5' Capping - The Starting Cap
- What: Addition of a 7-methylguanosine (m7G) cap to the 5' end of nascent pre-mRNA in the nucleus.
- How: An unusual 5'-to-5' triphosphate linkage is formed, catalyzed by guanylyltransferase.
- When: Occurs co-transcriptionally, shortly after transcription initiation by RNA polymerase II.
- Functions:
- Protects mRNA from degradation by 5' exonucleases.
- Essential for export from the nucleus to the cytoplasm.
- Acts as a binding site for the cap-binding complex (CBC).
⭐ The 5' cap is recognized by eukaryotic initiation factor eIF4E, which recruits the small ribosomal subunit (40S) to the mRNA, marking a critical step for initiating translation.

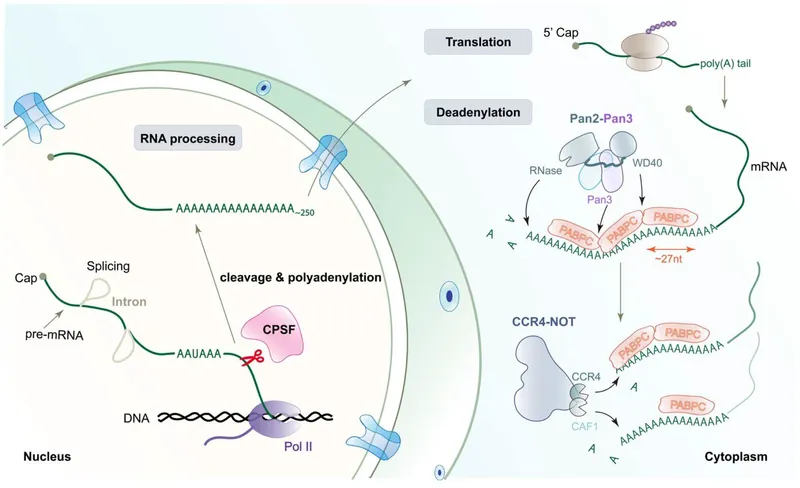
3' Polyadenylation - The Protective Tail
- Function: A long chain of adenine nucleotides (~250 A's) is added to the 3' end of pre-mRNA, creating the poly(A) tail.
- Mechanism:
- Triggered by the polyadenylation signal sequence (AAUAAA) near the 3' end.
- An endonuclease cleaves the pre-mRNA downstream of this signal.
- Polyadenylate polymerase (PAP) adds the adenine residues without a template.
- Purpose:
- ↑ mRNA stability by protecting it from 3' exonuclease degradation.
- Facilitates nuclear export.
- Required for efficient translation initiation.
⭐ The poly(A) tail interacts with Poly(A)-Binding Protein (PABP). PABP helps circularize the mRNA by binding to initiation factors on the 5' cap, significantly boosting translation efficiency.
Splicing - The Cutting Room Floor
-
Process of removing non-coding regions (introns) and joining coding regions (exons) from pre-mRNA. Occurs in the nucleus.
-
Carried out by the spliceosome, a large complex of small nuclear ribonucleoproteins (snRNPs).
-
Key Sites: Spliceosome recognizes consensus sequences.
- 5'-GU...AG-3': The intron begins with GU and ends with AG.
- 📌 Mnemonic: Got Under, After Going.
⭐ Autoantibodies against snRNPs (anti-Smith antibodies) are highly specific for Systemic Lupus Erythematosus (SLE).
- Alternative Splicing: A single gene can yield multiple mature mRNAs and proteins by selectively including or excluding exons. This greatly increases proteomic diversity.
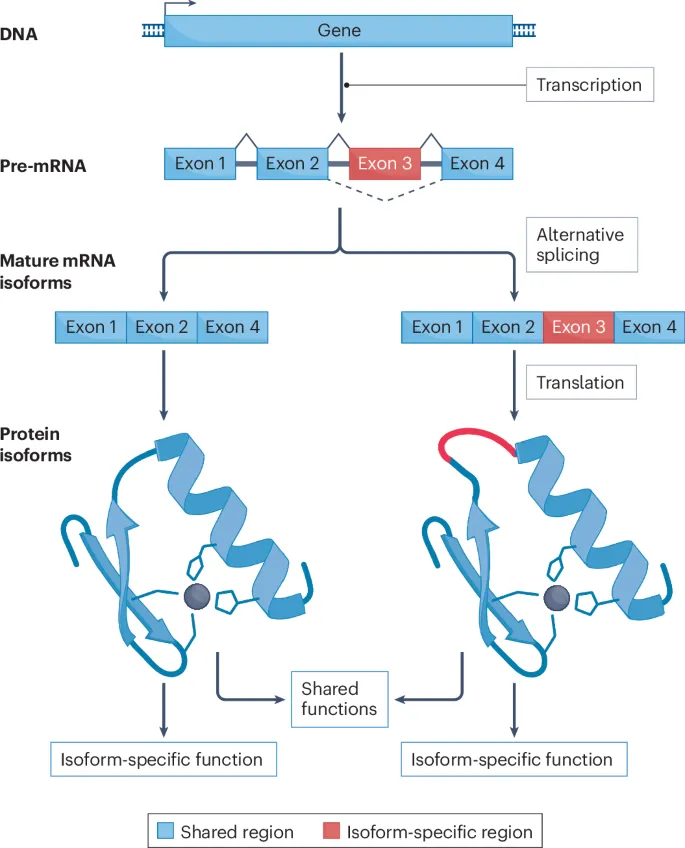
Splicing Variants - One Gene, Many Proteins
- Alternative Splicing: A regulated process where a single pre-mRNA transcript is processed in different ways, leading to distinct mature mRNA molecules.
- Mechanism:
- Spliceosomes selectively remove introns and join exons.
- By including or excluding certain exons, different combinations are made from the same gene.
- This vastly increases the proteomic diversity from a limited number of genes.
- Example: The tropomyosin gene produces multiple protein isoforms specific to different tissues (e.g., smooth vs. striated muscle) through alternative splicing.
⭐ The calcitonin gene produces calcitonin in thyroid C-cells but produces calcitonin gene-related peptide (CGRP) in neural tissue due to alternative splicing.
- Eukaryotic mRNA is processed in the nucleus before translation.
- 5' capping with 7-methylguanosine is crucial for translation initiation and mRNA protection.
- 3' polyadenylation (poly-A tail) stabilizes mRNA and facilitates its export from the nucleus.
- Splicing removes introns (non-coding regions) via the spliceosome; key sites are GU (5' end) and AG (3' end).
- Alternative splicing allows a single gene to code for multiple protein isoforms.
- Anti-Smith antibodies in SLE target snRNPs of the spliceosome.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

