Intro & Staphylococcal Chromosome - Superbug Blueprint
- MRSA (Methicillin-resistant Staphylococcus aureus): A quintessential "superbug" due to extensive antibiotic resistance, posing significant clinical challenges.
- Genetic Basis: Resistance is acquired via a mobile genetic element, not through simple mutation.
- Staphylococcal Cassette Chromosome mec (SCCmec): The core of the resistance blueprint.
- A large, mobile DNA element integrated into the S. aureus chromosome.
- Carries the mecA gene, the primary determinant of methicillin resistance.
- Contains ccr genes that mediate its own excision and integration.
 genetic map showing mecA and ccr genes)
genetic map showing mecA and ccr genes)
⭐ The size and composition of SCCmec cassettes vary, leading to different types (I-XIII). This variation has epidemiological significance for tracking MRSA clones.
MecA Gene & PBP2a - The Altered Target
- Core Mechanism: Resistance is mediated by the mecA gene, which encodes a novel Penicillin-Binding Protein, PBP2a.
- Altered Target: PBP2a has a very low affinity for β-lactam antibiotics.
- While standard PBPs are inhibited by drugs like methicillin and nafcillin, PBP2a remains functional.
- It takes over the crucial transpeptidation step in cell wall synthesis.
- Result: Continued peptidoglycan cross-linking allows the bacterium to survive, conferring resistance to nearly all β-lactams.
⭐ The mecA gene is located on a mobile genetic element, the Staphylococcal Cassette Chromosome mec (SCCmec), which allows for horizontal gene transfer between staphylococci.

Other Resistance Factors - Accessory Armor
-
Biofilm Formation:
- Mediated by polysaccharide intercellular adhesin (PIA), encoded by the ica operon.
- Creates a physical barrier against antibiotics and host immune cells.
- Crucial for infections on indwelling medical devices (e.g., catheters, prosthetic joints).
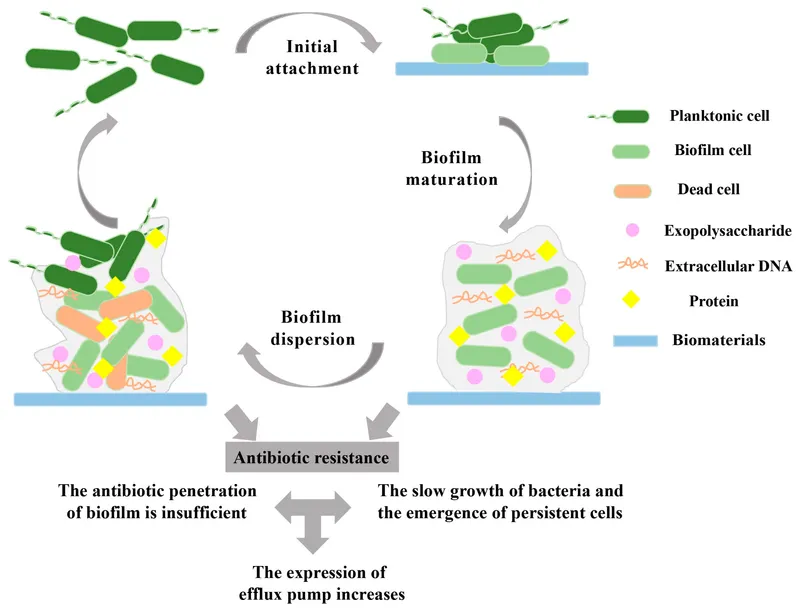
-
Efflux Pumps:
- Actively transport antibiotics (e.g., fluoroquinolones, tetracyclines) out of the bacterial cell.
- Key pumps: NorA, NorB, NorC.
- Contribute to low-level resistance and tolerance.
-
Toxin & Enzyme Production:
- Panton-Valentine Leukocidin (PVL): A cytotoxin destroying leukocytes, causing tissue necrosis. Strongly linked to community-associated MRSA (CA-MRSA).
- Accessory Gene Regulator (agr): Quorum-sensing system controlling virulence factor expression.
⭐ The agr quorum-sensing system is a key regulator of virulence in S. aureus. Its dysfunction is linked to persistent bacteremia and treatment failure, making it a potential therapeutic target.
High‑Yield Points - ⚡ Biggest Takeaways
- MRSA resistance is primarily mediated by the mecA gene, which encodes for a modified Penicillin-Binding Protein (PBP2a).
- PBP2a has a low affinity for β-lactam antibiotics, preventing them from inhibiting cell wall synthesis.
- The mecA gene is carried on a mobile genetic element, the Staphylococcal Cassette Chromosome mec (SCCmec).
- This allows for horizontal gene transfer, facilitating the rapid spread of resistance.
- Consequently, vancomycin is often the treatment of choice.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

