Biochemical Techniques

On this page
🔬 The Biochemical Detective Arsenal: Molecular Investigation Mastery
Biochemical techniques transform invisible molecular events into diagnostic certainty, giving clinicians the power to detect disease at its chemical roots. You'll master how each method-from spectrophotometry to immunoassays-exploits specific molecular properties, learn to select the right tool for each clinical question, and understand how these techniques integrate across diagnostic pathways. By connecting detection principles to real-world applications, you'll build the analytical framework needed to interpret lab results with precision and confidence.
Foundation Principles: The Molecular Toolkit
Biochemical techniques operate on four core principles that enable molecular detection and quantification:
- Separation Science
- Size-based separation: molecules 10-1000 kDa range
- Charge-based separation: pH 3.0-11.0 operational range
- Electrophoretic mobility: 1-15 cm²/V·s typical values
- Resolution power: distinguishes <5% molecular weight differences
- Detection Methods
- Spectroscopic detection: wavelengths 200-800 nm
- Fluorescence sensitivity: detects femtomolar (10⁻¹⁵ M) concentrations
- Signal-to-noise ratio: >100:1 for reliable quantification
- Dynamic range: 4-6 orders of magnitude
📌 Remember: SPEC - Separation, Purification, Electrophoresis, Chromatography - the four pillars supporting all biochemical analysis
| Technique Category | Separation Basis | Resolution Range | Sample Volume | Detection Limit | Clinical Application |
|---|---|---|---|---|---|
| Electrophoresis | Charge/Size | 1-2% difference | 1-50 μL | ng-μg range | Protein analysis |
| Chromatography | Polarity/Size | 0.1-1% difference | μL-mL range | pg-ng range | Drug monitoring |
| Spectrophotometry | Light absorption | 1-5% difference | 10-1000 μL | μg-mg range | Enzyme assays |
| Mass Spectrometry | Mass/charge | 0.01% difference | nL-μL range | fg-pg range | Metabolomics |
| Immunoassays | Antigen-antibody | High specificity | 1-100 μL | pg-ng range | Hormone levels |
💡 Master This: Every biochemical technique combines separation + detection + quantification - understanding this trinity predicts optimal method selection for any clinical scenario
The precision of biochemical techniques enables detection of molecular changes that occur hours before clinical symptoms appear, transforming these methods into predictive diagnostic tools that guide early intervention strategies.
🔬 The Biochemical Detective Arsenal: Molecular Investigation Mastery
⚡ Molecular Precision Engineering: Detection Mechanism Mastery
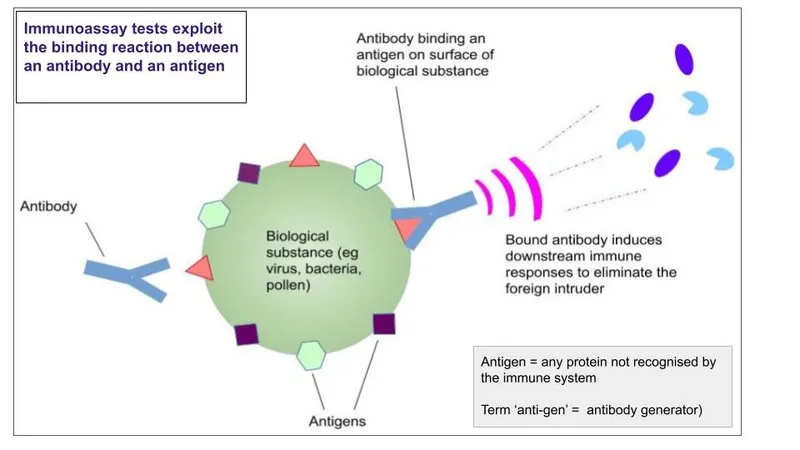
Signal Transduction Pathways: From Molecule to Measurement
Detection mechanisms operate through three amplification stages that enhance sensitivity:
- Primary Recognition
- Antibody-antigen binding: Kd values 10⁻⁹ to 10⁻¹² M
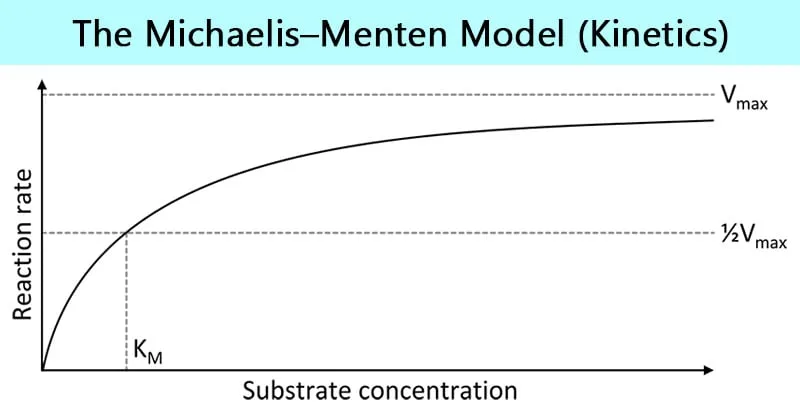
- Enzyme-substrate interaction: Km values 10⁻⁶ to 10⁻³ M
- Binding specificity: >99% for target molecules
- Cross-reactivity: <1% with similar structures
- Signal Amplification
- Enzymatic amplification: 10³-10⁶ fold signal enhancement
- Fluorescence amplification: 10²-10⁴ fold intensity increase
- Quantum yield: 0.1-0.9 efficiency range
- Stokes shift: 20-100 nm wavelength difference
📌 Remember: BIND-AMP-DETECT - Binding specificity, Interaction kinetics, Noise reduction, Detection sensitivity, Amplification efficiency, Measurement precision, Processing algorithms, Data interpretation, Error correction, Calibration standards, Time resolution
