Intrinsic Mechanisms - Bacterial Armor Up!
- Reduced Permeability: Bacteria armor up by making their membranes less porous.
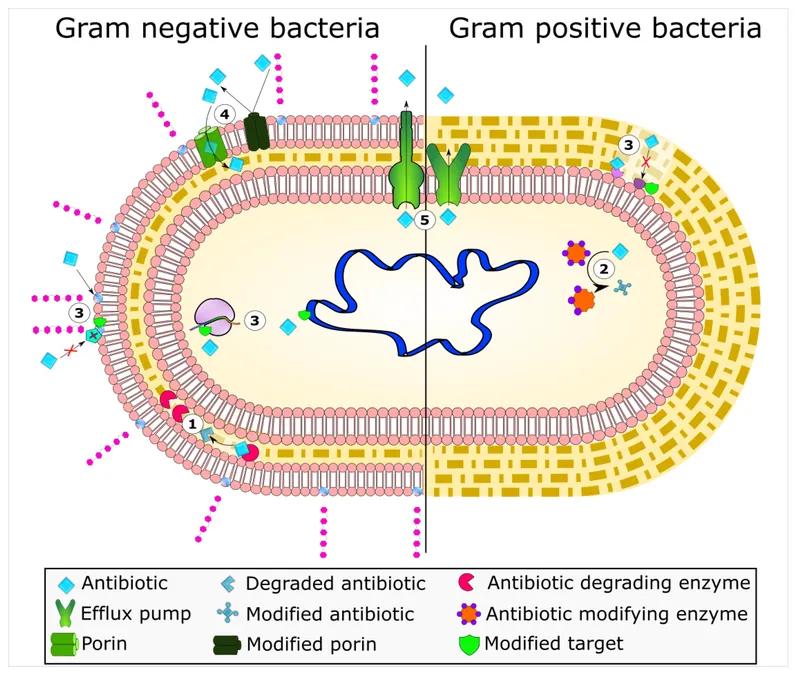
- Gram-negatives modify porin channels to block drug entry (e.g., Pseudomonas vs. Carbapenems).
- Thickened cell walls can slow drug access (e.g., VISA).
- Efflux Pumps: Actively pump antibiotics out of the cell before they can act.
- A major driver of multi-drug resistance (MDR) across many bacterial species.
- Enzymatic Inactivation: Bacteria produce enzymes that neutralize the antibiotic.
- Classic example: β-lactamases cleaving the β-lactam ring in penicillins.
⭐ Pseudomonas aeruginosa is notorious for its intrinsic resistance, combining low membrane permeability, multiple efflux pumps, and a chromosomally encoded AmpC β-lactamase.
Genetic Transmission - Spreading the Bad News
Resistance genes are shared between bacteria via horizontal gene transfer. This allows for rapid dissemination of resistance, even across different species. Key mechanisms include:
| Method | Mechanism | DNA Source |
|---|---|---|
| Transformation | Direct uptake of naked DNA from the environment | Lysed bacteria |
| Conjugation | Transfer via direct cell-to-cell contact (sex pilus) | Plasmid, transposon |
| Transduction | Bacteriophage (virus) acts as a vector | Bacterial chromosome |
⭐ High-Yield Fact: Conjugation is the most significant mechanism for spreading resistance in gram-negative bacteria, often transferring multi-drug resistance plasmids (R-plasmids).

The Usual Suspects - High-Yield Resistant Bugs

| Organism | Key Resistance Mechanism(s) | Genetic Basis | Clinical Pearls |
|---|---|---|---|
| MRSA | Altered Penicillin-Binding Protein (PBP2a) | mecA gene (transposon) | Resists all β-lactams. Vancomycin is a common treatment. |
| VRE | D-Ala-D-Ala → D-Ala-D-Lac in peptidoglycan | vanA or vanB operon (plasmid/transposon) | 📌 Vancomycin Resistance? Enterococci Laugh At Cell wall synthesis. |
| ESBL Producers | Extended-Spectrum β-Lactamases | Plasmids (e.g., SHV, TEM, CTX-M) | Hydrolyze most penicillins & cephalosporins. Treat with carbapenems. |
| CRE | Carbapenemases (e.g., KPC, NDM-1) | Plasmids | Often resistant to nearly all available antibiotics. |
| MDR-Pseudomonas | Efflux pumps, Porin loss, AmpC β-lactamase | Chromosomal mutations, plasmids | Intrinsic & acquired resistance. Common in nosocomial infections. |
| MDR-TB | Mutations in drug target genes | rpoB (Rifampin), katG (Isoniazid) | Requires multi-drug regimens for extended periods. |
High‑Yield Points - ⚡ Biggest Takeaways
- Antimicrobial resistance evolves via random mutation or horizontal gene transfer (conjugation, transformation, transduction).
- Selective pressure from antibiotic use is the primary driver for the proliferation of resistant bacteria.
- Common mechanisms include enzymatic inactivation (e.g., β-lactamases), target modification, and active efflux pumps.
- Plasmids and transposons are key mobile genetic elements that transfer resistance genes between bacteria, including across species.
- Inappropriate prescribing and agricultural use are major contributors.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

