Biochemical Techniques — MCQs

On this page
Gel filtration chromatography separates proteins based on which property?
Two proteins with the same charge can be separated by which of the following methods?
Which cation is commonly used in PCR?
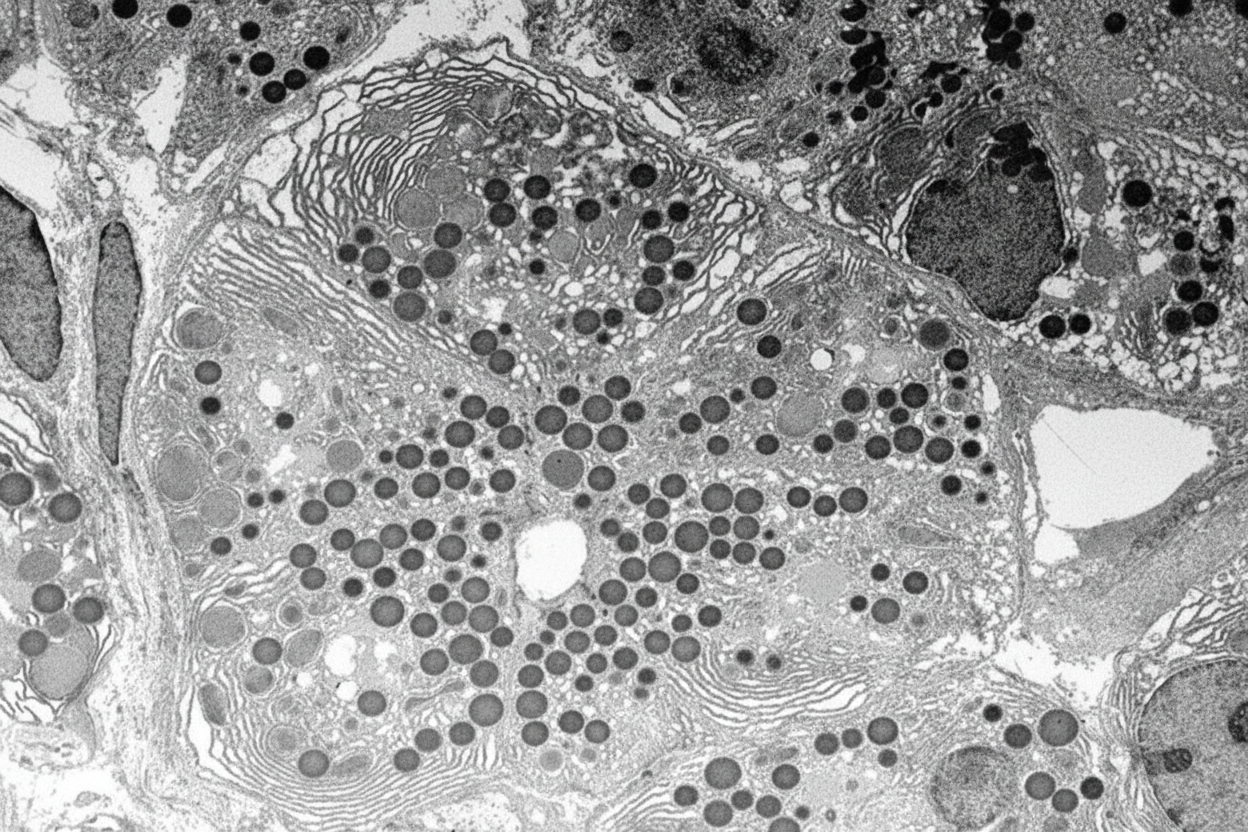
Which of the following enzymes is present inside the given organelle?
The Southern blot test is used for the detection of which of the following?
Which ion is essential in PCR?
Which blotting technique is used for RNA analysis?
All of the following assays are used for estimation of total serum protein, except?
Concentration of DNA is measured by:
Which of the following methods is used for the analysis of the C-terminal end of a polypeptide?
Practice by Chapter
Spectrophotometry and Colorimetry
Practice Questions
Chromatography Techniques
Practice Questions
Electrophoresis and Applications
Practice Questions
Centrifugation and Ultracentrifugation
Practice Questions
Radioisotope Techniques
Practice Questions
Enzyme-Linked Immunosorbent Assay (ELISA)
Practice Questions
Polymerase Chain Reaction (PCR)
Practice Questions
Blotting Techniques: Southern, Northern, Western
Practice Questions
Mass Spectrometry in Biochemistry
Practice Questions
Recombinant DNA Technology
Practice Questions
DNA Sequencing
Practice Questions
Proteomics and Metabolomics
Practice Questions
Want unlimited practice?
Get full access to all questions, explanations, and performance tracking.
Scan to download app