| Detection Method | Signal Type | Sensitivity Range | Response Time | Linear Range | Interference Level |
|---|---|---|---|---|---|
| Colorimetric | Absorbance change | μM-mM | 1-30 minutes | 2-3 orders | Moderate |
| Fluorometric | Emission intensity | nM-μM | Seconds-minutes | 3-4 orders | Low |
| Chemiluminescent | Light emission | pM-nM | 1-60 seconds | 4-5 orders | Very low |
| Electrochemical | Current/voltage | nM-mM | Milliseconds | 3-4 orders | Variable |
| Radiometric | Radiation count | fM-pM | Minutes-hours | 5-6 orders | Minimal |
💡 Master This: Detection sensitivity determines clinical utility - femtomolar detection enables early disease markers while micromolar sensitivity suits metabolic monitoring
Understanding detection mechanisms reveals why certain techniques excel in specific clinical applications, enabling optimal method selection based on required sensitivity, specificity, and turnaround time for patient care decisions.
⚡ Molecular Precision Engineering: Detection Mechanism Mastery
🎯 Pattern Recognition Protocols: Clinical Application Frameworks
Clinical Decision Matrix: Technique Selection Framework
-
Protein Analysis Patterns
- See: Suspected enzyme deficiency → Think: Spectrophotometric enzyme assays
- See: Protein expression changes → Think: Western blot analysis
- Sensitivity requirement: nanogram detection for Western blots
- Quantification range: 2-3 orders of magnitude linear response
- See: Protein-protein interactions → Think: Immunoprecipitation techniques
- Binding specificity: >95% target recognition required
-
Nucleic Acid Analysis Patterns
- See: Gene expression quantification → Think: RT-qPCR analysis
- See: DNA fragment analysis → Think: Gel electrophoresis
- Resolution power: distinguishes 10-50 bp size differences
- Detection limit: 1-10 ng DNA minimum requirement
- See: Mutation detection → Think: DNA sequencing methods
- Accuracy requirement: >99.9% base calling precision
📌 Remember: MATCH-TECH - Molecular target, Analytical sensitivity, Turnaround time, Cost considerations, Handling requirements, Technical expertise, Equipment availability, Clinical urgency, High-throughput needs
| Clinical Question | Primary Technique | Sensitivity Needed | Turnaround Time | Sample Volume | Cost Category |
|---|---|---|---|---|---|
| Hormone levels | ELISA/RIA | pg/mL range | 2-4 hours | 50-200 μL | Moderate |
| Enzyme activity | Spectrophotometry | U/L range | 15-30 minutes | 10-50 μL | Low |
| Protein expression | Western blot | ng range | 4-8 hours | 20-100 μg | High |
| Gene expression | RT-qPCR | Copy number | 2-3 hours | 1-10 μg RNA | Moderate |
| Metabolite profiling | LC-MS/MS | ng/mL range | 1-2 hours | 100-500 μL | High |
💡 Master This: Sample volume limitations often dictate technique selection - pediatric samples (<100 μL) require high-sensitivity methods while research samples allow volume-intensive procedures
Diagnostic Hierarchy: Screening to Confirmation
- Screening Level (High throughput, moderate sensitivity)
- Spectrophotometric assays: 96-384 well plate formats
- Immunoassays: automated platforms processing 200+ samples/hour
- Confirmatory Level (High specificity, precise quantification)
- Mass spectrometry: definitive identification with molecular fingerprinting
- DNA sequencing: base-level resolution for genetic variants
- Research Level (Maximum sensitivity, comprehensive analysis)
