HRI (heme deficiency). Global translation decreases 75%, while ATF4 translation increases through uORF-mediated mechanism, activating stress response genes.
Ribosome heterogeneity creates specialized ribosomes with distinct protein compositions and rRNA modifications. Ribosomal protein mutations cause tissue-specific phenotypes despite ubiquitous ribosome function - RPS19 mutations cause Diamond-Blackfan anemia, RPS26 mutations affect intellectual development, and RPL5 mutations cause T-cell immunodeficiency. Ribosomal RNA pseudouridylation (100+ sites) and 2'-O-methylation (100+ sites) fine-tune translation fidelity and mRNA selectivity.
Post-translational modifications begin co-translationally as nascent polypeptides emerge from ribosomal exit tunnel. Signal recognition particle (SRP) targets secretory proteins to endoplasmic reticulum, while nascent polypeptide-associated complex (NAC) prevents premature folding. Ribosome-associated quality control eliminates stalled ribosomes through ribosome-associated protein quality control (RQC) and no-go decay pathways.
Understanding translational mechanisms through epigenetic regulation reveals how DNA methylation and histone modifications create heritable gene expression changes without sequence alterations.
🏭 The Protein Assembly Line: Translation and Ribosomal Precision
🎨 The Epigenetic Canvas: Beyond the DNA Sequence
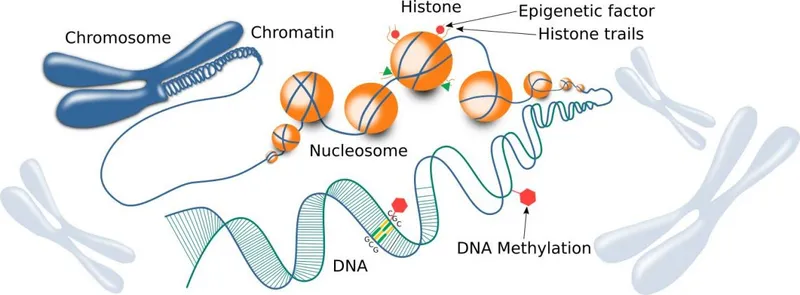
DNA methylation occurs primarily at CpG dinucleotides, which are 4-fold underrepresented in mammalian genomes due to spontaneous deamination of 5-methylcytosine to thymine. CpG islands (500-2000 base pairs with >50% GC content) remain unmethylated in normal cells and overlap 70% of gene promoters. DNA methyltransferases (DNMT1, DNMT3A, DNMT3B) establish and maintain methylation patterns, while TET enzymes catalyze active demethylation through 5-hydroxymethylcytosine intermediates.
📌 Remember: Histone modification code: H3K4me3 (active promoters), H3K27ac (active enhancers), H3K36me3 (gene bodies), H3K27me3 (Polycomb repression), H3K9me3 (heterochromatin). "Happy Histone Marks Make Gene Expression" - H3K4me3-H3K27ac-H3K36me3 mark active transcription.
Chromatin remodeling complexes use ATP hydrolysis to slide, eject, or exchange nucleosomes. SWI/SNF complexes (BAF and PBAF) facilitate transcriptional activation, while ISWI complexes promote chromatin assembly and gene repression. CHD complexes regulate development and differentiation, with CHD7 mutations causing CHARGE syndrome and CHD8 mutations associated with autism spectrum disorders.
| Epigenetic Mark | Genomic Location | Function | Enzymes | Disease Associations |
|---|---|---|---|---|
| H3K4me3 | Promoters | Transcription activation | SET1A/B, MLL | Leukemia |
| H3K27me3 | Developmental genes | Polycomb silencing | EZH2 | Cancer |
| H3K9me3 | Heterochromatin | Stable repression | SUV39H1/2 | Aging |
| H3K36me3 | Gene bodies | Transcription elongation | SETD2 | Renal cancer |
| DNA methylation | CpG islands | Gene silencing | DNMTs | Imprinting disorders |
- Polycomb and Trithorax Systems
- Polycomb Repressive Complex 1 (PRC1): H2AK119 ubiquitination
- Polycomb Repressive Complex 2 (PRC2): H3K27 trimethylation
- Trithorax complexes: H3K4 methylation and transcriptional memory
- MLL1-4 methyltransferases
- Leukemia fusion proteins (MLL-AF4, MLL-ENL)
- Developmental gene regulation
- Bivalent chromatin domains: H3K4me3 + H3K27me3
- Poised developmental genes in stem cells
- Resolution during differentiation
- Cancer stem cell maintenance
💡 Master This: Epigenetic clocks measure biological aging through DNA methylation patterns at 300-500 CpG sites. Accelerated epigenetic aging predicts mortality risk, age-related diseases, and response to interventions. Reprogramming factors (Oct4, Sox2, Klf4, c-Myc) can reverse epigenetic age, suggesting therapeutic potential for aging-related disorders.
