Protein Folding and Chaperones — MCQs

All are true about prions EXCEPT:
Which of the following reagents would be most useful in determining the N-terminal amino acid of a polypeptide?
Which factor stabilizes the alpha-helical structure of proteins?
All are true regarding G protein Except?
What type of bond is involved in the side chain linkage of proteoglycans?
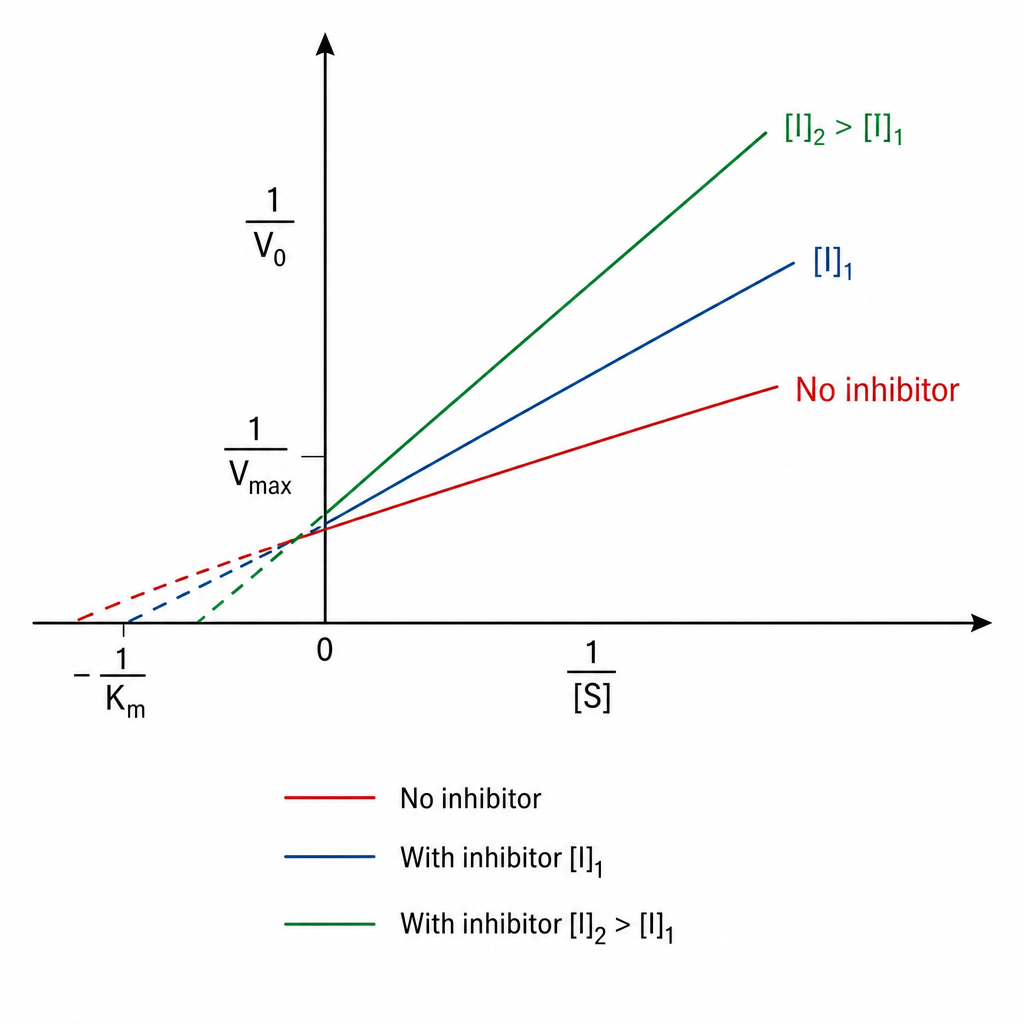
Identify the type of inhibition exhibited by A?
What is the primary role of calnexin and calreticulin in the endoplasmic reticulum?
Which of the following statements regarding prion diseases is NOT true?
In response to changes in Ca2+ concentration, which of the following Ca2+ binding proteins can modify the activity of many enzymes & proteins?
What do chaperones assist in?
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
