Basics & Principle - Glow & Go PCR
- Real-Time PCR (qPCR): Monitors PCR product amplification during each cycle for quantitative analysis, unlike conventional PCR (end-point).
- Principle: "Glow & Go" - fluorescence intensity, proportional to DNA amount, measured in real-time.
- Fluorescent reporters:
- SYBR Green: Binds non-specifically to dsDNA, fluoresces. Cost-effective.
- TaqMan Probes: Sequence-specific. Probe hydrolysis during extension separates fluorophore from quencher, emitting light. ↑Specificity.
- Fluorescent reporters:
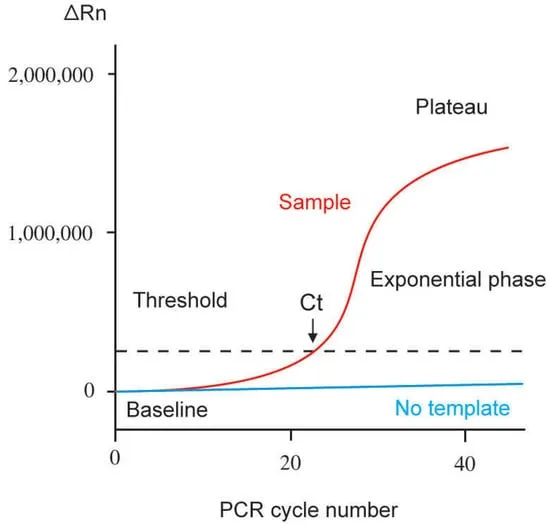
- Ct (Cycle threshold): Cycle number when fluorescence significantly exceeds background.
- Inversely proportional to initial template ($↓Ct = ↑initial DNA$).
⭐ qPCR allows precise quantification of starting nucleic acid, a key advantage over traditional PCR.
 oka
oka
Detection Chemistries - Tag, You're It!
- Non-Specific Dyes (e.g., SYBR Green I):
- Binds any dsDNA; fluorescence ↑ upon binding.
- Pros: Simple, cost-effective.
- Cons: Non-specific (binds primer-dimers). Requires melt curve analysis for specificity check.
- Specific Probes: ↑ Specificity; enables multiplexing.
- Hydrolysis Probes (e.g., TaqMan®):
- Probe: Reporter (R) + Quencher (Q).
- Taq's 5'→3' exonuclease activity cleaves probe during extension → R separates from Q → fluorescence ↑.
⭐ TaqMan® is the most common hydrolysis probe, offering high specificity and reliable quantification.
- Hybridization Probes (FRET Probes):
- Two probes: Donor & Acceptor. Fluorescence upon adjacent hybridization to target (FRET).
- Molecular Beacons:
- Stem-loop structure (R & Q close, quenched).
- Binding to target → opens hairpin → R & Q separate → fluorescence ↑. High specificity.
- Hydrolysis Probes (e.g., TaqMan®):
Data Interpretation - PCR's Crystal Ball
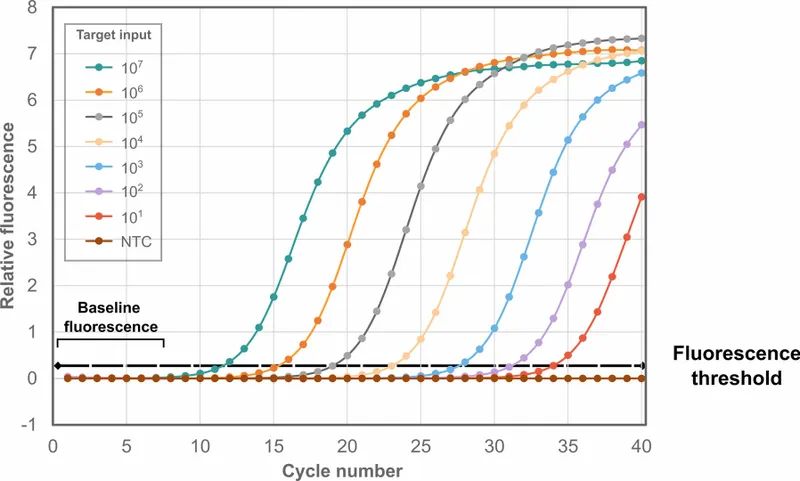
- Cycle Threshold ($C_t$ or $C_q$ Value):
- Cycle number at which fluorescence signal significantly exceeds background, crossing a set threshold.