X-chromosome inactivation demonstrates epigenetic dosage compensation through XIST long non-coding RNA coating one X chromosome in female cells. Random inactivation creates mosaic expression patterns, while skewed inactivation (>80% bias) occurs in 20% of women and influences X-linked disease severity. Reactivation of silenced X chromosome genes contributes to autoimmune diseases and cancer progression.
Genomic imprinting affects 150+ genes clustered in imprinted domains regulated by differentially methylated regions (DMRs). Parent-of-origin expression creates functional haploidy for imprinted genes, making them sensitive to mutations and epimutations. Beckwith-Wiedemann syndrome results from 11p15.5 imprinting defects, while Silver-Russell syndrome involves growth restriction from similar molecular mechanisms.
Understanding epigenetic regulation through genomic technologies reveals how high-throughput sequencing and genome editing transform molecular diagnosis and therapeutic development.
🎨 The Epigenetic Canvas: Beyond the DNA Sequence
🔬 The Genomic Revolution: Technologies Transforming Medicine
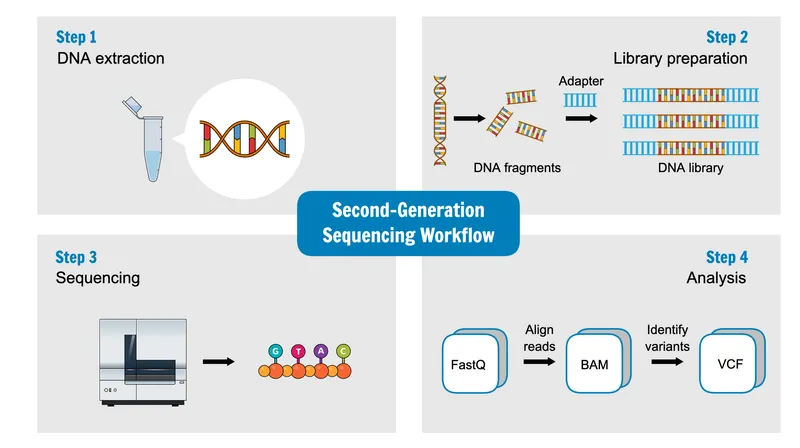
Next-generation sequencing (NGS) platforms achieve throughput of 1-6 terabases per run with error rates below 0.1%. Illumina sequencing-by-synthesis uses reversible terminators and fluorescent detection to sequence millions of clusters simultaneously. Library preparation involves DNA fragmentation, adapter ligation, and PCR amplification, while bioinformatics pipelines perform quality control, alignment, variant calling, and annotation.
📌 Remember: NGS applications: WGS (whole genome sequencing), WES (whole exome sequencing), RNA-seq (transcriptome), ChIP-seq (protein-DNA interactions), ATAC-seq (chromatin accessibility). "Next Generation Sequencing Widely Examines RNA, Chromatin, And Transcription".
CRISPR-Cas9 genome editing achieves precise DNA modifications through programmable nucleases. Guide RNAs direct Cas9 endonuclease to target sequences adjacent to protospacer adjacent motifs (PAMs), creating double-strand breaks repaired by non-homologous end joining (NHEJ) or homology-directed repair (HDR). Base editors enable single nucleotide changes without double-strand breaks, while prime editors allow insertions, deletions, and replacements with minimal off-target effects.
