Biochemical Techniques — MCQs

On this page
In PCR, DNA polymerase is used for which process?
What method is used to determine the tertiary structure of a protein?
Nephelometry is based on the principle of what?
Which amino acid migrates fastest on paper chromatography on methylcellulose medium?
Amplification of DNA uses the polymerase chain reaction (PCR) technique. Which cation is essential for PCR?
Match the following blotting techniques with the type of biomolecule they detect: Blotting Technique Detects A. Southern blot 1. RNA B. Northern blot 2. Lipids C. Western blot 3. DNA D. Eastern blot 4. Protein
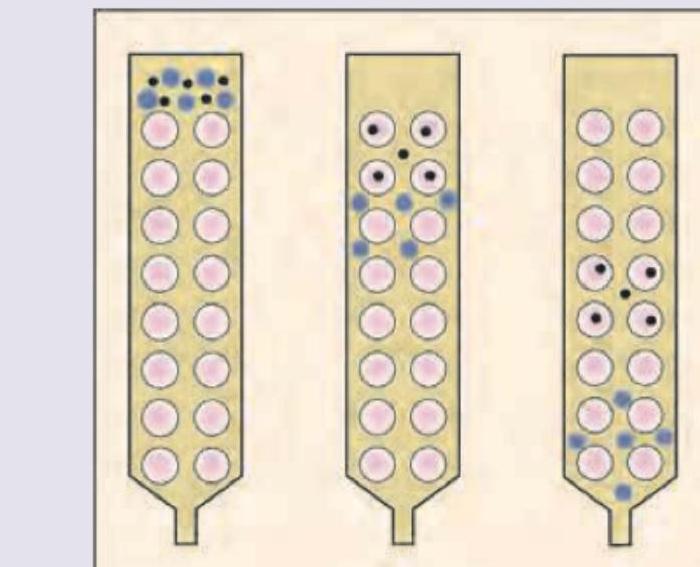
The following method of chromatography is called:
Which is the correct sequence of steps in isolating desirable protein using recombinant DNA technology? 1. Expression of protein and lysis of the bacterial cell 2. Incorporation of genes into bacteria 3. SDS PAGE 4. Protein elution 5. Column chromatography
Which of the following methods cannot be used to precipitate proteins?
Glycated hemoglobin (HbA1c) is best measured using?
Practice by Chapter
Spectrophotometry and Colorimetry
Practice Questions
Chromatography Techniques
Practice Questions
Electrophoresis and Applications
Practice Questions
Centrifugation and Ultracentrifugation
Practice Questions
Radioisotope Techniques
Practice Questions
Enzyme-Linked Immunosorbent Assay (ELISA)
Practice Questions
Polymerase Chain Reaction (PCR)
Practice Questions
Blotting Techniques: Southern, Northern, Western
Practice Questions
Mass Spectrometry in Biochemistry
Practice Questions
Recombinant DNA Technology
Practice Questions
DNA Sequencing
Practice Questions
Proteomics and Metabolomics
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