- Proteomics platforms: thousands of proteins simultaneously
- Metabolomics approaches: hundreds of metabolites in single analysis
Pattern recognition frameworks enable rapid technique selection that optimizes diagnostic accuracy while managing resource utilization, ensuring appropriate analytical approaches for diverse clinical scenarios requiring biochemical investigation.
🎯 Pattern Recognition Protocols: Clinical Application Frameworks
🔍 Analytical Discrimination Mastery: Technique Comparison Architecture
Performance Discrimination Matrix: Quantitative Comparisons
| Technique | Sensitivity Range | Specificity (%) | Throughput/Day | Setup Time | Cost/Sample | Expertise Level |
|---|---|---|---|---|---|---|
| ELISA | pg-ng/mL | 95-99% | 200-500 | 30 minutes | $5-15 | Moderate |
| Western Blot | ng-μg | 90-95% | 10-20 | 2-4 hours | $20-50 | High |
| RT-qPCR | 1-100 copies | 98-99.9% | 50-100 | 1 hour | $10-25 | High |
| Mass Spec | fg-pg | 99-99.9% | 100-300 | 2-3 hours | $25-75 | Very High |
| Spectrophotometry | μg-mg/mL | 80-95% | 500-1000 | 5 minutes | $1-5 | Low |
| Flow Cytometry | Single cell | 95-99% | 1000-5000 cells/sec | 30 minutes | $15-30 | High |
Discrimination Decision Trees: Method Selection Logic
-
Sensitivity-Driven Selection
- Ultra-high sensitivity (femtomolar): Mass spectrometry, single-molecule techniques
- Detection limit: 10⁻¹⁵ to 10⁻¹⁸ M concentration range
- Signal amplification: 10⁶-10⁹ fold enhancement possible
- High sensitivity (picomolar): Fluorescence, chemiluminescence
- Detection limit: 10⁻¹² to 10⁻⁹ M concentration range
- Background noise: <1% of signal intensity
- Moderate sensitivity (nanomolar): ELISA, radioimmunoassays
- Detection limit: 10⁻⁹ to 10⁻⁶ M concentration range
- Coefficient of variation: <10% for reliable quantification
- Ultra-high sensitivity (femtomolar): Mass spectrometry, single-molecule techniques
-
Throughput-Driven Selection
- High-throughput screening: >1000 samples/day capacity
- Automation level: >90% hands-off operation
- Processing time: <5 minutes per sample
- Medium-throughput analysis: 100-500 samples/day capacity
- Manual intervention: 20-50% of workflow
- Processing time: 10-30 minutes per sample
- Low-throughput research: <50 samples/day capacity
- Manual operation: >70% hands-on requirement
- Processing time: 1-4 hours per sample
- High-throughput screening: >1000 samples/day capacity
⭐ Clinical Pearl: Cost-per-result analysis reveals true economic efficiency - high initial equipment costs ($100K-500K) may provide lower per-sample costs ($1-5) compared to low-cost methods ($10-20/sample) for high-volume applications
💡 Master This: Analytical validation requirements determine technique acceptability - clinical diagnostics require >95% accuracy while research applications may accept 80-90% precision for exploratory studies
Quality Metrics: Performance Benchmarking
- Precision Metrics
- Intra-assay variation: <5% coefficient of variation
- Inter-assay variation: <10% coefficient of variation
- Day-to-day variation: <15% coefficient of variation
- Accuracy Metrics
- Recovery studies: 90-110% acceptable range
- Linearity correlation: r² >0.99 for quantitative methods
- Detection limit validation: signal-to-noise >3:1 minimum
Systematic discrimination frameworks enable objective technique selection based on quantitative performance criteria, ensuring optimal analytical approaches that balance sensitivity, throughput, cost, and accuracy requirements for specific clinical applications.
