Blotting Techniques - Spot the Difference!
📌 SNoW DRoP: Southern = DNA; Northern = RNA; Western = Protein.
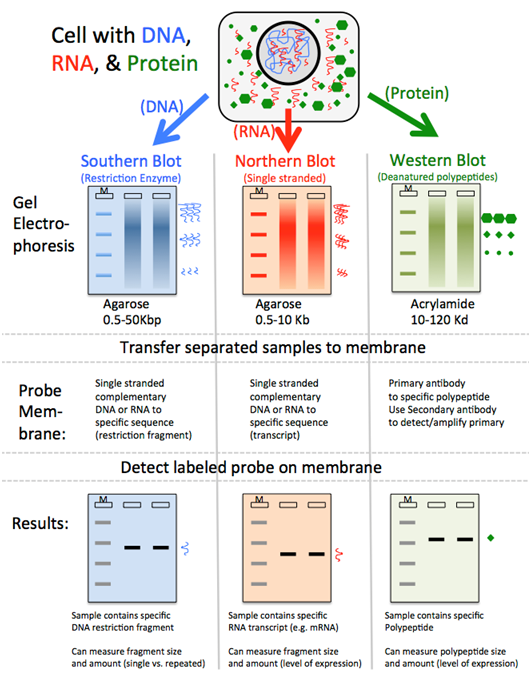
Blotting techniques identify specific DNA, RNA, or proteins post-electrophoresis. Molecules are transferred to a membrane, then detected with specific probes.
| Technique | Target Molecule | Probe Used | Key Clinical/Research Application(s) |
|---|---|---|---|
| Southern Blot | DNA | Labeled DNA/RNA | Genetic fingerprinting, RFLP, mutation detection |
| Northern Blot | RNA (typically mRNA) | Labeled DNA/RNA | Gene expression studies (mRNA abundance) |
| Western Blot | Protein | Labeled Antibody | Protein identification, quantification, disease diagnosis (e.g., HIV) |
| Southwestern | DNA-binding Proteins | Labeled dsDNA | Identifying transcription factors & interactions |
| Eastern Blot | Post-Translational Mods. | Lectins, specific Abs | Analyzing protein PTMs (glycosylation, etc.) |
| %%{init: {'flowchart': {'htmlLabels': true}}}%% | |||
| flowchart TD |
Start["<b>🧬 Sample Prep</b><br><span style='display:block; text-align:left; color:#555'>• Isolate DNA/RNA</span><span style='display:block; text-align:left; color:#555'>• Extract protein</span>"]
Electrophoresis["<b>🧪 Separation</b><br><span style='display:block; text-align:left; color:#555'>• Gel electrophoresis</span><span style='display:block; text-align:left; color:#555'>• Sort by size</span>"]
Transfer["<b>📄 Transfer</b><br><span style='display:block; text-align:left; color:#555'>• Move to membrane</span><span style='display:block; text-align:left; color:#555'>• NC or PVDF</span>"]
Blocking["<b>🛡️ Blocking</b><br><span style='display:block; text-align:left; color:#555'>• Prevent binding</span><span style='display:block; text-align:left; color:#555'>• BSA or milk</span>"]
Probing["<b>🔍 Probing</b><br><span style='display:block; text-align:left; color:#555'>• Primary antibody</span><span style='display:block; text-align:left; color:#555'>• Labelled probes</span>"]
Detection["<b>✨ Detection</b><br><span style='display:block; text-align:left; color:#555'>• Visualization</span><span style='display:block; text-align:left; color:#555'>• Band analysis</span>"]
Start --> Electrophoresis
Electrophoresis --> Transfer
Transfer --> Blocking
Blocking --> Probing
Probing --> Detection
style Start fill:#FFF7ED, stroke:#FFEED5, stroke-width:1.5px, rx:12, ry:12, color:#C2410C
style Electrophoresis fill:#FFF7ED, stroke:#FFEED5, stroke-width:1.5px, rx:12, ry:12, color:#C2410C
style Transfer fill:#FFF7ED, stroke:#FFEED5, stroke-width:1.5px, rx:12, ry:12, color:#C2410C
style Blocking fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534
style Probing fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534
style Detection fill:#F6F5F5, stroke:#E7E6E6, stroke-width:1.5px, rx:12, ry:12, color:#525252
> ⭐ Western blot is a highly specific confirmatory test for HIV, detecting antibodies against key viral proteins (e.g., p24, gp41, gp120).