- Inversely proportional to initial target nucleic acid amount: ↓$C_t$ implies ↑ initial template.
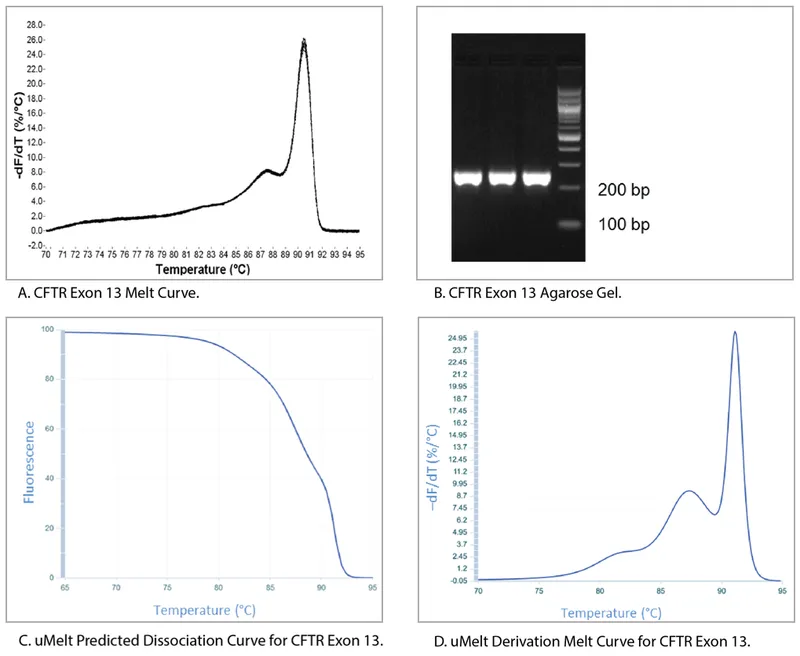
- Melting Curve Analysis:
- Performed post-PCR to assess product specificity and purity.
- Based on the unique melting temperature ($T_m$) of dsDNA, influenced by GC content and length.
- Helps distinguish specific products from non-specific amplicons or primer-dimers.
- Quantification Strategies:
- Absolute: Determines exact copy number using a standard curve generated from samples of known concentrations plotted against their $C_t$ values.
- Relative: Measures fold change in gene expression relative to a calibrator sample, normalized to a reference (housekeeping) gene, often using the $2^{-\Delta\Delta C_t}$ method.
⭐ A lower $C_t$ value signifies a higher initial concentration of the target nucleic acid.
Clinical Utility - Bug Detector Pro
- Rapid Pathogen ID:
- Bacteria (MTB/GeneXpert), Viruses (HIV, HCV, HBV, CMV), Fungi, Parasites.
- High sens/spec.
- Culture-negative detection (fastidious, prior Abx).
- Quantification (Load Monitoring):
- Viral load (HIV, HBV, HCV): monitors Rx efficacy, guides therapy.
- Bacterial load: assesses severity.
- Antimicrobial Resistance (AMR) Detection:
- Detects resistance genes (e.g., mecA, vanA/B, blaKPC).
- Guides targeted Abx therapy.
- Advantages:
- Rapid TAT (hours vs days).
- High throughput.
- Detects non-culturable.
- Closed system ↓ contamination.
- Limitations:
- DNA/RNA ≠ viable organism.
- Costly, expertise needed.
- False +/- risk.
- Infection vs. colonization challenge.
⭐ Real-Time PCR can detect as few as 1-10 copies of target DNA/RNA per reaction, making it exceptionally sensitive for early pathogen detection.
High‑Yield Points - ⚡ Biggest Takeaways
- Real-Time PCR (qPCR) enables quantification of DNA/RNA as it amplifies.
- Employs fluorescent reporters (e.g., SYBR Green, TaqMan probes) for real-time monitoring.
- Ct value is inversely proportional to initial target nucleic acid quantity.
- Melting curve analysis (SYBR Green) verifies product specificity, distinguishing from primer-dimers.
- Crucial for viral load quantification (HIV, HBV, HCV) and gene expression analysis.
- TaqMan probes provide specificity through 5'-3' exonuclease activity of Taq polymerase.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