| Technology | Resolution | Throughput | Applications | Clinical Use |
|---|---|---|---|---|
| Whole Genome Sequencing | Single nucleotide | 30-50x coverage | Rare diseases, cancer | Diagnostic testing |
| RNA sequencing | Transcript level | 20-100 million reads | Expression profiling | Cancer classification |
| ChIP-sequencing | 100-200 bp | 10-50 million reads | Protein-DNA binding | Epigenetic analysis |
| ATAC-sequencing | Single nucleotide | 25-75 million reads | Chromatin accessibility | Regulatory mapping |
| Single-cell RNA-seq | Cell-specific | 1,000-10,000 cells | Cell type identification | Developmental biology |
- Single-Cell Technologies
- Single-cell RNA sequencing: cell-type-specific expression profiles
- Single-cell ATAC-seq: chromatin accessibility at cellular resolution
- Single-cell multiomics: simultaneous RNA and chromatin profiling
- 10,000-100,000 cells per experiment
- Developmental trajectories and cell fate decisions
- Disease heterogeneity and therapeutic resistance
- Spatial transcriptomics: tissue architecture preservation
- 10-100 micrometer resolution
- Pathological processes in tissue context
- Tumor microenvironment characterization
💡 Master This: Liquid biopsies detect circulating tumor DNA (ctDNA) in plasma at concentrations as low as 0.01% of total cell-free DNA. Digital PCR and NGS-based approaches identify actionable mutations, monitor treatment response, and detect minimal residual disease with sensitivity exceeding tissue biopsies for metastatic cancers.
Genome-wide association studies (GWAS) identify disease-associated variants by comparing allele frequencies between cases and controls. Polygenic risk scores aggregate effects of thousands of variants to predict disease susceptibility, drug responses, and quantitative traits. Mendelian randomization uses genetic variants as instrumental variables to infer causal relationships between exposures and outcomes, avoiding confounding in observational studies.
Functional genomics approaches include CRISPR screens for gene function, massively parallel reporter assays for regulatory elements, and protein-protein interaction mapping through proximity labeling. Organoid models and tissue engineering enable functional validation of genomic findings in physiologically relevant systems, bridging molecular discoveries and clinical applications.
Understanding genomic technologies through precision medicine applications reveals how molecular profiling guides individualized treatment strategies and improves patient outcomes across diverse medical specialties.
🔬 The Genomic Revolution: Technologies Transforming Medicine
🎯 The Precision Medicine Arsenal: Molecular Tools for Clinical Mastery
Tumor molecular profiling identifies actionable mutations in 300+ cancer genes through comprehensive genomic profiling. Microsatellite instability (MSI) testing predicts immunotherapy response across tumor types, while homologous recombination deficiency (HRD) scores guide PARP inhibitor therapy. Tumor mutational burden (TMB) correlates with checkpoint inhibitor efficacy, and PD-L1 expression stratifies patients for anti-PD-1/PD-L1 treatments.
📌 Remember: Precision medicine biomarkers: Predictive (treatment response), Prognostic (disease outcome), Diagnostic (disease presence), Pharmacodynamic (drug effect), Safety (toxicity risk). "Precision Predicts Prognosis, Diagnoses Disease, Pharmacology, Safety".
Pharmacogenomics implementation reduces adverse drug reactions and improves therapeutic outcomes through pre-emptive testing of drug-metabolizing enzymes. CYP2D6 variants affect 25% of medications including codeine, tamoxifen, and antidepressants. CYP2C19 polymorphisms influence clopidogrel activation and proton pump inhibitor metabolism. DPYD deficiency increases 5-fluorouracil toxicity risk by 50-fold.
| Biomarker Category | Clinical Application | Testing Method | Actionable Findings | Implementation Rate |
|---|---|---|---|---|
| Germline variants | Hereditary cancer | Panel sequencing | BRCA1/2, Lynch syndrome | 15-25% |
| Somatic mutations | Targeted therapy | NGS panels | EGFR, KRAS, BRAF | 60-80% |
| Gene expression | Prognosis | RT-PCR, RNA-seq | Oncotype DX, PAM50 | 30-50% |
| Protein markers | Immunotherapy | IHC, flow cytometry | PD-L1, MSI | 40-70% |
| Pharmacogenomics | Drug selection | Genotyping arrays | CYP variants | 5-15% |
| %%{init: {'flowchart': {'htmlLabels': true}}}%% | ||||
| flowchart TD |
Start["🩺 Presentation
• Patient intake• Clinical assessment"]
Profile["🔬 Molecular Profile
• Genetic sequencing• Tumor analysis"]
Decision1["📋 Actionable?