🔍 Analytical Discrimination Mastery: Technique Comparison Architecture
⚖️ Treatment Protocol Engineering: Evidence-Based Method Optimization
Optimization Algorithm: Systematic Method Development
-
Critical Parameter Identification
- Buffer systems: pH optimization ±0.1 units for enzyme stability
- Temperature control: ±1°C precision for reaction consistency
- Enzyme activity: Q₁₀ = 2-3 for most biochemical reactions
- Denaturation risk: >5°C above optimum causes >50% activity loss
- Ionic strength: ±10 mM precision for electrophoretic separation
- Migration reproducibility: <2% variation in mobility
- Resolution maintenance: >1.5 peak separation factor
-
Interference Management
- Matrix effects: <10% signal suppression acceptable
- Cross-reactivity: <1% with structurally similar compounds
- Specificity testing: >20 related compounds evaluation
- Selectivity coefficient: >100:1 target vs. interferent
📌 Remember: OPTIMIZE-VALIDATE - Objective setting, Parameter identification, Testing design, Improvement tracking, Method comparison, Interference testing, Zero-drift monitoring, Error analysis, Validation studies, Accuracy assessment, Linearity testing, Inter-lab comparison, Documentation, Approval process, Training, Evaluation
| Optimization Parameter | Target Range | Acceptable Variation | Impact on Performance | Monitoring Frequency | Correction Action |
|---|---|---|---|---|---|
| pH Control | ±0.05 units | ±0.1 units | 5-20% activity change | Every batch | Buffer adjustment |
| Temperature | ±0.5°C | ±1.0°C | 10-30% rate change | Continuous | Incubator calibration |
| Reagent Concentration | ±2% | ±5% | 2-15% signal change | Weekly | Fresh preparation |
| Incubation Time | ±30 seconds | ±2 minutes | 5-25% completion | Each assay | Timer verification |
| Sample Volume | ±1% | ±2% | 1-10% precision | Each pipetting | Calibration check |
💡 Master This: Validation hierarchy follows analytical → clinical → regulatory progression - each level requires increasing evidence standards with >95% success rates for clinical implementation
Evidence Integration: Performance Validation Framework
- Analytical Validation
- Precision studies: n≥20 replicates for statistical power
- Accuracy assessment: certified reference materials comparison
- Bias evaluation: <5% deviation from true values
- Uncertainty calculation: 95% confidence intervals reporting
- Clinical Validation
- Patient sample correlation: n≥100 clinical specimens
- Reference method comparison: Deming regression analysis
- Correlation coefficient: r>0.95 for method equivalence
- Clinical concordance: >90% diagnostic agreement
- Implementation Validation
- Multi-operator studies: ≥3 technologists performance
- Long-term stability: 6-month performance monitoring
- Control limits: ±2 standard deviations from mean
- Trend analysis: <1%/month drift acceptable
Evidence-based optimization transforms biochemical techniques from variable research tools into reliable clinical platforms that deliver consistent, accurate results supporting confident diagnostic decisions and optimal patient care outcomes.
