Molecular Genetics — MCQs

On this page
An adult tall male presents with a long arm span, pectus excavatum, and cardiac abnormalities. What is the most likely defective protein?
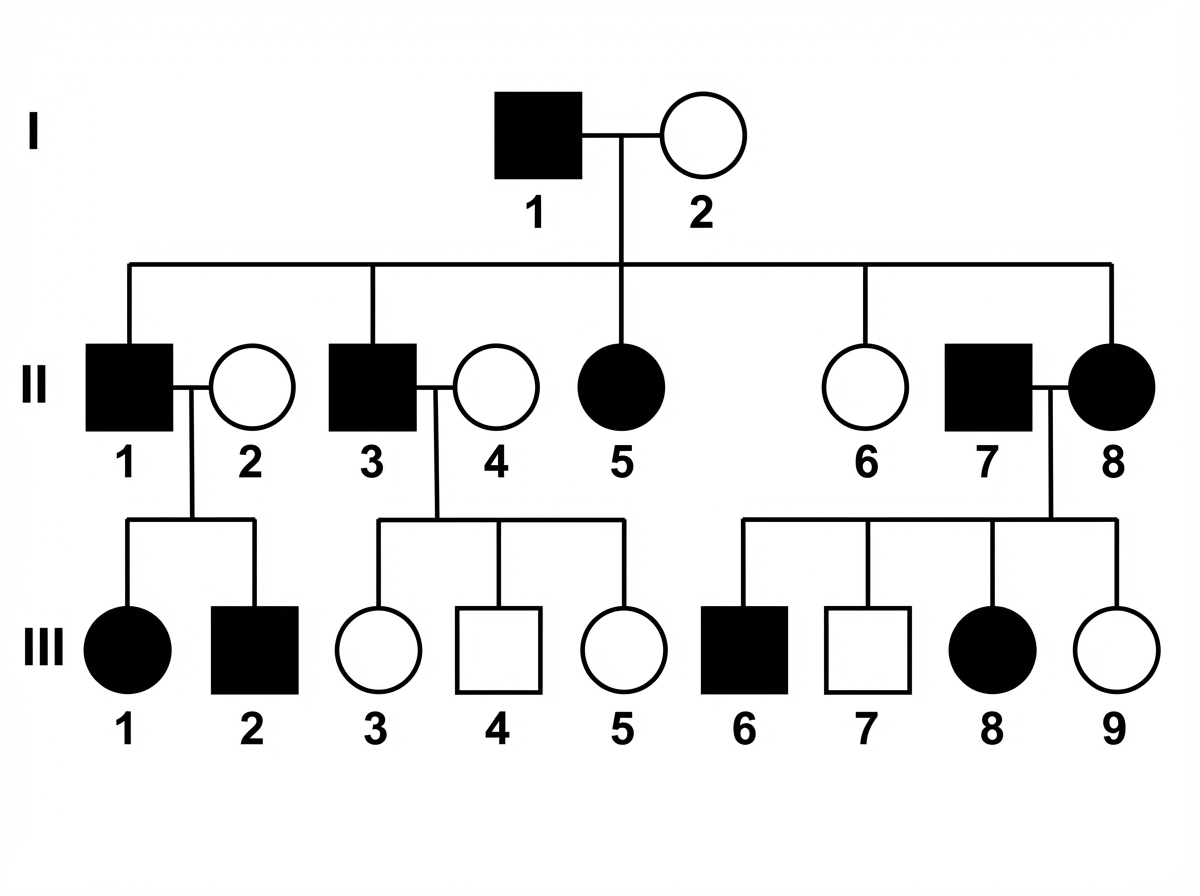
Which disease will show the mode of inheritance depicted in this pedigree?
A 12-year-old girl is brought to an oncologist, as she was recently diagnosed with a rare form of cancer. Cytogenetic studies reveal that the tumor is responsive to vinblastine, which is a cell-cycle specific anticancer agent. It acts on the M phase of the cell cycle and inhibits the growth of cells. Which of the following statements best describes the regulation of the cell cycle?
A 4-year-old male presents with a 1-year history of swaying while walking and recent episodes of tripping when ambulating. He has trouble trying to sit and get up from chairs, as well as walking up the stairs to his bedroom. On physical exam, the pediatrician notices nystagmus, absent deep tendon reflexes, significant loss of vibratory and proprioceptive sensation in his extremities, pes cavus, and slight kyphoscoliosis. A blood sample is sent for DNA sequencing and the results show a significant expansion of the trinucleotide GAA on chromosome 9. Which of the following diseases displays a similar mode of inheritance as the disease affecting this patient?
A 1-year-old child who was born outside of the United States is brought to a pediatrician for the first time because she is not gaining weight. Upon questioning, the pediatrician learns that the child has had frequent pulmonary infections since birth, and on exam the pediatrician appreciates several nasal polyps. Genetic testing is subsequently ordered to confirm the suspected diagnosis. Testing is most likely to show deletion of which of the following amino acids from the protein involved in this child's condition?
Practice by Chapter
DNA structure and organization
Practice Questions
Chromosomal structure
Practice Questions
DNA replication
Practice Questions
Cell cycle and mitosis
Practice Questions
Meiosis and genetic recombination
Practice Questions
Mutation types and consequences
Practice Questions
Mendelian inheritance patterns
Practice Questions
Non-Mendelian inheritance
Practice Questions
Genetic linkage and mapping
Practice Questions
Population genetics principles
Practice Questions
Genetic polymorphisms
Practice Questions
Gene therapy approaches
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
