Molecular Pathology — MCQs

On this page
Foci of granulomatous inflammation show all of the following except?
NKX3-1 immunohistochemistry is used for the diagnosis of which of the following?
Which of the following is considered a major pyrogenic cytokine?
A 60-year-old asymptomatic female shows the following change in the tunica media of blood vessels. What is the diagnosis?
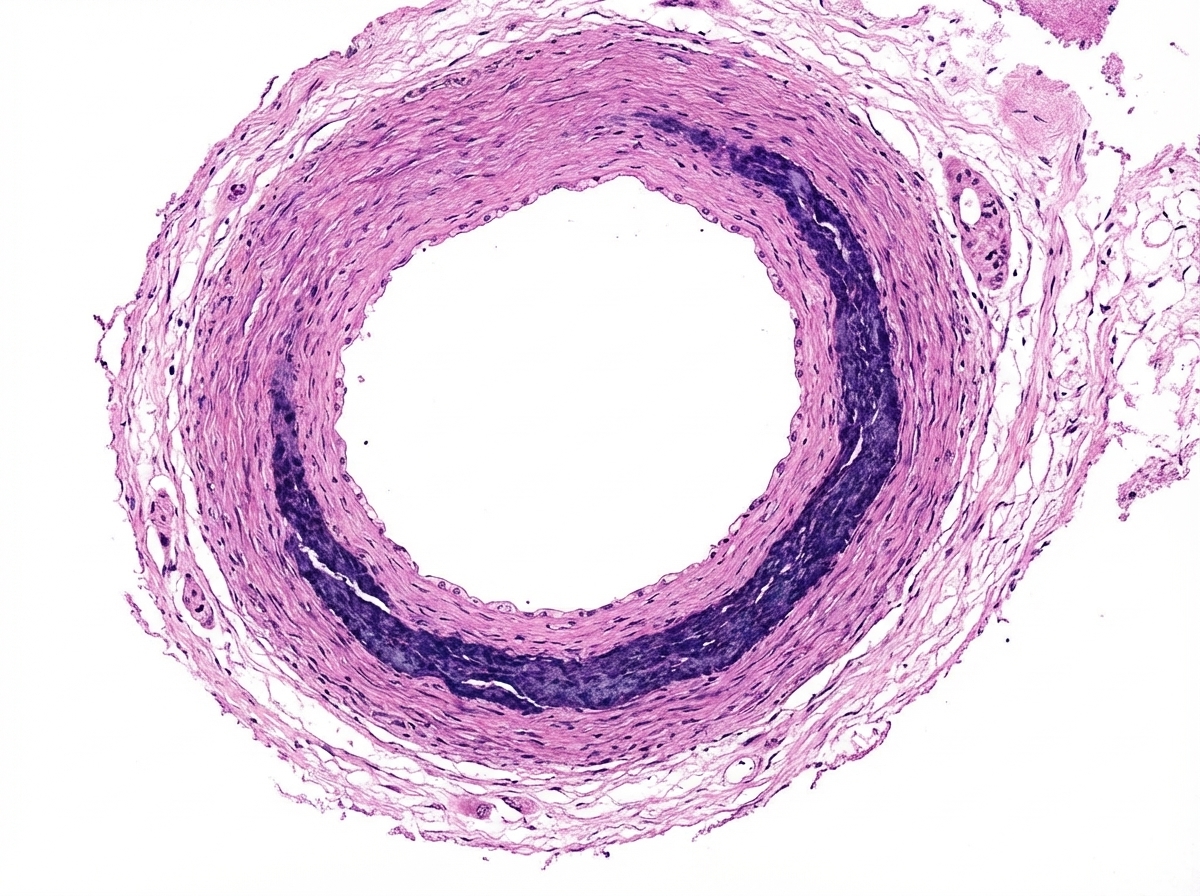
What is the primary molecular pathology associated with Marfan syndrome?
Practice by Chapter
Principles of Molecular Pathology
Practice Questions
DNA and RNA Analysis Techniques
Practice Questions
Cytogenetics
Practice Questions
Polymerase Chain Reaction Applications
Practice Questions
Next-Generation Sequencing
Practice Questions
Molecular Diagnosis of Infectious Diseases
Practice Questions
Molecular Oncology
Practice Questions
Pharmacogenomics
Practice Questions
Genetic Counseling and Risk Assessment
Practice Questions
Molecular Diagnostics Quality Control
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
