Enzyme Kinetics and Michaelis-Menten Equation — MCQs

Which of the following represents the most significant regulatory control point among these TCA cycle reactions?
Which of the following statements is correct regarding the given graph?
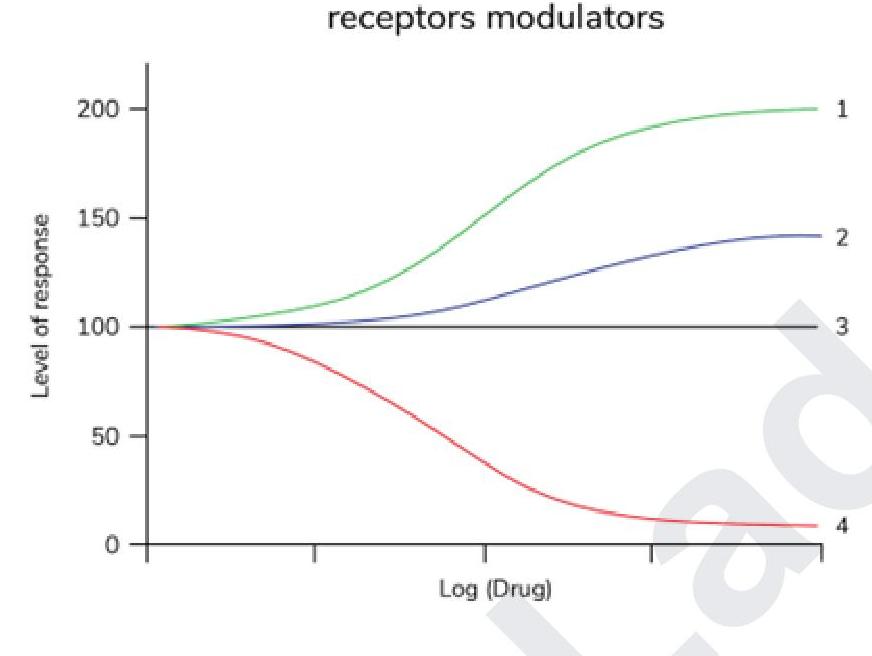
Km of an enzyme is
Identify the type of inhibition exhibited by A?
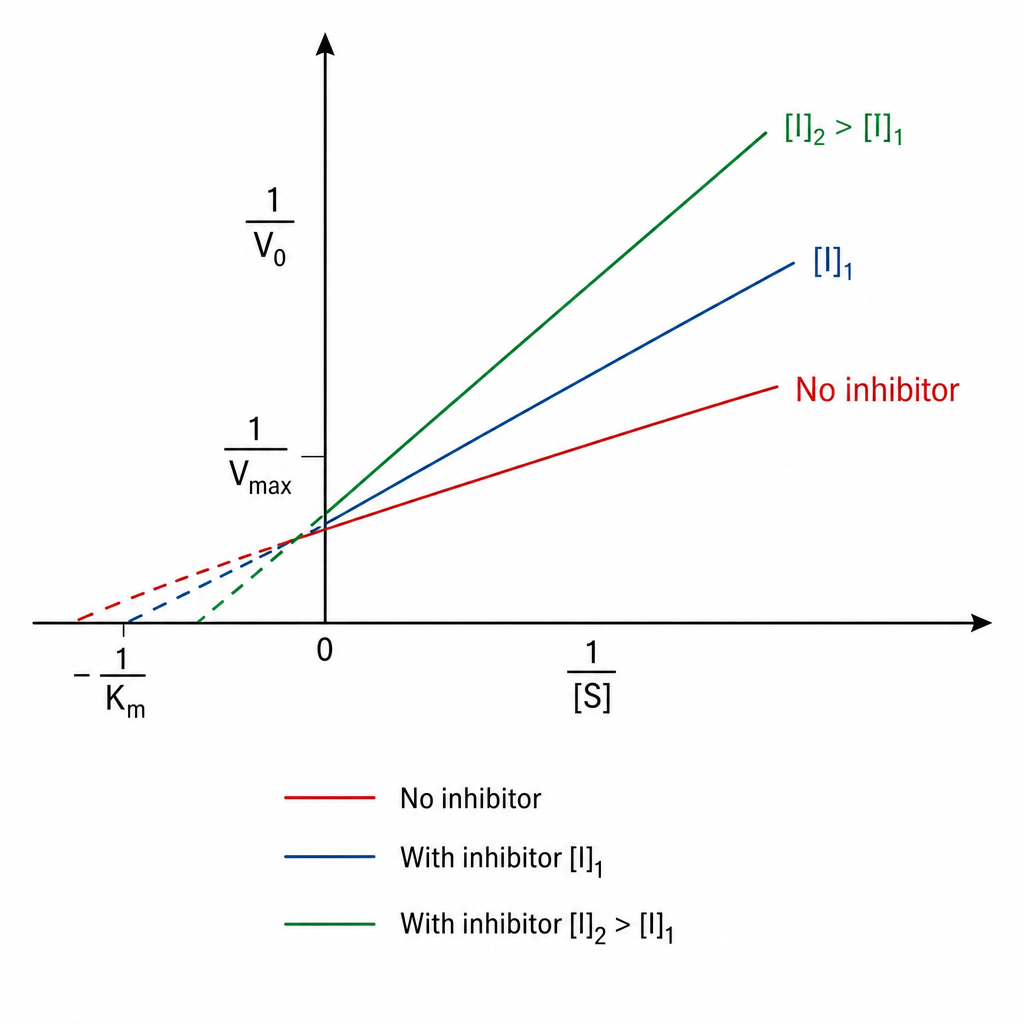
Which of the following statements best describes the mechanism of action of insulin on target cells?
Kcat/Km is a measure of which of the following?
Km increases, but Vmax remains same. This is which type of inhibition?
Which of the following statements about isozymes is true?
What is the specific activity of an enzyme?
Which kinetic parameter is primarily associated with enzyme specificity?
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