## PCR & Amplification - Multiply & Conquer!
Polymerase Chain Reaction (PCR): Rapid in-vitro amplification of specific DNA. (9 words)
* **Core Principle**: Exponential DNA copy increase. (5 words)
* **Key Components**: (2 words)
- DNA Template: Target DNA. (3 words)
- Primers (Fwd/Rev): Short ssDNA, define region. (6 words)
- Taq Polymerase: Heat-stable DNA synthesizing enzyme. (6 words)
- dNTPs: Building blocks (A,T,C,G). (4 words)
- Buffer + Mg²⁺: Optimal conditions. (4 words)
* **PCR Cycle (~25-35 repeats)**: (4 words)
1. **Denaturation**: ~**95°C**; dsDNA → ssDNA. (6 words)
2. **Annealing**: ~**50-65°C**; primers bind template. (6 words)
3. **Extension**: ~**72°C**; Taq extends primers. (6 words)
📌 Mnemonic: DAT (Denature, Anneal, Taq extend). (7 words)
```mermaid
%%{init: {'flowchart': {'htmlLabels': true}}}%%
flowchart TD
Start["<b>🧪 Setup</b><br><span style='display:block; text-align:left; color:#555'>• DNA Sample</span><span style='display:block; text-align:left; color:#555'>• PCR Reagents</span>"]
Cycle{"<b>🔄 PCR Cycle</b><br><span style='display:block; text-align:left; color:#555'>• Thermal cycles</span><span style='display:block; text-align:left; color:#555'>• 3-step process</span>"}
Denature["<b>🔥 Denaturing</b><br><span style='display:block; text-align:left; color:#555'>• High heat applied</span><span style='display:block; text-align:left; color:#555'>• ssDNA Strands</span>"]
Anneal["<b>❄️ Annealing</b><br><span style='display:block; text-align:left; color:#555'>• Primers bind</span><span style='display:block; text-align:left; color:#555'>• Cooler temp</span>"]
Extend["<b>🧬 Extension</b><br><span style='display:block; text-align:left; color:#555'>• New DNA built</span><span style='display:block; text-align:left; color:#555'>• Taq polymerase</span>"]
Final["<b>📈 Amplified DNA</b><br><span style='display:block; text-align:left; color:#555'>• Exponential gain</span><span style='display:block; text-align:left; color:#555'>• Final product</span>"]
Start --> Cycle
Cycle -->|Denaturation| Denature
Denature -->|Annealing| Anneal
Anneal -->|Elongation| Extend
Extend -.->|Repeat| Cycle
Cycle ==>|Final| Final
style Start fill:#FFF7ED, stroke:#FFEED5, stroke-width:1.5px, rx:12, ry:12, color:#C2410C
style Cycle fill:#FEF8EC, stroke:#FBECCA, stroke-width:1.5px, rx:12, ry:12, color:#854D0E
style Denature fill:#FDF4F3, stroke:#FCE6E4, stroke-width:1.5px, rx:12, ry:12, color:#B91C1C
style Anneal fill:#EEFAFF, stroke:#DAF3FF, stroke-width:1.5px, rx:12, ry:12, color:#0369A1
style Extend fill:#F1FCF5, stroke:#BEF4D8, stroke-width:1.5px, rx:12, ry:12, color:#166534
style Final fill:#F6F5F5, stroke:#E7E6E6, stroke-width:1.5px, rx:12, ry:12, color:#525252

⭐ Taq polymerase from Thermus aquaticus is crucial for PCR; its thermostability allows it to survive repeated high-temperature denaturation steps. (21 words)
- Yield: $2^n$ DNA copies (n = cycles). (7 words)
- Variants: RT-PCR (RNA template), qPCR (quantitative). (5 words)
(Total body word count: 101 words excluding heading, mermaid, image caption text itself, but including text within bullets and blockquote.)
Sequencing & Arrays - Code Breakers!
- DNA Sequencing: Determines precise nucleotide order (A, T, C, G).
- Sanger Sequencing (Dideoxy Method):
- Uses ddNTPs to terminate DNA chain elongation.
- Separates fragments by size via electrophoresis. Gold standard for small scale.
- Next-Generation Sequencing (NGS):
- Massively parallel; sequences millions of fragments simultaneously.
- Applications: Whole genome sequencing (WGS), RNA-Seq.
- Sanger Sequencing (Dideoxy Method):
- DNA Microarrays (Gene Chips):
- Glass slides with thousands of specific DNA probes.
- Detects gene expression levels (cDNA from mRNA) or SNPs.
- Principle: Hybridization of fluorescently labeled nucleic acids.
⭐ Sanger sequencing is crucial for validating NGS findings and sequencing PCR products, despite lower throughput.

Gene Manipulation - Designer Genes!
- Definition: Intentional modification of an organism’s genetic material to achieve desired traits or study gene function.
- Core Techniques:
- Recombinant DNA Technology: Combining DNA from different sources using restriction enzymes and ligases.
- Site-Directed Mutagenesis: Introducing specific nucleotide changes to study protein structure/function.
- Gene Knockout/Knock-in: Deleting a gene (knockout) or replacing it with a modified version/reporter gene (knock-in) to assess its physiological role.
- CRISPR-Cas9: A powerful, precise, and versatile gene editing tool for adding, deleting, or altering DNA sequences.

- Applications in Physiology:
- Creating transgenic animal models of human diseases.
- Investigating gene function and regulation in physiological processes.
- Developing gene therapies for genetic disorders.
⭐ High-Yield Fact: CRISPR-Cas9 system, derived from a bacterial immune defense mechanism, allows for highly specific genomic modifications, revolutionizing functional genomics and therapeutic development in physiology research for conditions like Duchenne Muscular Dystrophy (DMD).
High‑Yield Points - ⚡ Biggest Takeaways
- PCR: Amplifies DNA for genetic diagnosis & infection detection (e.g., TB, viral loads).
- ELISA: Detects proteins/antibodies; for hormone assays & viral screening (e.g., HIV).
- Western Blot: Confirms specific proteins (e.g., HIV) using antibodies after separation.
- Southern Blot for DNA analysis; Northern Blot for RNA analysis (detect specific sequences).
- Flow Cytometry: Analyzes cell properties; for immunophenotyping & CD4 counts.
- FISH: Visualizes DNA/RNA in situ for chromosomal abnormalities, gene mapping.
- CRISPR-Cas9: Precise gene editing tool for targeted DNA modification.
Unlock the full lesson and continue reading
Signup to continue reading this lesson and unlimited access questions, flashcards, AI notes, and more

