Transcription Factors and Gene Regulation — MCQs

miRNA binds to which part of the mRNA to inhibit translation?
Steroid hormone receptors have attachment site for all except:
Which of the following statements regarding Huntington’s chorea is true?
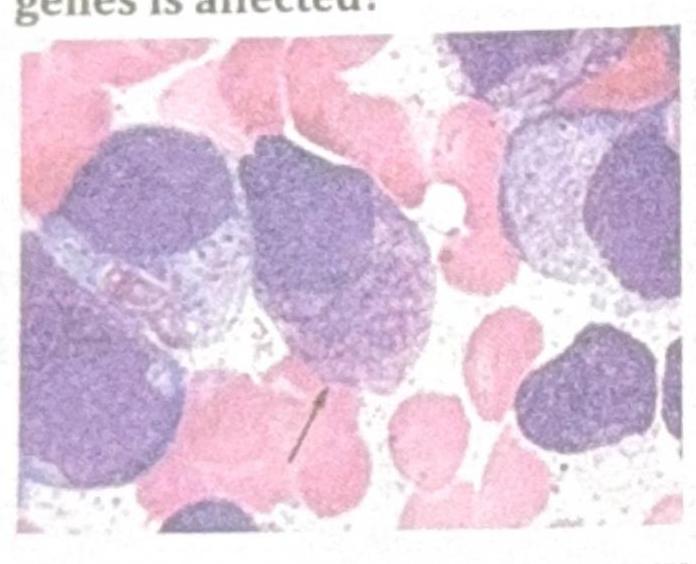
Identify the gene commonly involved in the condition shown in the image?
Which type of mutation can act as a suppressor to restore the wild-type phenotype in organisms carrying a mutant gene?
Which of the following is an X-linked dominant disorder?
Which one of the following is an autosomal dominant disorder?
Which one of the following statements about chromatin is not true?
Assertion: DNA methylation leads to gene silencing. Reason: Methylation prevents binding of transcription factors to promoter regions.
Phenotypic expression of a gene depending on the parent of origin is referred to as:
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
