AST Fundamentals - The Resistance Rules
-
Intrinsic Resistance: Innate, predictable non-susceptibility.
- Gram-negatives vs. Vancomycin (impermeable outer membrane).
- Anaerobes vs. Aminoglycosides (require O₂ for uptake).
- Mycoplasma vs. β-lactams (no cell wall).
-
Acquired Resistance: Gained via mutation or horizontal gene transfer (plasmids, transposons).
- mecA gene → MRSA
- ESBLs → Cephalosporin resistance
-
Inducible Resistance: Expressed only upon antibiotic exposure.
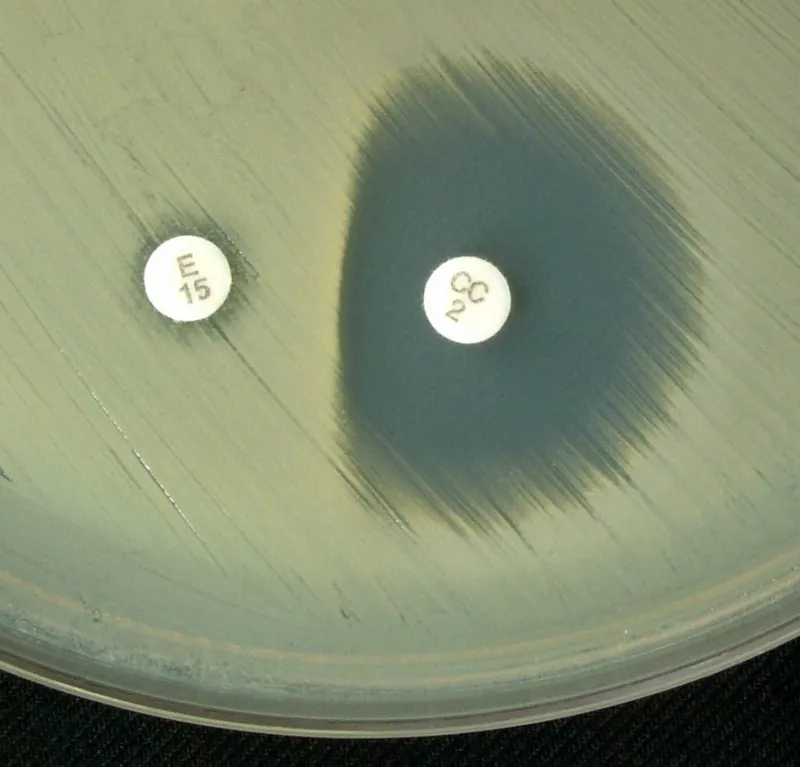
- D-Test: Detects inducible clindamycin resistance in S. aureus when exposed to erythromycin.
⭐ MRSA resistance is due to the mecA gene, which codes for Penicillin-Binding Protein 2a (PBP2a). PBP2a has a low affinity for β-lactam antibiotics, rendering them ineffective.
Phenotypic Methods - Reading the Zones
- Principle: Assess antimicrobial effectiveness by observing bacterial growth in the presence of the agent.
- Interpretation: Growth inhibition is measured to classify an organism as Susceptible (S), Intermediate (I), or Resistant (R) based on clinical breakpoints.

| Method | Principle | Measures | Use Case |
|---|---|---|---|
| Kirby-Bauer | Disk diffusion on agar | Zone of Inhibition (ZOI) diameter (mm) | Rapid, qualitative screening |
| Broth Dilution | Serial dilution in liquid media | Minimum Inhibitory Concentration (MIC) | Quantitative; gold standard for MIC |
| E-test | Gradient diffusion on agar | MIC (at ellipse intersection) | Quantitative; alternative to broth |
Special Screens - Unmasking Superbugs
- ESBL (Extended-Spectrum β-Lactamase) Screen:
- Suspect in E. coli or Klebsiella resistant to ceftriaxone/ceftazidime.
- Confirmation: Test with ceftazidime alone vs. ceftazidime + clavulanate. An increase in zone size (≥ 5 mm) with clavulanate confirms ESBL production.
-
D-Test (Inducible Clindamycin Resistance):
- For S. aureus isolates resistant to erythromycin but susceptible to clindamycin.
- Procedure: Place erythromycin and clindamycin disks adjacent on agar.
- Positive: Flattening of the clindamycin inhibition zone ("D" shape) indicates inducible erm gene resistance.
-
MRSA (Methicillin-Resistant S. aureus):
- Screen: Cefoxitin disk is a better predictor than oxacillin.
- Definitive: Detection of PBP2a (penicillin-binding protein 2a) or the mecA gene via PCR.
⭐ A positive D-test predicts in vivo clindamycin failure, even if the initial report shows susceptibility. Do not use clindamycin for treatment.
- CRE (Carbapenem-Resistant Enterobacteriaceae):
- Detected by tests like mCIM (modified Carbapenem Inactivation Method) or Carba NP.
High‑Yield Points - ⚡ Biggest Takeaways
- Minimum Inhibitory Concentration (MIC) is the lowest drug concentration that inhibits bacterial growth; it is the primary value for guiding dosage.
- Minimum Bactericidal Concentration (MBC) is the lowest concentration that kills 99.9% of bacteria, crucial for severe infections.
- The Kirby-Bauer test is a disk diffusion method that qualitatively assesses susceptibility based on the zone of inhibition size.
- The E-test is a gradient strip method that provides a quantitative MIC value.
- Clinical Breakpoints (S/I/R) are standardized MIC values used to predict clinical outcomes.
- Synergism and antagonism describe how drug combinations interact, impacting therapeutic choices.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