• Review findings• Identify variants"]
Standard["💊 Standard Care
• Guideline therapy• Baseline protocol"]
Adjustment["💊 Drug Adjustment
• PGx optimization• Dose modification"]
Immuno["💊 Immunotherapy
• PD-L1 markers• Immune activation"]
Targeted["💊 Targeted Therapy
• Mutation specific• Precision medicine"]
Monitor["👁️ Monitoring
• Response check• Follow-up imaging"]
Decision2["📋 Treatment Resp
• Evaluate efficacy• Check progress"]
Resistance["🔬 Resist. Testing
• Biopsy at relapse• Search escape mech"]
Continue["✅ Continue Rx
• Maintain therapy• Stable disease"]
Start --> Profile Profile --> Decision1
Decision1 -->|Negative| Standard Decision1 -->|PGx Var| Adjustment Decision1 -->|Biomarker+| Immuno Decision1 -->|Mutation| Targeted
Standard --> Monitor Adjustment --> Monitor Immuno --> Monitor Targeted --> Monitor
Monitor --> Decision2 Decision2 -->|Progression| Resistance Decision2 -->|Response| Continue
Resistance -.-> Profile
style Start fill:#F7F5FD, stroke:#F0EDFA, stroke-width:1.5px, rx:12, ry:12, color:#6B21A8 style Profile fill:#FFF7ED, stroke:#FFEED5, stroke-width:1.5px, rx:12, ry:12, color:#C2410C style Decision1 fill:#FEF8EC, stroke:#FBECCA, stroke-width:1.5px, rx:12, ry:12, color:#854D0E style Standard fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534 style Adjustment fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534 style Immuno fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534 style Targeted fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534 style Monitor fill:#EEFAFF, stroke:#DAF3FF, stroke-width:1.5px, rx:12, ry:12, color:#0369A1 style Decision2 fill:#FEF8EC, stroke:#FBECCA, stroke-width:1.5px, rx:12, ry:12, color:#854D0E style Resistance fill:#FFF7ED, stroke:#FFEED5, stroke-width:1.5px, rx:12, ry:12, color:#C2410C style Continue fill:#F6F5F5, stroke:#E7E6E6, stroke-width:1.5px, rx:12, ry:12, color:#525252
> ⭐ **Clinical Pearl**: **Circulating tumor DNA (ctDNA)** monitoring detects **disease recurrence** **2-12 months** earlier than **imaging** in **colorectal**, **breast**, and **lung cancers**. **Molecular residual disease** testing guides **adjuvant therapy** decisions and **surveillance strategies**, with **positive ctDNA** conferring **3-10 fold** increased **recurrence risk**.
* **Rare Disease Genomics**
- **Whole exome sequencing** achieves **25-30%** diagnostic yield
- **Whole genome sequencing** increases yield to **35-45%**
- **Structural variant** detection identifies **additional 5-10%**
+ **Copy number variants** (CNVs)
+ **Balanced translocations**
+ **Complex rearrangements**
- **Functional studies** validate **variants of uncertain significance**
+ **In vitro** assays for **enzyme activity**
+ **Cell-based** models for **pathway function**
+ **Animal models** for **phenotype validation**
> 💡 **Master This**: **Polygenic risk scores (PRS)** integrate **effects** of **thousands** to **millions** of **genetic variants** to predict **disease risk** and **treatment response**. **High PRS** individuals (**top 5%**) have **3-5 fold** increased risk for **coronary artery disease**, **diabetes**, and **breast cancer**, enabling **early intervention** and **enhanced screening** strategies.
**Liquid biopsy applications** extend beyond **oncology** to **prenatal testing**, **organ transplant monitoring**, and **infectious disease** detection. **Cell-free fetal DNA** testing achieves **>99% sensitivity** for **trisomy 21** with **<0.1% false positive rate**. **Donor-derived cell-free DNA** predicts **organ rejection** before **clinical symptoms** or **histological changes**. **Pathogen cell-free DNA** enables **culture-independent** diagnosis of **sepsis** and **meningitis**.
**Therapeutic genome editing** advances include **in vivo CRISPR** for **inherited diseases**, **CAR-T cell** engineering for **cancer immunotherapy**, and **base editing** for **sickle cell disease**. **CTX001** therapy uses **CRISPR** to **reactivate fetal hemoglobin** in **β-thalassemia** and **sickle cell disease** patients. **Leber congenital amaurosis** treatment with **EDIT-101** represents **first in vivo** CRISPR trial for **inherited blindness**.
**Implementation challenges** include **variant interpretation**, **clinical decision support**, **cost-effectiveness**, and **health disparities**. **Genomic medicine** requires **multidisciplinary teams**, **bioinformatics infrastructure**, and **continuous education** to translate **molecular discoveries** into **improved patient outcomes** while ensuring **equitable access** to **precision medicine** benefits.
🎯 The Precision Medicine Arsenal: Molecular Tools for Clinical Mastery
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app
























