Genetic Linkage - Genes Stick Together
- Genetic Linkage: Tendency for genes located close together on the same chromosome to be inherited together. This is an exception to Mendel's Law of Independent Assortment.
- Synteny: Genes located on the same chromosome, regardless of linkage.
| Feature | Independent Assortment | Genetic Linkage |
|---|---|---|
| Gene Location | Different chromosomes | Same chromosome, close |
| Gametes | Parental = Recombinant | Parental > Recombinant |
| Mechanism | Random alignment | Crossing over (recombination) |
⭐ Genes on different chromosomes or those very far apart on the same chromosome follow Mendel's law of independent assortment, producing a 50% recombination frequency.
- Recombination: The process of crossing over during meiosis that separates linked genes, creating recombinant gametes. The closer the genes, the lower the recombination frequency.
Recombination & LOD Score - Measuring Linkage
-
Recombination Frequency (θ): Measures the proportion of recombinant offspring, indicating the distance between linked genes.
- Formula: $θ = (Number of Recombinants) / (Total Progeny)$
- 1 Map Unit (m.u.) or 1 centiMorgan (cM) corresponds to a 1% recombination frequency. $1 cM ≈ 1% recombination frequency$.
-
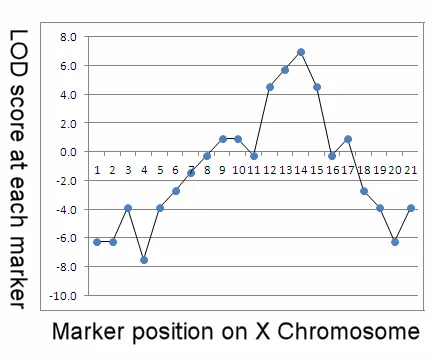
LOD Score (Logarithm of Odds): A statistical test to determine the probability that two genes are linked.
- Formula: $LOD(θ) = log10 [P(data | linkage at θ) / P(data | no linkage, θ=0.5)]$
- LOD score ≥ 3.0: Strong evidence FOR linkage.
- LOD score ≤ -2.0: Strong evidence AGAINST linkage.
⭐ A recombination frequency of 50% (θ = 0.5) means the genes are unlinked and assort independently.
- Linkage Phase: Describes the parental origin of alleles.
- Coupling: Dominant alleles on one chromosome, recessive on the other (e.g., AB/ab).
- Repulsion: A dominant and recessive allele on each chromosome (e.g., Ab/aB).
Linkage Disequilibrium - Population-Level Association
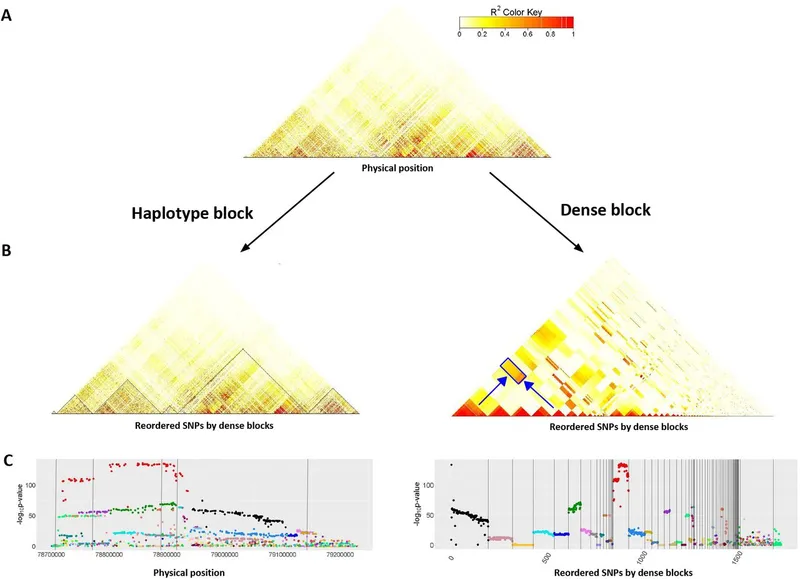
- Linkage Disequilibrium (LD): The non-random association of alleles at different loci on the same chromosome within a population. Alleles are seen together more or less frequently than expected by chance.
- Haplotype: A group of specific alleles on a single chromosome that are inherited together. LD leads to the persistence of specific haplotypes over generations.
- Allelic Association: LD is the basis for associating specific alleles (markers) with a particular trait or disease, even if the marker itself is not causative.
- Genome-Wide Association Studies (GWAS): Utilize LD to find disease-associated loci. By genotyping a few "tag" SNPs, entire haplotypes can be inferred, making large-scale studies feasible.
⭐ Linkage disequilibrium is the principle that allows GWAS to identify disease-associated loci without sequencing every individual's entire genome.
Clinical Applications - Finding Disease Genes
- Pedigree Analysis: Tracks disease inheritance within a family to establish a pattern.
- Genetic Markers: DNA variations (e.g., SNPs, RFLPs) used as landmarks to locate a gene's position on a chromosome through linkage.
- Positional Cloning: A method to identify a disease-causing gene by its location on a chromosome, rather than by its function.
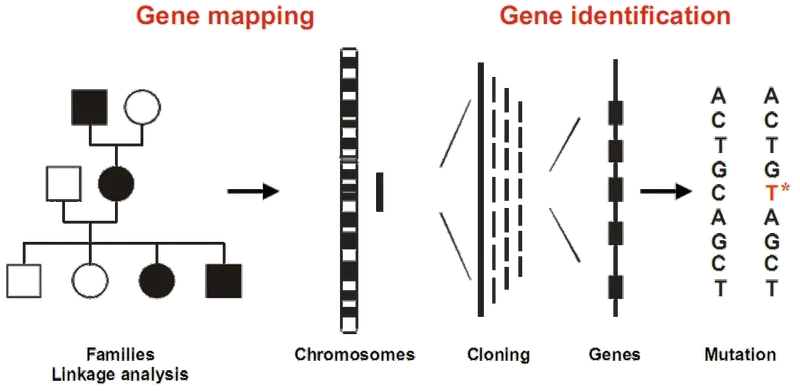
⭐ The gene for Huntington's disease was one of the first major disease genes mapped using linkage analysis with RFLP markers.
High‑Yield Points - ⚡ Biggest Takeaways
- Genetic linkage is the tendency for genes on the same chromosome to be inherited together.
- Recombination frequency (RF) is proportional to the physical distance between loci; ↓ RF means tighter linkage.
- One map unit (centimorgan) corresponds to a 1% recombination frequency.
- The maximum RF is 50%, at which point genes assort independently and are considered unlinked.
- Linkage disequilibrium is the non-random association of alleles at different loci.
- A LOD score > 3.0 provides strong statistical evidence for linkage.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

