VRE Basics - The Stubborn Germs
- Enterococci: Gram-positive cocci, part of normal gut flora. Intrinsically resistant to many antibiotics (e.g., cephalosporins).
- Enterococcus faecalis: More common, generally more susceptible.
- Enterococcus faecium: Less common, but more likely to be resistant.
- Vancomycin-Resistant Enterococci (VRE): Strains that have acquired resistance to vancomycin, a last-resort antibiotic.
- Clinical Impact: Major cause of nosocomial infections, particularly UTIs, bacteremia, and endocarditis in immunocompromised patients.
⭐ E. faecium is inherently more resistant and the predominant species in VRE infections compared to E. faecalis.
Vancomycin's MO - Wall-Wrecker at Work
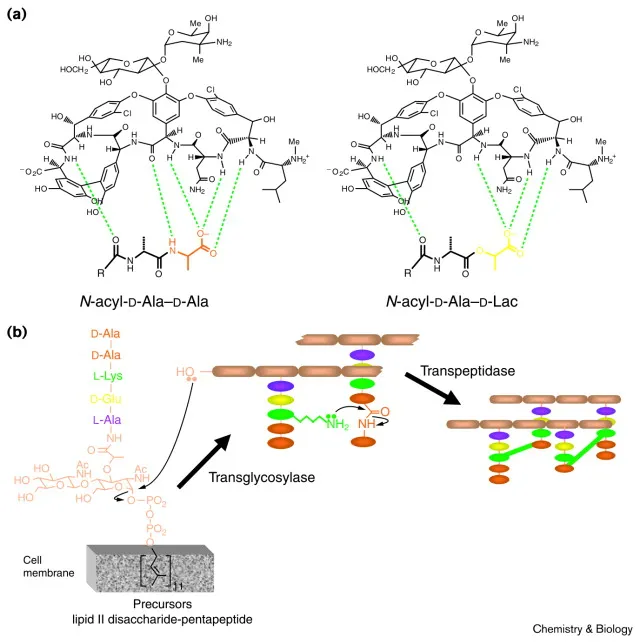
Vancomycin physically obstructs bacterial cell wall synthesis. It's a large glycopeptide that directly binds to the $D-Alanine-D-Alanine$ termini of nascent peptidoglycan chains.
- Mechanism: Forms a cap on the $D-Ala-D-Ala$ substrate.
- Effect: This steric hindrance blocks two crucial enzymes:
- Transglycosylase: Halts the elongation of the glycan chain.
- Transpeptidase (a PBP): Prevents cross-linking of the peptide side chains.
- Outcome: Inhibition of peptidoglycan synthesis → compromised cell wall integrity → cell lysis.
⭐ Because of its large size, vancomycin cannot penetrate the porin channels of gram-negative bacteria, making them intrinsically resistant.
The van Genes - Resistance Masterminds
Vancomycin resistance in enterococci (VRE) is primarily due to the alteration of the drug's target site in the bacterial cell wall. This is orchestrated by van gene clusters.
- Mechanism: The van operon codes for enzymes that synthesize a different peptidoglycan precursor.
- The normal target, $D-Ala-D-Ala$, is replaced with $D-Ala-D-Lac$.
- This substitution prevents vancomycin from binding effectively, as it disrupts crucial hydrogen bonds.
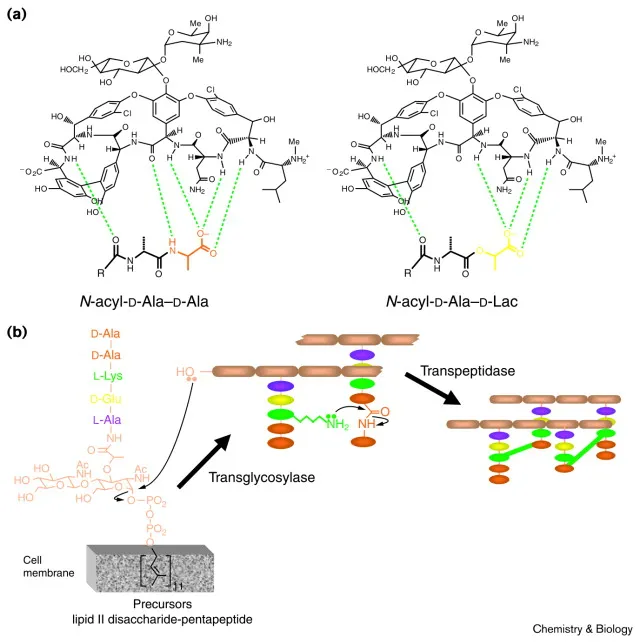
⭐ The vanA and vanB gene clusters are the most clinically significant and are typically located on mobile genetic elements (plasmids, transposons), facilitating their spread.
| Phenotype | Expression | Vancomycin Resistance | Teicoplanin Resistance |
|---|---|---|---|
| VanA | Inducible | High (MIC ≥64 µg/mL) | ↑ High |
| VanB | Inducible | ↑ Variable (High-Level) | Susceptible |
| VanC | Constitutive | ↓ Low-Level | Susceptible |
- Vancomycin-Resistant Enterococci (VRE) resistance is primarily due to the alteration of the peptidoglycan synthesis pathway.
- The terminal dipeptide target D-Ala-D-Ala is modified to D-Ala-D-Lac (conferred by VanA and VanB genes) or D-Ala-D-Ser (VanC).
- This change dramatically reduces the binding affinity of vancomycin, rendering it ineffective.
- VanA provides high-level resistance to both vancomycin and teicoplanin.
- Resistance genes are often located on mobile plasmids and transposons, facilitating horizontal gene transfer.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

