Base Excision Repair - The DNA Janitors
- Repairs non-bulky, single-base defects like depurination, deamination (C→U, A→hypoxanthine), and oxidation (e.g., 8-oxoguanine).
- Occurs constantly throughout the cell cycle to clean up spontaneous DNA damage.
- 📌 Mnemonic: Glycosylase, Endonuclease, Lyase, Polymerase, Ligase (GEL PLease).
⭐ Uracil (from cytosine deamination) is the most common lesion corrected by BER. Since uracil belongs in RNA, it's immediately recognized as an error in DNA.
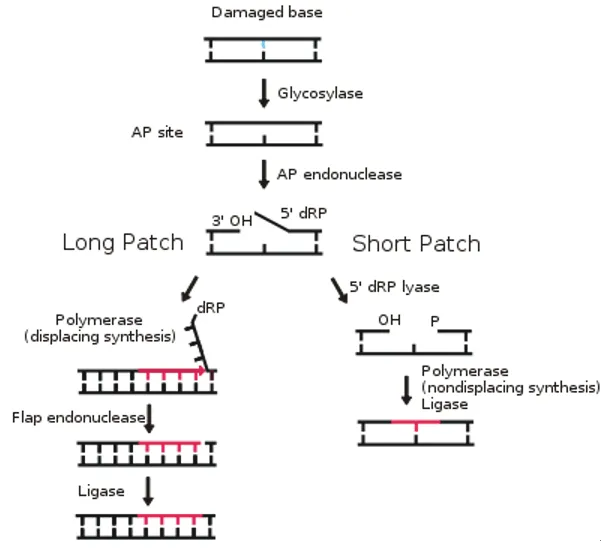
AP Site Cleavage - The Nick Creators
- An AP (apurinic/apyrimidinic) site is a sugar without a base in the DNA backbone, recognized by a specific endonuclease.
- AP Endonuclease 1 (APE1) is the key enzyme in humans that processes this lesion.
- APE1 incises the phosphodiester backbone on the 5' side of the AP site.
- This creates a single-strand break (a "nick") with two critical ends:
- A 3'-hydroxyl (3'-OH) group
- A 5'-deoxyribose phosphate (5'-dRP) residue
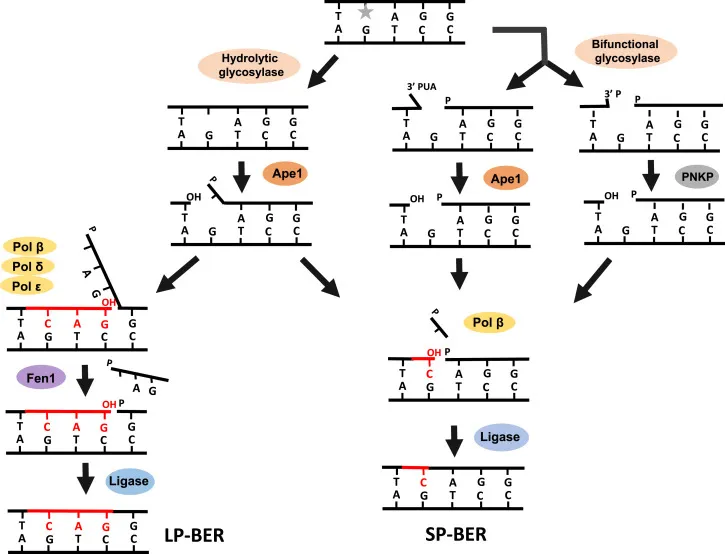
⭐ APE1 creates the necessary 3'-OH primer terminus, which is the essential substrate for DNA Polymerase β to initiate repair synthesis.
Synthesis & Ligation - The Fix-It Crew
- DNA Polymerase β (Pol β): Recruited to the AP site. It possesses both lyase activity to remove the baseless sugar and polymerase activity to insert a single, correct nucleotide into the gap.
- DNA Ligase: The final step. This enzyme seals the nick in the sugar-phosphate backbone by creating a phosphodiester bond, restoring the integrity of the DNA strand.
⭐ High-Yield: Eukaryotic DNA ligases use ATP as an energy source to catalyze the formation of the phosphodiester bond, a frequent exam topic comparing them to prokaryotic ligases which often use NAD+.
Clinical Correlations - When Repair Fails
- MUTYH-Associated Polyposis (MAP)
- Gene Defect: Biallelic mutation in the MUTYH gene, a key DNA glycosylase in the BER pathway.
- Inheritance: Autosomal Recessive.
- Pathophysiology: Failure to repair oxidative DNA damage, particularly 8-oxoguanine. This leads to an accumulation of G:C → T:A transversion mutations.
- Clinical Picture: Presents with multiple colorectal adenomatous polyps (typically 10-100s) and a significantly increased lifetime risk of colorectal cancer.
⭐ Unlike Familial Adenomatous Polyposis (FAP), which is autosomal dominant, MAP is autosomal recessive. This distinction is crucial for genetic counseling and screening family members.
High‑Yield Points - ⚡ Biggest Takeaways
- Base Excision Repair (BER) corrects non-bulky, single-base defects like spontaneous deamination (cytosine → uracil), alkylation, and oxidation.
- The key enzyme, base-specific Glycosylase, recognizes and removes the damaged base, creating an apurinic/apyrimidinic (AP) site.
- AP Endonuclease cleaves the 5' end of the AP site, and a Lyase removes the single, empty sugar-phosphate.
- DNA Polymerase β fills the gap, and DNA Ligase seals the nick.
- Think GEL PLease: Glycosylase, Endonuclease, Lyase, Polymerase, Ligase.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