⚖️ Treatment Protocol Engineering: Evidence-Based Method Optimization
🔗 Multi-System Integration Hub: Advanced Biochemical Networks

Integration Architecture: Synergistic Platform Design
-
Multi-Omics Integration
- Genomics + Proteomics: DNA-to-protein correlation analysis
- Data integration: >10,000 genes with >50,000 proteins simultaneously
- Correlation algorithms: machine learning pattern recognition
- Metabolomics + Transcriptomics: pathway-level system analysis
- Metabolite coverage: >1,000 compounds per analysis
- Pathway mapping: >300 biochemical pathways integration
- Temporal resolution: minutes to hours dynamic tracking
- Genomics + Proteomics: DNA-to-protein correlation analysis
-
Automated Workflow Integration
- Sample-to-result automation: >95% hands-off processing
- Quality control integration: real-time monitoring across platforms
- Error detection: <1% false positive rate
- System uptime: >98% operational availability
- Throughput capacity: >1,000 samples/day multi-technique processing
📌 Remember: INTEGRATE-ANALYZE-DECIDE - Input standardization, Network protocols, Throughput optimization, Error handling, Guality monitoring, Real-time processing, Automated workflows, Technology convergence, End-to-end tracking, Algorithmic analysis, Network security, Alert systems, Long-term storage, Yield optimization, Zero-downtime design, Emergency protocols, Data validation, Expert systems, Clinical integration, Information delivery, Decision support, Evaluation metrics
| Integration Level | Technique Combination | Data Complexity | Analysis Time | Clinical Value | Implementation Cost |
|---|---|---|---|---|---|
| Basic Dual | 2 complementary methods | 10²-10³ data points | 2-4 hours | Moderate | $50K-100K |
| Multi-Platform | 3-5 analytical systems | 10⁴-10⁵ data points | 4-8 hours | High | $200K-500K |
| Omics Integration | 5-10 comprehensive platforms | 10⁶-10⁷ data points | 8-24 hours | Very High | $1M-5M |
| AI-Enhanced | Full laboratory automation | 10⁷-10⁸ data points | Real-time | Transformative | $5M-20M |
💡 Master This: Integration success depends on standardized data formats - platforms using common ontologies achieve >90% interoperability while proprietary systems require custom interfaces reducing efficiency
Advanced Applications: Cutting-Edge Integration Examples
-
Precision Medicine Platforms
- Pharmacogenomics integration: genetic variants + drug metabolism analysis
- Variant detection: >1,000 pharmacogenes simultaneous screening
- Drug interaction prediction: >95% accuracy for major medications
- Biomarker discovery: multi-omics pattern identification
- Discovery success: >80% validation rate for integrated approaches
- Time-to-biomarker: 50% reduction compared to single-technique studies
- Pharmacogenomics integration: genetic variants + drug metabolism analysis
-
Real-Time Monitoring Systems
- Point-of-care integration: bedside multi-parameter analysis
- Response time: <15 minutes for critical care decisions
- Accuracy maintenance: >95% compared to central laboratory
- Continuous monitoring: wearable biosensors with laboratory correlation
- Sampling frequency: every 1-60 minutes depending on analyte
- Predictive capability: 6-24 hours advance warning for clinical changes
- Point-of-care integration: bedside multi-parameter analysis
Future Integration Trends: Next-Generation Platforms
- Artificial Intelligence Integration
- Pattern recognition: deep learning algorithms for complex data
- Predictive analytics: outcome prediction from biochemical patterns
- Prediction accuracy: >85% for major clinical outcomes
- False positive rate: <5% for critical alerts
- Miniaturization Integration
- Lab-on-chip platforms: complete workflows on cm² devices
- Portable integration: field-deployable multi-technique systems
- Power consumption: <10W for battery operation
- Analysis time: <30 minutes for complex panels
Multi-system integration transforms biochemical techniques from isolated analytical tools into comprehensive diagnostic ecosystems that provide unprecedented molecular insights, enabling precision medicine approaches that optimize patient outcomes through integrated data analysis.
🔗 Multi-System Integration Hub: Advanced Biochemical Networks
🎯 Clinical Mastery Command Center: Rapid-Fire Diagnostic Arsenal
Essential Arsenal: Critical Numbers for Instant Recall
📌 Remember: RAPID-FIRE-RECALL - Reference ranges, Analytical limits, Performance specs, Interference factors, Detection thresholds, False positive rates, Incubation times, Reagent stability, Equipment specs, Recovery percentages, Error rates, Calibration intervals, Accuracy targets, Linearity ranges, Limit specifications
| Technique | Detection Limit | Linear Range | Precision (CV%) | Turnaround Time | Sample Volume |
|---|---|---|---|---|---|
| ELISA | 1-10 pg/mL | 3-4 orders | <5% | 2-4 hours | 50-200 μL |
| Western Blot | 1-10 ng | 2-3 orders | <10% | 4-8 hours | 20-100 μg |
| RT-qPCR | 1-100 copies | 6-8 orders | <2% | 2-3 hours | 1-10 μg RNA |
| Mass Spec | 0.1-1 pg/mL | 4-5 orders | <3% | 1-2 hours | 100-500 μL |
| Spectrophotometry | 1-10 μg/mL | 2-3 orders | <3% | 5-30 minutes | 10-100 μL |
💡 Master This: Sample volume limitations in pediatrics (<100 μL total) restrict technique options to high-sensitivity methods requiring femtomolar-picomolar detection capabilities
Rapid Decision Matrix: Instant Technique Selection
-
Emergency Scenarios (Results needed <1 hour)
- Cardiac markers: Point-of-care troponin (15 minutes)
- Glucose monitoring: Glucometer (30 seconds)
- Blood gases: Automated analyzer (2 minutes)
-
Routine Diagnostics (Results needed 2-24 hours)
- Hormone panels: Automated immunoassays (2-4 hours)
- Protein analysis: Turbidimetry/nephelometry (30 minutes)
- Enzyme activities: Kinetic spectrophotometry (15 minutes)
-
Specialized Testing (Results needed 1-7 days)
- Genetic analysis: DNA sequencing (24-48 hours)
- Proteomics: Mass spectrometry (4-8 hours)
- Metabolomics: LC-MS/MS (2-4 hours)
📌 Remember: TIME-SENSITIVITY-ACCURACY triangle determines technique selection - emergency situations prioritize speed, research applications prioritize accuracy, screening programs prioritize throughput
Quality Control Commandments: Non-Negotiable Standards
-
Calibration Requirements
- Daily: Spectrophotometers, pH meters, pipettes
- Weekly: Automated analyzers, centrifuges, incubators
- Monthly: Balances, thermometers, timers
- Quarterly: Mass spectrometers, HPLC systems
-
Control Limits
- ±2 SD: Warning limits for investigation
- ±3 SD: Control limits for rejection
- Westgard rules: 1₃ₛ, 2₂ₛ, R₄ₛ, 4₁ₛ, 10ₓ for quality monitoring
⭐ Clinical Pearl: Control failure requires immediate investigation - >5% of results outside control limits indicates systematic error requiring method shutdown until resolution
💡 Master This: Proficiency testing requires >80% acceptable performance for laboratory accreditation - failure rates >20% trigger regulatory investigation and potential license suspension
Clinical mastery transforms biochemical techniques from complex procedures into rapid-fire diagnostic tools that deliver precise, timely results supporting optimal patient care decisions across emergency, routine, and specialized clinical scenarios.
🎯 Clinical Mastery Command Center: Rapid-Fire Diagnostic Arsenal
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more
Have doubts about this lesson?
Ask Rezzy, your AI Study Mate, to explain anything you didn't understand
Everything you need for NEET-PG prep
Get full Oncourse access with lessons, practice questions, flashcards and AI study tools.
Scan to download app
























